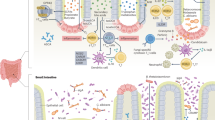
Abstract
The healthy intestine is characterized by a low level of oxygen and by the presence of large bacterial communities of obligate anaerobes. Dysbiosis of the gut microbiota has been reported in patients suffering from inflammatory bowel diseases (IBDs), but the mechanisms causing this imbalance remain unknown. Observations have included a decrease in obligate anaerobes of the phylum Firmicutes and an increase in facultative anaerobes, including members of the family Enterobacteriaceae. The shift of bacterial communities from obligate to facultative anaerobes strongly suggests a disruption in anaerobiosis and points to a role for oxygen in intestinal dysbiosis. Proposals to evaluate this hypothesis of a role for oxygen in IBD dysbiosis are provided. If this hypothesis is confirmed, decreasing oxygen in the intestine could open novel means to rebalance the microbiota and could provide novel preventative or therapeutic strategies for IBD patients in whom current treatments are ineffective.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Adlerberth I, Wold AE . (2009). Establishment of the gut microbiota in Western infants. Acta Paediatr 98: 229–238.
Arumugam M, Raes J, Pelletier E, Le Paslier D, Yamada T, Mende DR et al (2011). Enterotypes of the human gut microbiome. Nature 473: 174–180.
Baughn AD, Malamy MH . (2004). The strict anaerobe Bacteroides fragilis grows in and benefits from nanomolar concentrations of oxygen. Nature 427: 441–444.
Baumgart M, Dogan B, Rishniw M, Weitzman G, Bosworth B, Yantiss R et al (2007). Culture independent analysis of ileal mucosa reveals a selective increase in invasive Escherichia coli of novel phylogeny relative to depletion of Clostridiales in Crohn’s disease involving the ileum. ISME J 1: 403–418.
Corthier G, Muller MC, L'Haridon R . (1996). Selective enumeration of Bacteroides vulgatus and B. distasonis organisms in the predominant human fecal flora by using monoclonal antibodies. Appl Environ Microbiol 62: 735–738.
Darfeuille-Michaud A, Boudeau J, Bulois P, Neut C, Glasser AL, Barnich N et al (2004). High prevalence of adherent-invasive Escherichia coli associated with ileal mucosa in Crohn’s disease. Gastroenterology 127: 412–421.
Duncan SH, Louis P, Flint HJ . (2007). Cultivable bacterial diversity from the human colon. Lett Appl Microbiol 44: 343–350.
Erickson AR, Cantarel BL, Lamendella R, Darzi Y, Mongodin EF, Pan C et al (2012). Integrated metagenomics/metaproteomics reveals human host-microbiota signatures of Crohn’s disease. PLoS One 7: e49138.
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR . (2007). Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci USA 104: 13780–13785.
Grutte FK, Horn R, Haenel H . (1965). Ernahrung und biochemischmikrookologische voorgange in enddarm von sauglingen (Nutrition and biochemical microecology processes occuring in the colon of infants). Z Kinderheikld 93: 28–39.
Handa K, Ohmura M, Nishime C, Hishiki T, Nagahata Y, Kawai K et al (2010). Phosphorescence-assisted microvascular O(2) measurements reveal alterations of oxygen demand in human metastatic colon cancer in the liver of superimmunodeficient NOG mice. Adv Exp Med Biol 662: 423–429.
Hartman AL, Lough DM, Barupal DK, Fiehn O, Fishbein T, Zasloff M et al (2009). Human gut microbiome adopts an alternative state following small bowel transplantation. Proc Natl Acad Sci USA 106: 17187–17192.
He G, Shankar RA, Chzhan M, Samouilov A, Kuppusamy P, Zweier JL . (1999). Noninvasive measurement of anatomic structure and intraluminal oxygenation in the gastrointestinal tract of living mice with spatial and spectral EPR imaging. Proc Natl Acad Sci USA 96: 4586–4591.
