Abstract
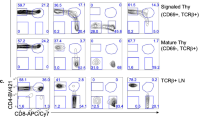
CD8 engagement with class I major histocompatibility antigens greatly enhances T-cell activation, but it is not clear how this is achieved. We address the question of whether or not the antibody-mediated ligation of CD8 alone induces transcriptional remodeling in a T-cell clone, using serial analysis of gene expression. Even though it fails to induce overt phenotypic changes, we find that CD8 ligation profoundly alters transcription in the T-cell clone, at a scale comparable to that induced by antibody-mediated ligation of CD3. The character of the resulting changes is distinct, however, with the net effect of CD8 ligation being substantially inhibitory. We speculate that ligating CD8 induces weak, T-cell receptor (TCR)-mediated inhibitory signals reminiscent of the effects of TCR antagonists. Our results imply that CD8 ligation alone is incapable of activating the T-cell clone because it fails to fully induce NFAT-dependent transcription.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Holler PD, Kranz DM . Quantitative analysis of the contribution of TCR/pepMHC affinity and CD8 to T cell activation. Immunity 2003; 18:255–264.
Wyer JR, Willcox BE, Gao GF, et al. T cell receptor and coreceptor CD8 alphaalpha bind peptide-MHC independently and with distinct kinetics. Immunity 1999; 10:219–225.
Davis SJ, Ikemizu S, Evans EJ, Fugger L, Bakker TR, van der Merwe PA . The nature of molecular recognition by T cells. Nat Immunol 2003; 4:217–224.
Leishman AJ, Naidenko OV, Attinger A, et al. T cell responses modulated through interaction between CD8alphaalpha and the nonclassical MHC class I molecule, TL. Science 2001; 294:1936–1939.
Tsujimura K, Obata Y, Matsudaira Y, et al. The binding of thymus leukemia (TL) antigen tetramers to normal intestinal intraepithelial lymphocytes and thymocytes. J Immunol 2001; 167:759–764.
Spruyt LL, Glennie MJ, Beyers AD, Williams AF . Signal transduction by the CD2 antigen in T cells and natural killer cells: requirement for expression of a functional T cell receptor or binding of antibody Fc to the Fc receptor, Fc gamma RIIIA (CD16). J Exp Med 1991; 174:1407–1415.
Luhder F, Huang Y, Dennehy KM, et al. Topological requirements and signaling properties of T cell-activating, anti-CD28 antibody superagonists. J Exp Med 2003; 197:955–966.
Moss PA, Rowland-Jones SL, Frodsham PM, et al. Persistent high frequency of human immunodeficiency virus-specific cytotoxic T cells in peripheral blood of infected donors. Proc Natl Acad Sci USA 1995; 92:5773–5777.
Evans EJ, Hene L, Sparks LM, et al. The T cell surface – how well do we know it? Immunity 2003; 19:213–223.
Velculescu VE, Zhang L, Vogelstein B, Kinzler KW . Serial analysis of gene expression. Science 1995; 270:484–487.
Beyers AD, Spruyt LL, Williams AF . Molecular associations between the T-lymphocyte antigen receptor complex and the surface antigens CD2, CD4, or CD8 and CD5. Proc Natl Acad Sci USA 1992; 89:2945–2949.
Gygi SP, Rochon Y, Franza BR, Aebersold R . Correlation between protein and mRNA abundance in yeast. Mol Cell Biol 1999; 19:1720–1730.
Williams AF, Barclay AN . Glycoprotein antigens of the lymphocyte surface and their purification by antibody affinity chromatography. In: Weir DM, Herzenberg LA, eds. Handbook of experimental immunology. Oxford: Blackwell Scientific Publishing, 1985: 22.1–22.24.
Wahl SM, Wen J, Moutsopoulos N . TGF-beta: a mobile purveyor of immune privilege. Immunol Rev 2006; 213:213–227.
Zhong XP, Hainey EA, Olenchock BA, et al. Enhanced T cell responses due to diacylglycerol kinase zeta deficiency. Nat Immunol 2003; 4:882–890.
Zhu J, Shibasaki F, Price R, et al. Intramolecular masking of nuclear import signal on NF-AT4 by casein kinase I and MEKK1. Cell 1998; 93:851–61.
