Key Points
-
Electrospray ionization mass spectrometry (ESI-MS) is an emerging technology for studying noncovalent ligand–macromolecular target interactions. Characterization of these interactions could provide new insights into undesirable binding interactions at earlier stages in the drug discovery process, enabling termination of the development of such compounds at an earlier stage and at less overall expense.
-
Mass spectrometry has several potential advantages over other methods used to study noncovalent complexes. Ligands or targets do not have to be labelled for detection of the complex, smaller quantities of material are required for analysis, and this technology is rapid and automatable. All these features make mass spectrometry well suited for high-throughput applications in the drug discovery process.
-
ESI-MS can be used to study ligand interactions in protein, multiprotein, DNA and RNA systems, and can provide information about binding specificity, binding affinity, binding stoichiometry, dissociation constants and gas-phase stability of ligand–target complexes. The key information generated by ESI-MS can contribute to, or drive, primary compound screening activities and structure–activity relationship (SAR) optimization.
-
ESI-MS can be used to measure the gas-phase stabilities of protein–ligand complexes, which can then be compared with solution-phase stabilities. Typically, electrostatic and H-bond contacts of inhibitors are modified during the drug development phase, and so the ability to measure the strength of these interactions by gas-phase stability measurements is useful. In the gas phase, electrostatic and H-bond interactions are favoured over hydrophobic interactions, and data interpretation must take this bias into consideration.
-
Multitarget affinity/specificity screening (MASS) enables the discovery of small-molecule ligands that bind to structured regions of RNA. In a single assay, MASS analysis can determine the chemical composition of ligands that bind to an RNA target, the relative/absolute dissociation constants, and the specificity of binding to one RNA target relative to other RNA targets.
-
MASS can also be implemented in a ligand-based lead discovery strategy in the optimization of SARs. Screening of a panel of small molecules consisting of various chemical motifs against a target using MASS identifies structural motifs that bind the target at different locations. The information generated on motif-binding sites and orientation on the target can then be used to design a single structure with higher affinity for the target.
-
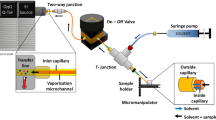
Nanospray ionization uses a smaller-diameter ESI emitter (typically 2–20 μm) and offers several potential advantages over aspects of ESI-MS-based drug discovery. Significantly less sample is consumed in an analysis, with meaningful measurements being made from total sample volumes in the 1–3 μl range. Moreover, nanospray ionization is thought to be a gentler method of ionization than ESI in a conventional format, and in some cases has been shown to provide greater sensitivity than ESI (based on both less sample consumption and more efficient ion desolvation).
Abstract
For many years, analytical mass spectrometry has had numerous supporting roles in the drug development process, including the assessment of compound purity; quantitation of absorption, distribution, metabolism and excretion; and compound-specific pharmacokinetic analyses. More recently, mass spectrometry has emerged as an effective technique for identifying lead compounds on the basis of the characterization of noncovalent ligand–macromolecular target interactions. This approach offers several attractive properties for screening applications in drug discovery compared with other strategies, including the small quantities of target and ligands required, and the capacity to study ligands or targets without having to label them. Here, we review the application of electrospray ionization mass spectrometry to the interrogation of noncovalent complexes, highlighting examples from drug discovery efforts aimed at a range of target classes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lee, M. S. Integrated Strategies for Drug Discovery Using Mass Spectrometry (John Wiley & Sons, Hoboken New Jersey, 2005).
Zhang, J., McCombie, G., Guenat, C. & Knochenmuss, R. FT-ICR mass spectrometry in the drug discovery process. Drug Discov. Today 10, 635–642 (2005).
Siegel, M. M. Early discovery drug screening using mass spectrometry. Curr. Topics Med. Chem. 2, 13–33 (2002).
Hofstadler, S. A. & Griffey, R. H. Analysis of noncovalent complexes of DNA and RNA by mass spectrometry. Chem. Rev. 101, 377–390 (2001).
Hofstadler, S. A. & Griffey, R. H. Mass spectrometry as a drug discovery platform against RNA targets. Curr. Opin. Drug Discov. Dev. 3, 423–431 (2000).
De Stasio, E. A., Moazed, D., Noller, H. F. & Dahlberg, A. E. Mutations in 16S ribosomal RNA disrupt antibiotic--RNA interactions. EMBO J. 8, 1213–1216 (1989).