Khan MT, Duncan SH, Stams AJ, van Dijl JM, Flint HJ, Harmsen HJ . (2012). The gut anaerobe Faecalibacterium prausnitzii uses an extracellular electron shuttle to grow at oxic-anoxic interphases. ISME J 6: 1578–1585.
Kotlowski R, Bernstein CN, Sepehri S, Krause DO . (2007). High prevalence of Escherichia coli belonging to the B2+D phylogenetic group in inflammatory bowel disease. Gut 56: 669–675.
Lepage P, Hasler R, Spehlmann ME, Rehman A, Zvirbliene A, Begun A et al (2011). Twin study indicates loss of interaction between microbiota and mucosa of patients with ulcerative colitis. Gastroenterology 141: 227–236.
Lind Due V, Bonde J, Kann T, Perner A . (2003). Extremely low oxygen tension in the rectal lumen of human subjects. Acta Anaesth Scand 47: 372.
Lozupone CA, Stombaugh JI, Gordon JI, Jansson JK, Knight R . (2012). Diversity, stability and resilience of the human gut microbiota. Nature 489: 220–230.
Lupp C, Robertson ML, Wickham ME, Sekirov I, Champion OL, Gaynor EC et al (2007). Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe 2: 204.
Macho Fernandez E, Pot B, Grangette C . (2011). Beneficial effect of probiotics in IBD: are peptidogycan and NOD2 the molecular key effectors? Gut Microbes 2: 280–286.
Mackie RI, Sghir A, Gaskins HR . (1999). Developmental microbial ecology of the neonatal gastrointestinal tract. Am J Clin Nutr 69: 1035S–1045S.
Manichanh C, Rigottier-Gois L, Bonnaud E, Gloux K, Pelletier E, Frangeul L et al (2006). Reduced diversity of faecal microbiota in Crohn’s disease revealed by a metagenomic approach. Gut 55: 205–211.
Manichanh C, Borruel N, Casellas F, Guarner F . (2012). The gut microbiota in IBD. Nature reviews Gastroenterology & hepatology 9: 599–608.
Meynell GG . (1963). Antibacterial mechanisms of the mouse gut. II. The role of Eh and volatile fatty acids in the normal gut. Br J Exper Pathol 44: 209–219.
Mondot S, Kang S, Furet JP, Aguirre de Carcer D, McSweeney C, Morrison M et al (2011). Highlighting new phylogenetic specificities of Crohn’s disease microbiota. Inflamm Bowel Dis 17: 185–192.
Mukhopadhya I, Hansen R, El-Omar EM, Hold GL . (2012). IBD-what role do Proteobacteria play? Nat Rev Gastroenterol Hepatol 9: 219–230.
Nagalingam NA, Lynch SV . (2012). Role of the microbiota in inflammatory bowel diseases. Inflamm Bowel Dis 18: 968–984.
Naser SA, Ghobrial G, Romero C, Valentine JF . (2004). Culture of Mycobacterium avium subspecies paratuberculosis from the blood of patients with Crohn’s disease. Lancet 364: 1039–1044.
Ott SJ, Musfeldt M, Wenderoth DF, Hampe J, Brant O, Folsch UR et al (2004). Reduction in diversity of the colonic mucosa associated bacterial microflora in patients with active inflammatory bowel disease. Gut 53: 685–693.
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C et al (2010). A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464: 59–65.
Sartor RB . (2008). Microbial influences in inflammatory bowel diseases. Gastroenterology 134: 577–594.
Sekirov I, Russell SL, Antunes LC, Finlay BB . (2010). Gut microbiota in health and disease. Physiol Rev 90: 859–904.
Seksik P, Rigottier-Gois L, Gramet G, Sutren M, Pochart P, Marteau P et al (2003). Alterations of the dominant faecal bacterial groups in patients with Crohn’s disease of the colon. Gut 52: 237–242.
