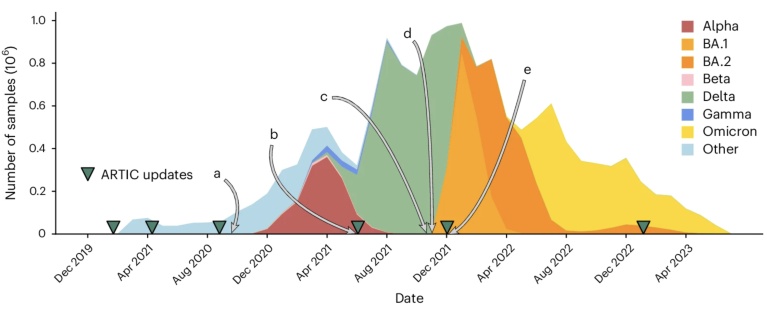
Key moments in the COVID-19 pandemic when sequencing errors muddled the viral family tree, issues now corrected by the Viridian assembler. Credit: Hunt, M. et al. Nat. Methods (2026)
During the COVID-19 pandemic, scientists sequenced millions of SARS-CoV-2 genomes by breaking them into overlapping fragments, a method that was fast but prone to errors. When mutations disrupted these fragments, many assembly pipelines simply filled the gaps with the original reference sequence. The result: false “reversions” that scrambled the virus’s family tree, skewed growth-rate estimates, and exaggerated the number of viral introductions.
Now, an international team of researchers — including scientists from India — has tackled the problem with a new genome assembler called Viridian1. Unlike earlier methods, Viridian identifies the fragment layout for each sample, assembles the genome piece by piece using a robust alignment approach, and rigorously checks for errors.
The team reprocessed more than 4.4 million SARS-CoV-2 genomes from Illumina, Nanopore, and Ion Torrent datasets. Compared with existing GenBank assemblies, Viridian drastically reduced errors, smoothed out misleading reversions, and tripled the precision of growth-rate estimates. In countries with heavy sampling, like the United States, it also cut the number of falsely inferred viral introductions.
Beyond COVID-19, Viridian offers a standardized, high-accuracy workflow for sequencing viruses with limited structural variation. By unifying assembly and quality control, it lays a foundation for evolutionary studies, epidemiology, and rapid genomic surveillance in future outbreaks.