Bryan, L. E. in New dimensions in Antimicrobial Therapy (eds Root, R. K. & Sande, M. A.) 17–35 (Churchill Livingstone, New York, 1984).
Wong, C.-H., Hendrix, M., Priestley, E. S. & Greenberg, W. A. Specificity of aminoglycoside antibiotics for the A-site of the decoding region of ribosomal RNA. Chem. Biol. 5, 397–406 (1998).
Ganem, B., Li, Y.-T. & Henion, J. D. Detection of Noncovalent Receptor-Ligand Complexes by Mass Spectrometry. J. Am. Chem. Soc. 113, 6294–6296 (1991). Seminal paper demonstrating that specific noncovalent complexes can be studied with ESI-MS.
Smith, R. D., Light-Wahl, K. J., Winger, B. E. & Loo, J. A. Preservation of noncovalent associations in electrospray ionization mass-spectrometry — multiply charged polypeptide and protein dimers. Organic Mass Spectrometry 27, 811–821 (1992).
Loo, J. A. Bioanalytical mass spectrometry: many flavors to choose. Bioconjug. Chem. 6, 644–665 (1995).
Loo, J. A. Studying noncovalent protein complexes by electrospray ionization mass spectrometry. Mass Spectrom. Rev. 16, 1–23 (1997).
Bruce, J. E. et al. Bio-affinity characterization mass spectrometry. Rapid Commun. Mass Spectrom. 9, 644–650 (1995).
van den Heuvel, R. H. & Heck, A. J. Native protein mass spectrometry: from intact oligomers to functional machineries. Curr. Opin. Chem. Biol. 8, 519–526 (2004).
Breuker, K. The study of protein–ligand interactions by mass spectrometry — a personal view. Int. J. Mass Spectrom. 239, 33–41 (2004).
Beck, J. L., Colgrave, M. L., Ralph, S. F. & Sheil, M. M. Electrospray ionization mass spectrometry of oligonucleotide complexes with drugs, metals, and proteins. Mass Spectrom. Rev. 20, 61–87 (2001).
Kempen, E. C. & Brodbelt, J. S. A method for the determination of binding constants by electrospray ionization mass spectrometry. Anal. Chem. 72, 5411–5416 (2000).
Griffey, R. H. et al. Characterization of low-affinity complexes between rna and small molecules using electrospray ionization mass spectrometry. J. Am. Chem. Soc. 122, 9933–9938 (2000).
Griffey, R. H., Hofstadler, S. A., Sannes-Lowery, K. A., Ecker, D. J. & Crooke, S. T. Determinants of aminoglycoside-binding specificity for rRNA by using mass spectrometry. Proc. Natl Acad. Sci. USA 96, 10129–10133 (1999). Demonstrates that detailed information, including approximate binding sites, can be gained from studying noncovalent complexes with mass spectrometry.
Sannes-Lowery, K. A., Griffey, R. H. & Hofstadler, S. A. Measuring dissociation constants of RNA and aminoglycoside antibiotics by electrospray ionization mass spectrometry. Anal. Biochem. 280, 264–271 (2000).
Hensley, P. Defining the structure and stability of macromolecular assemblies in solution: the re-emergence of analytical ultracentrifugation as a practical tool. Structure 4, 367–373 (1996).
Smith, D. L. & Zhang, Z. Q. Probing noncovalent structural features of proteins by mass spectrometry. Mass Spectrom. Rev. 13, 411–429 (1994).
Lakey, J. H. & Raggett, E. M. Measuring protein–protein interactions. Curr. Opin. Struct. Biol. 8, 119 (1998).
Lim, H. K., Hsieh, Y. L., Ganem, B. & Henion, J. Recognition of cell-wall peptide ligands by vancomycin group antibiotics: studies using ion spray mass spectrometry. J. Mass Spectrom. 30, 708–714 (1995). One of the first papers to study antibiotic interactions with peptides and provide insights about the mechanism of action of the antibiotic.
Jorgensen, T. J. D. & Roepstorff, P. Direct determination of solution binding constants for noncovalent complexes between bacterial cell wall peptide analogues and vancomycin group antibiotics by electrospray ionization mass spectrometry. Anal. Chem 70, 4427–4432 (1998).