Gao GF, Rao Z, Bell JI . Molecular coordination of alphabeta T-cell receptors and coreceptors CD8 and CD4 in their recognition of peptide-MHC ligands. Trends Immunol 2002; 23:408–413.
Doucey MA, Goffin L, Naeher D, et al. CD3 delta establishes a functional link between the T cell receptor and CD8. J Biol Chem 2003; 278:3257–3264.
Davis SJ, van der Merwe PA . The kinetic-segregation model: TCR triggering and beyond. Nat Immunol 2006; 7:803–809.
Klenerman P, Rowland-Jones S, McAdam S, et al. Cytotoxic T-cell activity antagonized by naturally occurring HIV-1 Gag variants. Nature 1994; 369:403–407.
Sykulev Y, Vugmeyster Y, Brunmark A, Ploegh HL, Eisen HN . Peptide antagonism and T cell receptor interactions with peptide-MHC complexes. Immunity 1998; 9:475–483.
Zhao G, Shi L, Qiu D, Hu H, Kao PN . NF45/ILF2 tissue expression, promoter analysis, and interleukin-2 transactivating function. Exp Cell Res 2005; 305:312–323.
Reichman TW, Muniz LC, Mathews MB . The RNA binding protein nuclear factor 90 functions as both a positive and negative regulator of gene expression in mammalian cells. Mol Cell Biol 2002; 22:343–356.
Shim J, Lim HR, Yates J, Karin M . Nuclear export of NF90 is required for interleukin-2 mRNA stabilization. Mol Cell 2002; 10:1331–1344.
Dittel BN, Stefanova I, Germain RN, Janeway CA Jr . Cross-antagonism of a T cell clone expressing two distinct T cell receptors. Immunity 1999; 11:289–298.
Fong SS, Joyce AR, Palsson BO . Parallel adaptive evolution cultures of Escherichia coli lead to convergent growth phenotypes with different gene expression states. Genome Res 2005; 15:1365–1372.
Stern S, Dror T, Stolovicki E, Brenner N, Braun E . Genome-wide transcriptional plasticity underlies cellular adaptation to novel challenge. Mol Syst Biol 2007; 3:106.
Pelosi L, Kuhn L, Guetta D, et al. Parallel changes in global protein profiles during long-term experimental evolution in Escherichia coli. Genetics 2006; 173:1851–1869.
Koonin EV . Chance and necessity in cellular response to challenge. Mol Syst Biol 2007; 3:107.
Riley JL, Mao M, Kobayashi S, et al. Modulation of TCR-induced transcriptional profiles by ligation of CD28, ICOS, and CTLA-4 receptors. Proc Natl Acad Sci USA 2002; 99:11790–11795.
Diehn M, Alizadeh AA, Rando OJ, et al. Genomic expression programs and the integration of the CD28 costimulatory signal in T cell activation. Proc Natl Acad Sci USA 2002; 99:11796–11801.
McMichael AJ, Beverley PCL, Cobbold S, et al., eds. Leucocyte typing III: white cell differentiation antigens. Oxford: Oxford University Press, 1987.
Audic S, Claverie JM . The significance of digital gene expression profiles. Genome Res 1997; 7:986–995.
Romualdi C, Bortoluzzi S, Danieli GA . Detecting differentially expressed genes in multiple tag sampling experiments: comparative evaluation of statistical tests. Hum Mol Genet 2001; 10:2133–2141.
Lash AE, Tolstoshev CM, Wagner L, et al. SAGEmap: a public gene expression resource. Genome Res 2000; 10:1051–1060.
Boon K, Osorio EC, Greenhut SF, et al. An anatomy of normal and malignant gene expression. Proc Natl Acad Sci USA 2002; 99:11287–11292.
Wheeler DL, Barrett T, Benson DA, et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 2006; 34:D173–D180.
Pruitt KD, Maglott DR . RefSeq and LocusLink: NCBI gene-centered resources. Nucleic Acids Res 2001; 29:137–140.
Riggins GJ, Strausberg RL . Genome and genetic resources from the Cancer Genome Anatomy Project. Hum Mol Genet 2001; 10:663–667.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Abidi, S., Dong, T., Vuong, M. et al. Differential remodeling of a T-cell transcriptome following CD8- versus CD3-induced signaling. Cell Res 18, 641–648 (2008). https://doi.org/10.1038/cr.2008.56
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/cr.2008.56