Abstract
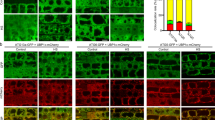
Arabidopsis thaliana histone deacetylase 1 (AtHD1 or AtHDA19), a homolog of yeast RPD3, is a global regulator of many physiological and developmental processes in plants. In spite of the genetic evidence for a role of AtHD1 in plant gene regulation and development, the biochemical and cellular properties of AtHD1 are poorly understood. Here we report cellular localization patterns of AtHD1 in vivo and histone deacetylase activity in vitro. The transient and stable expression of a green fluorescent protein (GFP)-tagged AtHD1 in onion cells and in roots, seeds and leaves of the transgenic Arabidopsis, respectively, revealed that AtHD1 is localized in the nucleus presumably in the euchromatic regions and excluded from the nucleolus. The localization patterns of AtHD1 are different from those of AtHD2 and AtHDA6 that are involved in nucleolus formation and silencing of transgenes and repeated DNA elements, respectively. In addition, a histone deacetylase activity assay showed that the recombinant AtHD1 produced in bacteria demonstrated a specific histone deacetylase activity in vitro. The data suggest that AtHD1 is a nuclear protein and possesses histone deacetylase activities responsible for global transcriptional regulation important to plant growth and development.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Brownell JE, Allis CD . Special HATs for special occasions: linking histone acetylation to chromatin assembly and gene activation. Curr Opin Genet Dev 1996; 6:176–184.
Jenuwein T, Allis CD . Translating the histone code. Science 2001; 293:1074–1080.
Berger SL . Histone modifications in transcriptional regulation. Curr Opin Genet Dev 2002; 12:142–148.
Taunton J, Hassig CA, Schreiber SL . A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science 1996; 272:408–511.
Pandey R, Muller A, Napoli CA, et al. Analysis of histone acetyltransferase and histone deacetylase families of Arabidopsis thaliana suggests functional diversification of chromatin modification among multicellular eukaryotes. Nucleic Acids Res 2002; 30:5036–5055.
Rundlett SE, Carmen AA, Kobayashi R, et al. HDA1 and RPD3 are members of distinct yeast histone deacetylase complexes that regulate silencing and transcription. Proc Natl Acad Sci U S A 1996; 93:14503–14508.
Loidl P . A plant dialect of the histone language. Trends Plant Sci 2004; 9:84–90.
Lusser A, Brosch G, Loidl A, Haas H, Loidl P . Identification of maize histone deacetylase HD2 as an acidic nucleolar phosphoprotein. Science 1997; 277:88–91.
Alland L, Muhle R, Hou H Jr, et al. Role for N-CoR and histone deacetylase in Sin3-mediated transcriptional repression. Nature 1997; 387:49–55.
Hassig CA, Fleischer TC, Billin AN, Schreiber SL, Ayer DE . Histone deacetylase activity is required for full transcriptional repression by mSin3A. Cell 1997; 89:341–347.
Kadosh D, Struhl K . Repression by Ume6 involves recruitment of a complex containing Sin3 corepressor and Rpd3 histone deacetylase to target promoters. Cell 1997; 89:365–371.
Vidal M, Gaber RF . RPD3 encodes a second factor required to achieve maximum positive and negative transcriptional states in Saccharomyces cerevisiae. Mol Cell Biol 1991; 11:6317–6327.
McKenzie EA, Kent NA, Dowell SJ, et al. The centromere and promoter factor, 1, CPF1, of Saccharomyces cerevisiae modulates gene activity through a family of factors including SPT21, RPD1 (SIN3), RPD3 and CCR4. Mol Gen Genet 1993; 240:374–386.
Stillman DJ, Dorland S, Yu Y . Epistasis analysis of suppressor mutations that allow HO expression in the absence of the yeast SW15 transcriptional activator. Genetics 1994; 136:781–788.
Jackson JC, Lopes JM . The yeast UME6 gene is required for both negative and positive transcriptional regulation of phospholipid biosynthetic gene expression. Nucleic Acids Res 1996; 24:1322–1329.
De Nadal E, Zapater M, Alepuz PM, et al. The MAPK Hog1 recruits Rpd3 histone deacetylase to activate osmoresponsive genes. Nature 2004; 427:370–374.
Stockinger EJ, Mao Y, Regier MK, Triezenberg SJ, Thomashow MF . Transcriptional adaptor and histone acetyltransferase proteins in Arabidopsis and their interactions with CBF1, a transcriptional activator involved in cold-regulated gene expression. Nucleic Acids Res 2001; 29:1524–1533.
Kim HJ, Hyun Y, Park JY, et al. A genetic link between cold responses and flowering time through FVE in Arabidopsis thaliana. Nat Genet 2004; 36:167–171.
Rossi V, Hartings H, Motto M . Identification and characterisation of an RPD3 homologue from maize (Zea mays L.) that is able to complement an rpd3 null mutant of Saccharomyces cerevisiae. Mol Gen Genet 1998; 258:288–296.