Jorgensen, T. J. D., Delforge, D., Remacle, J., Bojesen, G. & Roepstorff, P. Collision-induced dissociation of noncovalent complexes between vancomycin antibiotics and peptide ligand stereoisomers: evidence for molecular recognition in the gas phase. Int. J. Mass Spectrom. 188, 63–85 (1999).
Jorgensen, T. J. D., Hvelplund, P., Andersen, J. U. & Roepstorff, P. Tandem mass spectrometry of specific vs. nonspecific noncovalent complexes of vancomycin antibiotics and peptide ligands. Int. J. Mass Spectrom. 219, 659–670 (2002).
Jorgensen, T. J. D., Staroske, T., Roepstorff, P., Williams, D. H. & Heck, A. J. R. Subtle differences in molecular recognition between modified gylcopeptide antibiotics and bacterial receptor peptides identified by electrospray ionization mass spectrometry. J. Chem. Soc., Perkin Trans. 2, 1859–1863 (1999).
van de Kerk-van Hoof, A. & Heck, A. J. Interactions of a- and b-avoparcin with bacterial cell-wall receptor-mimicking peptides studied by electrospray ionization mass spectrometry. J. Antimicrob. Chemother. 44, 593–599 (1999).
van de Kerk-van Hoof, A. & Heck, A. J. R. Covalent and non-covalent dissociations of gas-phase complexes of avoparcin and bacterial receptor mimicking precursor peptides studied by collisionally activated decomposition mass spectrometry. J. Mass Spectrom. 34, 813–819 (1999).
Coulson, C. J. Molecular Mechanisms fo Drug Action (Taylor & Francis, London, 1994).
Robinson, C. V. et al. Probing the nature of noncovalent interactions by mass spectrometry. A study of protein-CoA ligand binding and assembly. J. Am. Chem. Soc. 118, 8646–8653 (1996).
Wu, Q. Y. et al. Carbonic anhydrase-inhibitor binding: from solution to the gas phase. J. Am. Chem. Soc. 119, 1157–1158 (1997).
Rogniaux, H. et al. Binding of aldose reductase inhibitors: correlation of crystallographic and mass spectrometric studies. J. Am. Soc. Mass Spectrom. 10, 635–647 (1999).
Nesatyy, V. J. Gas-phase binding of non-covalent protein complexes between bovine pancreatic trypsin inhibitor and its target enzymes studied by electrospray ionization tandem mass spectrometry. J. Mass Spectrom. 36, 950–959 (2001).
Daniel, J. M., Friess, S. D., Rajagopalan, S., Wendt, S. & Zenobi, R. Quantitative determination of noncovalent binding interactions using soft ionization mass spec-trometry. Int. J. Mass Spectrom. 216, 1–27 (2002).
Cheng, X. et al. Using electrospray ionization FTICR mass spectrometry to study competitive binding of inhibitors to carbonic anhydrase. J. Am. Chem. Soc. 117, 8859–8860 (1995). Demonstrates how high-resolution mass spectrometry can be used to simultaneously screen multiple compounds against a target.
Gao, J. et al. Screening derivatized peptide libraries for tight binding inhibitors to carbonic anhydrase II by electrospray ionization-mass spectrometry. J. Med. Chem. 39, 1949–1955 (1996).
Gao, J. et al. Probing the energetics of dissociation of carbonic anhydrase-ligand complexes in the gas phase. Biophys. J. 76, 3253–3260 (1999).
Wigger, M., Eyler, J. R., Benner, S. A., Li, W. & Marshall, A. G. Fourier transform-ion cyclotron resonance mass spectrometric resolution, identification, and screening of non-covalent complexes of Hck Src homology 2 domain receptor and ligands from a 324-member peptide combinatorial library. J. Am. Soc. Mass Spectrom. 13, 1162–1169 (2002).
Grucza, R. A., Bradshaw, J. M., Futterer, K. & Waksman, G. SH2 domains: from structure to energetics, a dual approach to the study of structure-function relationships. Med. Res. Rev. 19, 273–293 (1999).
Koch, C. A., Anderson, D., Moran, M. F., Ellis, C. & Pawson, T. SH2 and SH3 domains: elements that control interactions of cytoplasmic signaling proteins. Science 252, 668–674 (1991).
Courtneidge, S. A. Protein tyrosine kinases, with emphasis on the Src family. Semin. Cancer Biol. 5, 239–246 (1994).