Sokol H, Seksik P, Rigottier-Gois L, Lay C, Lepage P, Podglajen I et al (2006). Specificities of the fecal microbiota in inflammatory bowel disease. Inflamm Bowel Dis 12: 106–111.
Sokol H, Pigneur B, Watterlot L, Lakhdari O, Bermudez-Humaran LG, Gratadoux JJ et al (2008). Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc Natl Acad Sci USA 105: 16731–16736.
Sokol H, Seksik P, Furet JP, Firmesse O, Nion-Larmurier I, Beaugerie L et al (2009). Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm Bowel Dis 15: 1183–1189.
Swidsinski A, Loening-Baucke V, Theissig F, Engelhardt H, Bengmark S, Koch S et al (2007). Comparative study of the intestinal mucus barrier in normal and inflamed colon. Gut 56: 343–350.
Swidsinski A, Loening-Baucke V, Verstraelen H, Osowska S, Doerffel Y . (2008). Biostructure of fecal microbiota in healthy subjects and patients with chronic idiopathic diarrhea. Gastroenterology 135: 568–579.
Van Nuenen HMC . (2005), Composition and Activity of Gut Microbiota in Inflammatory Bowel Disease. PhD thesis, Erasmus University Rotterdam, The Netherlands.
Willing B, Halfvarson J, Dicksved J, Rosenquist M, Jarnerot G, Engstrand L et al (2009). Twin studies reveal specific imbalances in the mucosa-associated microbiota of patients with ileal Crohn’s disease. Inflamm Bowel Dis 15: 653–660.
Willing BP, Dicksved J, Halfvarson J, Andersson AF, Lucio M, Zheng Z et al (2010). A pyrosequencing study in twins shows that gastrointestinal microbial profiles vary with inflammatory bowel disease phenotypes. Gastroenterology 139: 1844–1854e1.
Winter SE, Thiennimitr P, Winter MG, Butler BP, Huseby DL, Crawford RW et al (2010). Gut inflammation provides a respiratory electron acceptor for Salmonella. Nature 467: 426–429.
Wu GD, Chen J, Hoffmann C, Bittinger K, Chen YY, Keilbaugh SA et al (2011). Linking long-term dietary patterns with gut microbial enterotypes. Science 334: 105–108.
Xavier RJ, Podolsky DK . (2007). Unravelling the pathogenesis of inflammatory bowel disease. Nature 448: 427–434.
Yong E . (2012). Gut microbial ‘enterotypes’ become less clear-cut. Nature News (e-pub ahead of print 21 March 2012 doi:10.1038/nature.2012.10276).
Zhu H, Li YR . (2012). Oxidative stress and redox signaling mechanisms of inflammatory bowel disease: updated experimental and clinical evidence. Exp Biol Med 237: 474–480.
Acknowledgements
I thank Dr William Blackhall for the English editing and revision of the text.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no conflict of interest.
Rights and permissions
About this article
Cite this article
Rigottier-Gois, L. Dysbiosis in inflammatory bowel diseases: the oxygen hypothesis. ISME J 7, 1256–1261 (2013). https://doi.org/10.1038/ismej.2013.80
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ismej.2013.80
Keywords
This article is cited by
-
Metabolic network of the gut microbiota in inflammatory bowel disease
Inflammation and Regeneration (2024)
-
An in vitro platform for study of the human gut microbiome under an oxygen gradient
Biomedical Microdevices (2023)
-
Mapping microbiome-redox spectrum and evaluating Microbial-Redox Index in chronic gastritis
Scientific Reports (2022)
-
Impact of gut Microbiome alteration in Ulcerative Colitis patients on disease severity and outcome
Clinical and Experimental Medicine (2022)
-
Quantitative sequencing clarifies the role of disruptor taxa, oral microbiota, and strict anaerobes in the human small-intestine microbiome
Microbiome (2021)