Abstract
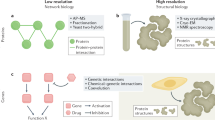
Structural genomics projects aim to solve the experimental structures of all possible protein folds. Such projects entail a conceptual shift from traditional structural biology in which structural information is obtained on known proteins to one in which the structure of a protein is determined first and the function assigned only later. Whereas the goal of converting protein structure into function can be accomplished by traditional sequence motif-based approaches, recent studies have shown that assignment of a protein's biochemical function can also be achieved by scanning its structure for a match to the geometry and chemical identity of a known active site. Importantly, this approach can use low-resolution structures provided by contemporary structure prediction methods. When applied to genomes, structural information (either experimental or predicted) is likely to play an important role in high-throughput function assignment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Clark, M.S. Comparative genomics: the key to understanding the Human Genome Project. Bioessays 21, 121–130 ( 1999).
DellaPenna, D. Nutritional genomics: manipulating plant micronutrients to improve human health . Science 285, 375–379 (1999).
Wiley, S.R. Genomics in the real world. Curr. Pharm. Des. 4, 417–422 (1998).
Lin, J. et al. Whole-genome shotgun optical mapping of Deinococcus radiodurans . Science 285, 1558– 1562 (1999).
Carulli, J. P. et al. High throughput analysis of differential gene expression. J. Cell Biochem. Suppl. 31, 286–96 (1998).
Chothia, C. & Finkelstein, A. The classification and origins of protein folding patterns. Annu. Rev. Biochem. 59 , 1007–1039 (1990).
Murzin, A.G., Lesk, A.M. & Chothia, C. Principles determining the structure of beta-sheet barrels in proteins. II. The observed structures. J. Mol. Biol. 236, 1382–1400 (1994).
Chothia, C., Hubbard, T., Brenner, S., Barns, H. & Murzin, A. Protein folds in the all-beta and all-alpha classes. Annu. Rev. Biophys. Biomol. Struct. 26, 597– 627 (1997).
Sali, A. 100,000 protein structures for the biologist (see comments). Nat. Struct. Biol. 5, 1029–1032 ( 1998).
Holm, L. & Sander, C. Protein folds and families: sequence and structure alignments. Nucleic Acids Res. 27, 244–247 (1999).
Dodge, C., Schneider, R. & Sander, C. The HSSP database of protein structure-sequence alignments and family profiles. Nucleic Acids Res. 26, 313–315 (1998).
Holm, L. & Sander, C. Dali/FSSP classification of three-dimensional folds. Nucleic Acids Res. 25, 231– 234 (1997).
Orengo, C.A. et al. CATH—a hierarchic classification of protein domain structures . Structure 5, 1093–1108 (1997).
Murzin, A.G., Brenner, S.E., Hubbard, T. & Chothia, C. Scop: a structural classification of proteins database for the investigation of sequences and structures. J. Mol. Biol. 247, 536–540 (1995).
Sanchez, R. & Sali, A. Evaluation of comparative protein structure modeling by MODELLER-3. Proteins Suppl. 50– 58 (1997).
Briem, H. & Kuntz, I.D. Molecular similarity based on DOCK-generated fingerprints. J. Med. Chem. 39, 3401– 3408 (1996).
Fetrow, J.S. & Skolnick, J. Method for prediction of protein function from sequence using the sequence-to-structure-to-function paradigm with application to glutaredoxins/ thioredoxins and T1 ribonucleases. J. Mol. Biol. 281, 949–968 (1998).
Fetrow, J.S., Godzik, A. & Skolnick, J. Functional analysis of the Escherichia coli genome using the sequence-to-structure-to-function paradigm: identification of proteins exhibiting the glutaredoxin/thioredoxin disulfide oxidoreductase activity . J. Mol. Biol. 282, 703– 711 (1998).
Fetrow, J.S., Siew, N. & Skolnick, J. Structure-based functional motif identifies a potential disulfide oxidoreductase active site in the serine/threonine protein phosphatase-1 subfamily. FASEB J. 13, 1866– 1874 (1999).