Courtneidge, S. A. Role of Src in signal transduction pathways. The Jubilee Lecture. Biochem. Soc. Trans. 30, 11–17 (2002).
Guan, S. H. & Marshall, A. G. Stored waveform inverse Fourier transform (SWIFT) ion excitation in trapped-ion mass spectometry: theory and applications. Int. J. Mass Spectrom. Ion Process. 158, 5–37 (1996).
Little, D. P., Speir, J. P., Senko, M. W., O'Connor, P. B. & McLafferty, F. W. Infrared multiphoton dissociation of large multiply charged ions for biomolecule sequencing. Anal. Chem. 66, 2809–2815 (1994).
Dufresne, C. P., Wood, T. D. & Hendrickson, C. L. High- resolution electrospray ionization Fourier transform mass spectrometry with infrared multiphoton disso-ciation of glucokinase from Bacillus stearothermophilus. J. Am. Soc. Mass Spectrom. 9, 1222–1225 (1998).
Loo, J. A., Hu, P., McConnell, P. & Mueller, W. T. A study of Src SH2 domain protein-phosphopeptide binding interactions by electrospray ionization mass spectrometry. J. Am. Soc. Mass Spectrom. 8, 234–243 (1997).
Chung, E. et al. Mass spectrometric and thermodynamic studies reveal the role of water molecules in complexes formed between SH2 domains and tyrosyl phosphopeptides. Structure 6, 1141–1151 (1998).
Chung, E. W. et al. Probing the nature of interactions in SH2 binding interfaces--evidence from electrospray ionization mass spectrometry. Protein Sci. 8, 1962–1970 (1999).
Bligh, S. W., Haley, T. & Lowe, P. N. Measurement of dissociation constants of inhibitors binding to Src SH2 domain protein by non-covalent electrospray ionization mass spectrometry. J. Mol. Recognit. 16, 139–148 (2003).
Larson, E. R., Lipinski, C. A. & Sarges, R. Medicinal chemistry of aldose reductase inhibitors. Med. Res. Rev. 8, 159–186 (1988).
Nesatyy, V. J. Mass spectrometry evaluation of the solution and gas-phase binding properties of noncovalent protein complexes. Int. J. Mass Spectrom. 221, 147–161 (2002).
McCammon, M. G. et al. Screening transthyretin amyloid fibril inhibitors: characterization of novel multiprotein, multiligand complexes by mass spectrometry. Structure 10, 851–863 (2002). Demonstrates the potential to examine binding of ligands to large complex multi-protein systems.
McCammon, M. G., Hernandez, H., Sobott, F. & Robinson, C. V. Tandem mass spectrometry defines the stoichiometry and quaternary structural arrangement of tryptophan molecules in the multiprotein complex TRAP. J. Am. Chem. Soc. 126, 5950–5951 (2004).
Pepys, M. B. et al. Targeting C-reactive protein for the treatment of cardiovascular disease. Nature 440, 1217–1221 (2006).
Kelly, J. W. Amyloid fibril formation and protein misassembly: a structural quest for insights into amyloid and prion diseases. Structure 5, 595–600 (1997).
Koo, E. H., Lansbury, P. T., Jr. & Kelly, J. W. Amyloid diseases: abnormal protein aggregation in neurodegeneration. Proc. Natl Acad. Sci. USA 96, 9989–9990 (1999).
Petrassi, H. M., Klabunde, T., Sacchettini, J. & Kelly, J. W. Structure-based design of N-phenyl phenoxazine transthyretin amyloid fibril inhibitors. J. Am. Chem. Soc. 122, 2178–2192 (2000).
Benkestock, K., Edlund, P. O. & Roeraade, J. Electrospray ionization mass spectrometry as a tool for determination of drug binding sites to human serum albumin by noncovalent interaction. Rapid Commun. Mass Spectrom. 19, 1637–1643 (2005).
Gabelica, V., De Pauw, E. & Rosu, F. Interaction between antitumor drugs and a double-stranded oligonucleotide studied by electrospray ionization mass spectrometry. J. Mass Spectrom. 34, 1328–1337 (1999).
Rosu, F., Gabelica, V., Houssier, C. & De Pauw, E. Determination of affinity, stoichiometry and sequence selectivity of minor groove binder complexes with double-stranded oligodeoxynucleotides by electrospray ionization mass spectrometry. Nucleic Acids Res. 30, e82 (2002). Demonstrates how mass spectrometry can be used to assess the preference of drugs for binding to sequences that have only minor differences.