Murfett J, Wang XJ, Hagen G, Guilfoyle TJ . Identification of Arabidopsis histone deacetylase hda6 mutants that affect transgene expression. Plant Cell 2001; 13:1047–1061.
Aufsatz W, Mette MF, Van Der Winden J, Matzke M, Matzke AJ . HDA6, a putative histone deacetylase needed to enhance DNA methylation induced by double-stranded RNA. EMBO J 2002; 2002:6832–6841.
Lippman Z, May B, Yordan C, Singer T, Martienssen R . Distinct mechanisms determine transposon inheritance and methylation via small interfering RNA and histone modification. PLoS Biol 2003; 1:E67.
Probst AV, Fagard M, Proux F, et al. Arabidopsis histone deacetylase HDA6 is required for maintenance of transcriptional gene silencing and determines nuclear organization of rDNA repeats. Plant Cell 2004; 16:1021–1034.
Tian L, Wang J, Fong MP, et al. Genetic control of developmental changes induced by disruption of Arabidopsis histone deacetylase 1 (AtHD1) expression. Genetics 2003; 165:399–409.
Tian L, Chen ZJ . Blocking histone deacetylation in Arabidopsis induces pleiotropic effects on plant gene regulation and development. Proc Natl Acad Sci U S A 2001; 98:200–205.
Tian L, Fong MP, Wang JJ, et al. Reversible histone acetylation and deacetylation mediate genome-wide, promoter-dependent and locus-specific changes in gene expression during plant development. Genetics 2005; 169:337–345.
Zhou C, Zhang L, Duan J, Miki B, Wu K . HISTONE DEACETYLASE19 Is Involved in Jasmonic Acid and Ethylene Signaling of Pathogen Response in Arabidopsis. Plant Cell 2005; 17:1196–1204.
Clough SJ, Bent AF . Floral dip: a simlified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 1998; 16:735–743.
Bechtold N, Pelletier G . In planta Agrobacterium-mediated transformation of adult Arabidopsis thaliana plants by vacuum infiltration. Methods Mol Biol 1998; 82:259–266.
Bent AF, Clough SJ . Agrobacterium germ-line transformation: Transfromation of Arabidopsis without tissue culture. In: Gelvin SB, Schilperoort RA. Eds. Plant Molecular Biology Manual. Kluwer Academic Pub.: Dordrecht, The Netherlands 1998; 1–14.
Lawrence RJ, Earley K, Pontes O, et al. A concerted DNA methylation/histone methylation switch regulates rRNA gene dosage control and nucleolar dominance. Mol Cell 2004; 13:599–609.
Wu K, Malik K, Tian L, Brown D, Miki B . Functional analysis of a RPD3 histone deacetylase homologue in Arabidopsis thaliana. Plant Mol Biol 2000; 44:167–176.
He Y, Michaels SD, Amasino RM . Regulation of flowering time by histone acetylation in Arabidopsis. Science 2003; 302:1751–1754.
Ausin I, Alonso-Blanco C, Jarillo JA, Ruiz-Garcia L, Martinez-Zapater JM . Regulation of flowering time by FVE, a retinoblastoma-associated protein. Nat Genet 2004; 36:162–166.
He Y, Amasino RM . Role of chromatin modification in flowering-time control. Trends Plant Sci 2005; 10:30–35.
Grozinger CM, Schreiber SL . Regulation of histone deacetylase 4 and 5 and transcriptional activity by 14-3-3-dependent cellular localization. Proc Natl Acad Sci U S A 2000; 97:7835–7840.
McKinsey TA, Zhang CL, Olson EN . Activation of the myocyte enhancer factor-2 transcription factor by calcium/calmodulin-dependent protein kinase-stimulated binding of 14-3-3 to histone deacetylase 5. Proc Natl Acad Sci U S A 2000; 97:14400–14405.
Wang AH, Kruhlak MJ, Wu J, et al. Regulation of histone deacetylase 4 by binding of 14-3-3 proteins. Mol Cell Biol 2000; 20:6904–6912.
Kao HY, Verdel A, Tsai CC, et al. Mechanism for nucleocytoplasmic shuttling of histone deacetylase 7. J Biol Chem 2001; 276:47496–47507.
Miska EA, Karlsson C, Langley E, et al. HDAC4 deacetylase associates with and represses the MEF2 transcription factor. Embo J 1999; 18:5099–5107.
Sengupta N, Seto E . Regulation of histone deacetylase activities. J Cell Biochem 2004; 93:57–67.
Baek SH, Ohgi KA, Rose DW, et al. Exchange of N-CoR corepressor and Tip60 coactivator complexes links gene expression by NF-kappaB and beta-amyloid precursor protein. Cell 2002; 110:55–67.
Varotto S, Locatelli S, Canova S, et al. Expression profile and cellular localization of maize Rpd3-type histone deacetylases during plant development. Plant Physiol 2003; 133:606–617.
Heim R, Prasher DC, Tsien RY . Wavelength mutations and posttranslational autoxidation of green fluorescent protein. Proc Natl Acad Sci U S A 1994; 91:12501–12504.