Zhang, L., Godzik, A., Skolnick, J. & Fetrow, J.S. Functional analysis of E. coli proteins for members of the a/b hydrolase family. Folding and Design 3, 535–548 (1998).
Orengo, C.A., Todd, A.E. & Thornton, J.M. From protein structure to function. Curr. Opin. Struct. Biol. 9, 374–382 (1999).
Montelione, G.T. & Anderson, S. Structural genomics: keystone for a Human Proteome Project (news). Nat. Struct. Biol. 6, 11–12 (1999 ).
Kim, S.H. Shining a light on structural genomics. Nat. Struct. Biol. 5 Suppl, 643–645 ( 1998).
Gaasterland, T. Structural genomics taking shape. Trends Genet. 14, 135 (1998).
Sanchez, R. & Sali, A. Large-scale protein structure modeling of the Saccharomyces cerevisiae genome. Proc. Natl. Acad. Sci. USA 95, 13597–13602 (1998).
Terwilliger, T.C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999).
Wallin, E. & Heijne, G.V. Genome-wide analysis of intergral membrane proteins from eubacterial, archaen, and eukaryotic organismc. Prot. Sci. 7, 1029–1038 (1998).
Goffeau, A. et al. Life with 6000 genes (see comments). Science 274, 546, 563–567 ( 1996).
Elofsson, A. & Sonnhammer, E.L. A comparison of sequence and structure protein domain families as a basis for structural genomics. Bioinformatics 15, 480–500 (1999).
Rost, B., Schneider, R. & Sander, C. Protein fold recognition by prediction-based threading . J. Mol. Biol. 270, 471– 480 (1997).
Jones, D.T. GenTHREADER: an efficient and reliable protein fold recognition method for genomic sequences. J. Mol. Biol. 287, 797 –815 (1999).
Marchler-Bauer, A. & Brenner, S. Comparison of prediction quality in the three CASPs. Proteins Suppl. 3, 218–225 (1999).
Fischer, D. & Eisenberg, D. Assigning folds to the proteins encoded by the genome of Mycoplasma genitalium. Proc. Natl. Acad Sci USA 94, 11929–11934 (1997).
Kolinski, A., Rotkiewicz, P., Ilkowski, I. & Skolnick, J. A method for the improvement of threading based protein models. Proteins 37, 592–610 ( 1999).
Lee, J., Liwo, A., Ripoll, D.R., Pillardy, J. & Scheraga, H.A. Calculation of protein conformation by global optimiation of a potential energy function. Proteins Suppl. 3, 204–208 (1999).
Simons, K.T., Bonneau, R., Ruczinski, I. & Baker, D. Ab initio structure prediction of CASP III targets using ROSETTA. Proteins Suppl. 3, 171–176 (1999).
Ortiz, A., Kolinski, A., Rotkiewicz, P., Ilkowski, B. & Skolnick, J. Ab intio folding of proteins using restraints derived from evolutionary information. Proteins Suppl. 3, 177–185 ( 1999).
Osguthorpe, D.J. Improved ab initio predictions with a simplified, flexible geometry model . Proteins Suppl. 3, 186– 193 (1999).
Samudrala, R., Xia, Y., Huang, E. & Levitt, M. Ab initio protein structure prediction using a combined hierarchical approach. Proteins Suppl 3, 194–198 ( 1999).
Orengo, C., Bray, J.E., LoConte, L. & Sillitoe, I. Analysis and assessment of ab initio three-dimensional prediction, secondary structure and contacts prediction. Proteins Suppl. 3, 149–170 (1999).
Murzin, A. Structure classification-based assessement of CASP3 prediction for the fold recognition targets. Proteins Suppl. 3, 88–103 (1999).
Venclovas, C., Zemla, A., Fidelis, K. & Moult, J. Some measures of comparative performance in the three CASPs. Proteins Suppl. 3, 231–227 (1999).