Kapur, A., Beck, J. L. & Sheil, M. M. Observation of daunomycin and nogalamycin complexes with duplex DNA using electrospray ionization mass spectrometry. Rapid Commun. Mass Spectrom. 13, 2489–2497 (1999).
Gupta, R., Kapur, A., Beck, J. L. & Sheil, M. M. Positive ion electrospray ionization mass spectrometry of double-stranded DNA/drug complexes. Rapid Communications in Mass Spectrometry 15, 2472–2480 (2001).
Gupta, R., Beck, J. L., Ralph, S. F., Sheil, M. M. & Aldrich-Wright, J. R. Comparison of the binding stoichiometries of positively charged DNA-binding drugs using positive and negative ion electrospray ionization mass spectrometry. J. Am. Soc. Mass Spectrom. 15, 1382–1391 (2004).
Wan, K. X., Gross, M. L. & Shibue, T. Gas-phase stability of double-stranded oligodeoxynucleotides and their noncovalent complexes with DNA-binding drugs as revealed by collisional activation in an ion trap. J. Am. Soc. Mass Spectrom. 11, 450–457 (2000).
Wan, K. X., Shibue, T. & Gross, M. L. Non-covalent complexes between DNA-binding drugs and double-stranded oligodeoxynucleotides: a study by ESI ion-trap mass spectrometry. J. Am. Chem. Soc. 122, 300–307 (2000).
Oehlers, L., Mazzitelli, C. L., Brodbelt, J. S., Rodriguez, M. & Kerwin, S. Evaluation of complexes of DNA duplexes and novel benzoxazoles or benzimidazoles by electrospray ionization mass spectrometry. J. Am. Soc. Mass Spectrom. 15, 1593–1603 (2004).
Triolo, A., Arcamone, F. M., Raffaelli, A. & Salvadori, P. Non-covalent complexes between DNA-binding drugs and double-stranded deoxyoligonucleotides: a study by ionspray mass spectrometry. J. Mass Spectrom. 32, 1186–1194 (1997).
Greig, M. J. & Robinson, J. M. Detection of oligonucleotide-ligand complexes by ESI-MS (DOLCE-MS) as a component of high throughput screening. J. Biomol. Screening 5, 441–454 (2000). Describes a mass spectrometry-based method for screening ligand binding to single-stranded or double-stranded DNA.
Hermann, T. & Patel, D. J. Adaptive recognition by nucleic acid aptamers. Science 287, 820–825 (2000).
Ramos, A., Gubser, C. C. & Varani, G. Recent solution structures of RNA and its complexes with drugs, peptides and proteins. Curr. Opin. Struct. Biol. 7, 317–323 (1997).
Conn, G. L. & Draper, D. E. RNA structure. Curr. Opin. Struct. Biol. 8, 278–285 (1998).
Hofstadler, S. A. et al. Multiplexed screening of neutral mass-tagged rna targets against ligand libraries with electrospray ionization FTICR MS: a paradigm for high-throughput affinity screening. Anal. Chem. 71, 3436–3440 (1999). Describes the simultaneous screening of multiple macromolecular targets against multiple ligands.
Sannes-Lowery, K. A., Drader, J. J., Griffey, R. H. & Hofstadler, S. A. Fourier transform ion cyclotron resonance mass spectrometry as a high throughput affinity screen to identify RNA-binding ligands. Trends Anal. Chem. 19, 481–491 (2000). Reports a highly automated ESI-MS-based high-throughput screening approach to find ligands that bind to structured RNA targets.
Wang, X., Migawa, M. T., Sannes-Lowery, K. A. & Swayze, E. E. The synthesis and 16S A-site rRNA recognition of carbohydrate-free aminoglycosides. Bioorg. Med. Chem. Lett. 15, 4919–4922 (2005).
He, Y. et al. Synthesis and evaluation of novel bacterial rRNA-binding benzimidazoles by mass spectrometry. Bioorg. Med. Chem. Lett. 14, 695–699 (2004).
Seth, P. P. et al. SAR by MS: discovery of a new class of RNA-binding small molecules for the hepatitis C virus: internal ribosome entry site IIA subdomain. J. Med. Chem. 48, 7099–7102 (2005).