Chiu W, Niwa Y, Zeng W, et al. Engineered GFP as a vital reporter in plants. Curr Biol 1996; 6:325–330.
Koroleva OA, Tomlinson ML, Leader D, Shaw P, Doonan JH . High-throughput protein localization in Arabidopsis using Agrobacterium-mediated transient expression of GFP-ORF fusions. Plant J 2005; 41:162–174.
Grebenok RJ, Pierson E, Lambert GM, et al. Green-fluorescent protein fusions for efficient characterization of nuclear targeting. Plant J 1997; 11:573–586.
Haasen D, Kohler C, Neuhaus G, Merkle T . Nuclear export of proteins in plants: AtXPO1 is the export receptor for leucine-rich nuclear export signals in Arabidopsis thaliana. Plant J 1999; 20:695–705.
Kruhlak MJ, Hendzel MJ, Fischle W, et al. Regulation of global acetylation in mitosis through loss of histone acetyltransferases and deacetylases from chromatin. J Biol Chem 2001; 276:38307–38319.
Cimini D, Mattiuzzo M, Torosantucci L, Degrassi F . Histone hyperacetylation in mitosis prevents sister chromatid separation and produces chromosome segregation defects. Mol Biol Cell 2003; 14:3821–3833.
Li DW, Yang Q, Chen JT, et al. Dynamic distribution of Ser-10 phosphorylated histone H3 in cytoplasm of MCF-7 and CHO cells during mitosis. Cell Res 2005; 15:120–126.
Brosch G, Georgieva EI, Lopez-Rodas G, Lindner H, Loidl P . Specificity of Zea mays histone deacetylase is regulated by phosphorylation. J Biol Chem 1992; 267:20561–20564.
Cai R, Kwon P, Yan-Neale Y, et al. Mammalian histone deacetylase 1 protein is posttranslationally modified by phosphorylation. Biochem Biophys Res Commun 2001; 283:445–453.
Pflum MK, Tong JK, Lane WS, Schreiber SL . Histone deacetylase 1 phosphorylation promotes enzymatic activity and complex formation. J Biol Chem 2001; 276:47733–47741.
Tsai SC, Seto E . Regulation of histone deacetylase 2 by protein kinase CK2. J Biol Chem 2002; 277:31826–31833.
Colombo R, Boggio R, Seiser C, Draetta GF, Chiocca S . The adenovirus protein Gam1 interferes with sumoylation of histone deacetylase 1. EMBO Rep 2002; 3:1062–1068.
David G, Neptune MA, DePinho RA . SUMO-1 modification of histone deacetylase 1 (HDAC1) modulates its biological activities. J Biol Chem 2002; 277:23658–663.
Kirsh O, Seeler JS, Pichler A, et al. The SUMO E3 ligase RanBP2 promotes modification of the HDAC4 deacetylase. Embo J 2002; 21:2682–2691.
Pipal A, Goralik-Schramel M, Lusser A, et al. Regulation and processing of maize histone deacetylase Hda1 by limited proteolysis. Plant Cell 2003; 15:1904–1917.
Carmen AA, Griffin PR, Calaycay JR, et al. Yeast HOS3 forms a novel trichostatin A-insensitive homodimer with intrinsic histone deacetylase activity. Proc Natl Acad Sci U S A 1999; 96:12356–12361.
Hu E, Chen Z, Fredrickson T, et al. Cloning and characterization of a novel human class I histone deacetylase that functions as a transcription repressor. J Biol Chem 2000; 275:15254–15264.
Lee H, Rezai-Zadeh N, Seto E . Negative regulation of histone deacetylase 8 activity by cyclic AMP-dependent protein kinase A. Mol Cell Biol 2004; 24:765–773.
Acknowledgements
We thank Mary Bryk and Timothy Hall for critical suggestions to improve the manuscript, David Stelly and Keerti Rathore for assistance in GFP localization studies in onion cells, and Stanislav Vitha in the Microscopy and Imaging Center at Texas A&M University for technical support for epifluorescence microscopic image analysis in the transgenic plants. The work is supported by grants from the National Institutes of Health (GM067015) and the National Science Foundation Plant Genome Research Program (DBI0077774) to Z J C.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fong, P., Tian, L. & Chen, Z. Arabidopsis thaliana histone deacetylase 1 (AtHD1) is localized in euchromatic regions and demonstrates histone deacetylase activity in vitro. Cell Res 16, 479–488 (2006). https://doi.org/10.1038/sj.cr.7310059
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/sj.cr.7310059
Keywords
This article is cited by
-
An efficient chromatin immunoprecipitation (ChIP) protocol for studying histone modifications in peach reproductive tissues
Plant Methods (2022)
-
Cloning PIP genes in drought-tolerant vetiver grass and responses of transgenic VzPIP2;1 soybean plants to water stress
Biologia plantarum (2016)
-
Parallel action of AtDRB2 and RdDM in the control of transposable element expression
BMC Plant Biology (2015)
-
Histone deacetylases and their functions in plants
Plant Cell Reports (2013)
-
Role of HD2 genes in seed germination and early seedling growth in Arabidopsis
Plant Cell Reports (2011)