Brutlag, D.L. Genomics and computational molecular biology. Curr. Opin. Microbiol. 1, 340–345 ( 1998).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Pearson, W.R. Empirical statistical estimates for sequence similarity searches. J. Mol. Biol. 276, 71–84 (1998).
Brenner, S.E. Errors in genome annotation. Trends Genet. 15, 132–133 (1999).
Attwood, T.K. et al. Novel developments with the PRINTS protein fingerprint database . Nucleic Acids Res. 25, 212– 216 (1997).
Bairoch, A. Prosite: a dictionary of sites and patterns in proteins. Nucleic Acids Res. Suppl, 19, 2241–2245 (1991).
Henikoff, J.G., Henikoff, S. & Pietrokovski, S. New features of the Blocks database servers. Nucleic Acids Res. 27, 226–228 (1999).
Hofmann, K., Bucher, P., Falquet, L. & Bairoch, A. The Prosite database, its status in 1999. Nucleic Acids Res. 27, 215–219 (1999).
Pietrovski, S., Henikoff, J.G. & Henikoff, S. The Blocks database—a system for protein classification . Nucleic Acids Res. 24, 197– 200 (1996).
Yu, L., White, J.V. & Smith, T.F. A homology identification method that combines protein sequence and structure information. Protein Sci. 7, 2499–2510 (1998).
Kasuya, A. & Thornton, J.M. Three-dimensional structure analyis of Prosite patterns. J. Mol. Biol. 286, 1673–1691 (1999).
Hegyi, H. & Gerstein, M. The relationship between protein structure and function: a comprehensive survey with application to the yeast genome. J. Mol. Biol. 288, 147– 164 (1999).
Wallace, A.C., Laskowski, R.A. & Thornton, J.M. Derivation of 3D coordinate templates for searching structural databases: application to Ser-His-Asp catalytic triads in the serine proteinases and lipases. Protein Sci. 5, 1001–1013 (1996).
Fischer, D., Wolfson, H., Lin, S.L. & Nussinov, R. Three-dimensional, sequence order-independent structural comparison of a serine protease against the crystallographic database reveals active site similarities: potential implications to evolution and to protein folding. Protein Sci. 3, 769–778 ( 1994).
Matthews, D.A. et al. Structure of human rhinovirus 3C protease reveals a trypsin-like polypeptide fold, RNA-binding site, and means for cleaving precursor polyprotein . Cell 77, 761–771 (1994).
Zarembinski, T.I. et al. Structure-based assignment of the biochemical function of a hypothetical protein: a test case of structural genomics. Proc. Natl. Acad. Sci. USA 95, 15189–15193 (1998).
Brenner, S.E., Barken, D. & Levitt, M. The PRESAGE database for structural genomics. Nucleic Acids Res. 27, 251–253 (1999).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Skolnick, J., Fetrow, J. & Kolinski, A. Structural genomics and its importance for gene function analysis. Nat Biotechnol 18, 283–287 (2000). https://doi.org/10.1038/73723
Received:
Accepted:
Issue date:
DOI: https://doi.org/10.1038/73723
This article is cited by
-
Homology Modeling and Analysis of Vacuolar Aspartyl Protease from a Novel Yeast Expression Host Meyerozyma guilliermondii Strain SO
Arabian Journal for Science and Engineering (2023)
-
CSN: unsupervised approach for inferring biological networks based on the genome alone
BMC Bioinformatics (2020)
-
In silico characterisation, homology modelling and structure-based functional annotation of blunt snout bream (Megalobrama amblycephala) Hsp70 and Hsc70 proteins
Journal of Animal Science and Technology (2015)
-
Terbium promotes adhesion and osteogenic differentiation of mesenchymal stem cells via activation of the Smad-dependent TGF-β/BMP signaling pathway
JBIC Journal of Biological Inorganic Chemistry (2014)
-
A simple recipe for the non-expert bioinformaticist for building experimentally-testable hypotheses for proteins with no known homologs
Journal of Structural and Functional Genomics (2012)