Swayze, E. E. et al. SAR by MS: A ligand based technique for drug lead discovery against structured RNA targets. J. Med. Chem. 45, 3816–3819 (2002). Describes the use of the SAR-by-MS technique.
Recht, M. I., Fourmy, D., Blanchard, S. C., Dahlquist, K. D. & Puglisis, J. D. RNA sequence determinants for aminoglycoside binding to an A-site rRNA model oligonucleotide. J. Mol. Biol. 262, 421–436 (1996).
Cummins, L. L. et al. Multitarget affinity/specificity screening of natural products: finding and characterizing high affinity ligands from complex mixtures by using high performance mass spectrometry. J. Nat. Prod. 66, 1186–1190 (2003).
Ockey, D. A. et al. Structure–activity relationships by mass spectrometry: identification of novel MMP-3 inhibitors. Bioorg. Med. Chem. 12, 37–44 (2004).
Gabelica, V. et al. Advantages and drawbacks of nanospray for studying noncovalent protein-DNA complexes by mass spectrometry. Rapid Commun. Mass Spectrom. 16, 1723–1728 (2002).
Rostom, A. A. & Robinson, C. V. Disassembly of intact multiprotein complexes in the gas phase. Curr. Opin. Struct. Biol. 9, 135–141 (1999).
Benkestock, K., Sundqvist, G., Edlund, P. O. & Roeraade, J. Influence of droplet size, capillary-cone distance and selected instrumental parameters for the analysis of noncovalent protein-ligand complexes by nano-electrospray ionization mass spectrometry. J. Mass Spectrom. 39, 1059–1067 (2004).
Keetch, C. A. et al. Use of a microchip device coupled with mass spectrometry for ligand screening of a multi-protein target. Anal. Chem. 75, 4937–4941 (2003).
Zhang, S., Van Pelt, C. K. & Wilson, W. D. Quantitative determination of noncovalent binding interactions using automated nanoelectrospray mass spectrometry. Anal. Chem. 75, 3010–3018 (2003). Describes the use of an automated microfabricated nanoelectrospray platform to characterize noncovalent complexes.
Benkestock, K. et al. Automated nano-electrospray mass spectrometry for protein-ligand screening by noncovalent interaction applied to human H-FABP and A-FABP. J. Biomol. Screen. 8, 247–256 (2003).
Sun, Y. & Cheng, J. Hydrolysis of lignocllulosic materials for ethanol production:review. Bioresour. Technol. 83, 1–11 (2002).
Gidden, J. et al. Application of ion mobility to the gas-phase conformational analysis of polyhedral oligomeric silsesquioxanes (POSS). Int. J. Mass Spectrom. 222, 63–73 (2003).
Koeniger, S. L., Merenbloom, S. I. & Clemmer, D. E. Evidence for many resolvable structures within conformation types of electrosprayed ubiquitin ions. J. Phys. Chem. B 110, 7017–7021 (2006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Ion mobility spectrometry
-
The separation of ions according to their velocity through a buffer gas under the influence of an electric field.
- Collisionally activated dissociation
-
(CAD). Dissociation is caused by collisions between ions and gaseous molecules that result in the conversion of translational energy into internal vibrational energy of the ion.
- SH2 domain
-
(Src homology 2 domain). A protein motif that recognizes and binds tyrosine-phosphorylated sequences, and therefore has a key role in relaying cascades of signal transduction.
Rights and permissions
About this article
Cite this article
Hofstadler, S., Sannes-Lowery, K. Applications of ESI-MS in drug discovery: interrogation of noncovalent complexes. Nat Rev Drug Discov 5, 585–595 (2006). https://doi.org/10.1038/nrd2083
Issue date:
DOI: https://doi.org/10.1038/nrd2083
This article is cited by
-
The Advance of Plasmonic-Electric Nanopipette Sensing in Single Cells
Current Pharmacology Reports (2021)
-
Screening for natural inhibitors of human topoisomerases from medicinal plants with bio-affinity ultrafiltration and LC–MS
Phytochemistry Reviews (2020)
-
Native mass spectrometry of human carbonic anhydrase I and its inhibitor complexes
JBIC Journal of Biological Inorganic Chemistry (2020)
-
An In Vitro Study of Aromatic Stacking of Drug Molecules
Journal of the American Society for Mass Spectrometry (2019)
-
Nanoparticle-based surface assisted laser desorption ionization mass spectrometry: a review
Microchimica Acta (2019)