Abstract
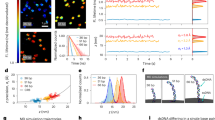
Bacteria can acquire genetic diversity, including antibiotic resistance and virulence traits, by horizontal gene transfer. In particular, many bacteria are naturally competent for uptake of naked DNA from the environment in a process called transformation. Here, we used optical tweezers to demonstrate that the DNA transport machinery in Bacillus subtilis is a force-generating motor. Single DNA molecules were processively transported in a linear fashion without observable pausing events. Uncouplers inhibited DNA uptake immediately, suggesting that the transmembrane proton motive force is needed for DNA translocation. We found an uptake rate of 80 ± 10 bp s−1 that was force-independent at external forces <40 pN, indicating that a powerful molecular machine supports DNA transport.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ochman, H., Lawrence, J.G. & Groisman, E.A. Lateral gene transfer and the nature of bacterial innovation. Nature 405, 299–304 (2000).
Dubnau, D. DNA uptake in bacteria. Annu. Rev. Microbiol. 53, 217–244 (1999).
Chen, I. & Dubnau, D. DNA uptake during bacterial transformation. Nat. Rev. Microbiol. 3, 241–249 (2004).
Smith, D.E. et al. The bacteriophage straight phi29 portal motor can package DNA against a large internal force. Nature 413, 748–752 (2001).
Salman, H., Zbaida, D., Rabin, Y., Chatenay, D. & Elbaum, M. Kinetics and mechanism of DNA uptake into the cell nucleus. Proc. Natl. Acad. Sci. USA 98, 7247–7252 (2001).
Sauer-Budge, A.F., Nyamwanda, J.A., Lubensky, D.K. & Branton, D. Unzipping kinetics of double-stranded DNA in a nanopore. Phys. Rev. Lett. 90, 238101 (2003).
Li, J. et al. Ion-beam sculpting at nanometre length scales. Nature 412, 166–169 (2001).
Hoa, T.T., Tortosa, P., Albano, M. & Dubnau, D. Rok (YkuW) regulates genetic competence in Bacillus subtilis by directly repressing comK. Mol. Microbiol. 43, 15–26 (2002).
Provvedi, R., Chen, I. & Dubnau, D. NucA is required for DNA cleavage during transformation of Bacillus subtilis. Mol. Microbiol. 40, 634–644 (2001).
Provvedi, R. & Dubnau, D. ComEA is a DNA receptor for transformation of competent Bacillus subtilis. Mol. Microbiol. 31, 271–280 (1999).
Bouchiat, C. et al. Estimating the persistence length of a worm-like chain molecule from force-extension measurements. Biophys. J. 76, 409–413 (1999).
Nieuwenhoven, M.H., Hellingwerf, K.J., Venema, G. & Konings, W.N. Role of proton motive force in genetic transformation of Bacillus subtilis. J. Bacteriol. 151, 771–776 (1982).
Mejean, V. & Claverys, J.P. DNA processing during entry in transformation of Streptococcus pneumoniae. J. Biol. Chem. 268, 5594–5599 (1993).
Neupert, W. & Brunner, M. The protein import motor of mitochondria. Nat Rev Mol. Cell. Biol. 3, 555–565 (2002).
Chauwin, J.F., Oster, G. & Glick, B.S. Strong precursor-pore interactions constrain models for mitochondrial protein import. Biophys. J. 74, 1732–1743 (1998).
Oster, G. & Wang, H. Rotary protein motors. Trends Cell Biol. 13, 114–121 (2003).
Londono-Vallejo, J.A. & Dubnau, D. Mutation of the putative nucleotide binding site of the Bacillus subtilis membrane protein ComFA abolishes the uptake of DNA during transformation. J. Bacteriol. 176, 4642–4645 (1994).
Perkins, T.T., Dalal, R.V., Mitsis, P.G. & Block, S.M. Sequence-dependent pausing of single lambda exonuclease molecules. Science 301, 1914–1918 (2003).
Davenport, R.J., Wuite, G.J., Landick, R. & Bustamante, C. Single-molecule study of transcriptional pausing and arrest by E. coli RNA polymerase. Science 287, 2497–2500 (2000).
Maier, B., Bensimon, D. & Croquette, V. Replication by a single DNA polymerase of a stretched single-stranded DNA. Proc. Natl. Acad. Sci. USA 97, 12002–12007 (2000).
Maier, B. et al. Single pilus motor forces exceed 100 pN. Proc. Natl. Acad. Sci. USA 99, 16012–16017 (2002).
Kaiser, D. Bacterial motility: how do pili pull? Curr. Biol. 10, R777–R780 (2000).
Albano, M., Breitling, R. & Dubnau, D.A. Nucleotide sequence and genetic organization of the Bacillus subtilis comG operon. J. Bacteriol. 171, 5386–5404 (1989).
Liu, L., Nakano, M.M., Lee, O.H. & Zuber, P. Plasmid-amplified comS enhances genetic competence and suppresses sinR in Bacillus subtilis. J. Bacteriol. 178, 5144–5152 (1996).
Anagnostopoulos, C. & Spizizen, J. Requirements for transformation in Bacillus subtilis. J. Bacteriol. 81, 3389–4302 (1961).
Albano, M., Hahn, J. & Dubnau, D. Expression of competence genes in Bacillus subtilis. J. Bacteriol. 169, 3110–3117 (1987).
Fung, D.C. & Berg, H.C. Powering the flagellar motor of Escherichia coli with an external voltage source. Nature 375, 809–812 (1995).
Simmons, R.M., Finer, J.T., Chu, S. & Spudich, J.A. Quantitative measurements of force and displacement using an optical trap. Biophys. J. 70, 1813–1822 (1996).
Marko, J.F. & Siggia, E.D. Stretching DNA. Macromolecules 28, 8759–8770 (1995).
Acknowledgements
We thank J. Hahn for strain BD3773 and invaluable discussions. The project was supported by grant GM3677 and D.D. by grant GM43756. B.M. acknowledges support by the Deutsche Forschungsgemeinschaft (Emmy Noether Programm).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Example for the distance between the bead in the optical trap and the bacterium attached to the glass coverslide during DNA uptake. (PDF 372 kb)
Supplementary Fig. 2
Elasticity of DNA in competence medium. (PDF 84 kb)
Rights and permissions
About this article
Cite this article
Maier, B., Chen, I., Dubnau, D. et al. DNA transport into Bacillus subtilis requires proton motive force to generate large molecular forces. Nat Struct Mol Biol 11, 643–649 (2004). https://doi.org/10.1038/nsmb783
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/nsmb783
This article is cited by
-
Natural transformation occurs independently of the essential actin-like MreB cytoskeleton in Legionella pneumophila
Scientific Reports (2015)
-
What renders Bacilli genetically competent? A gaze beyond the model organism
Applied Microbiology and Biotechnology (2015)
-
Three Powerful Research Tools from Single Cells into Single Molecules: AFM, Laser Tweezers, and Raman Spectroscopy
Applied Biochemistry and Biotechnology (2011)
-
Force–Velocity Curves of Motor Proteins Cooperating In Vivo
Cell Biochemistry and Biophysics (2008)