Abstract
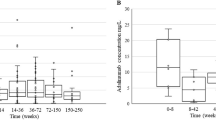
Given the substantial degree of inter-individual variability in treatment responses in patients with inflammatory bowel disease (IBD), treatment optimization is warranted. We provide an overview of pharmacogenetic variants that can affect the efficacy or toxicity of drugs commonly used for IBD. We used The Danish Register of Medical Product Statistics for information about medical treatment from 2000 – 2024. Of the most used drugs Azathioprine, Mesalazine, and Sulfasalazine have pharmacogenetic recommendation guidelines and/or FDA annotations. Approximately 19,241 Danish individuals treated with azathioprine—around 801 annually—may carry a genetic variant for which pharmacogenetic dosing guidelines exist. Up to 13,934 Danish individuals using infliximab or adalimumab (~580 individuals each year) have a potential risk of developing immunogenicity related to monoclonal antibodies. This study shows the possibilities of pharmacogenetic testing as a supportive clinical decision tool to optimize treatment and minimize risk of e.g., serious adverse effects, remission, and other serious complications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Roden DM, McLeod HL, Relling MV, Williams MS, Mensah GA, Peterson JF, et al. Pharmacogenomics. Lancet. 2019;394:521–32.
Westergaard N, Nielsen RS, Jørgensen S, Vermehren C. Drug use in Denmark for drugs having pharmacogenomics (PGx) based dosing guidelines from CPIC or DPWG for CYP2D6 and CYP2C19 drug–gene pairs: Perspectives for introducing PGx test to polypharmacy patients. J Pers Med. 2020;10:3.
Vermehren C, Nielsen RS, Jørgensen S, Drastrup AM, Westergaard N. Drug use among nursing home residents in denmark for drugs having pharmacogenomics based (PGX) dosing guidelines: potential for preemptive PGX testing. J Pers Med. 2020;10:1–11.
Relling MV, Klein TE. CPIC: Clinical pharmacogenetics implementation consortium of the pharmacogenomics research network. Clin Pharmacol Ther. 2011;89:464–7.
Bank PCD, Caudle KE, Swen JJ, Gammal RS, Whirl-Carrillo M, Klein TE, et al. Comparison of the guidelines of the clinical pharmacogenetics implementation consortium and the dutch pharmacogenetics working group. Clin Pharmacol Ther. 2018;103:599–618.
Barbarino JM, Whirl-Carrillo M, Altman RB, Klein TE. PharmGKB: a worldwide resource for pharmacogenomic information. Wiley Interdiscip Rev Syst Biol Med. 2018;10:e1417.
European Medicines Agency. EMA recommendations on DPD testing prior to treatment with fluorouracil, “ EMA recommendations on DPD testing prior to treatment with fluorouracil, capecitabine, tegafur and flucytosine,.” 2020. https://www.fagg.be/sites/default/files/content/dhpc_fluorouracil_nl_-_.
Westergaard N, Tarnow L, Vermehren C. Use of clopidogrel and proton pump inhibitors alone or in combinations in persons with diabetes in Denmark; potential for CYP2C19 genotype-guided drug therapy. Metabolites. 2021;11:1–11.
Lucafò M, Franca R, Selvestrel D, Curci D, Pugnetti L, Decorti G, et al. Pharmacogenetics of treatments for inflammatory bowel disease. Expert Opin Drug Metab Toxicol. 2018;14:1209–23.
Van Den Bosch BJC, Coenen MJH. Pharmacogenetics of inflammatory bowel disease. Pharmacogenomics. 2021;22:55–66.
Frolkis AD, Dykeman J, Negrón ME, Debruyn J, Jette N, Fiest KM, et al. Risk of surgery for inflammatory bowel diseases has decreased over time: a systematic review and meta-analysis of population-based studies. Gastroenterology. 2013;145:996–1006.
Yamamoto-Furusho JK. Pharmacogenetics in inflammatory bowel disease: understanding treatment response and personalizing therapeutic strategies. Pharmacogenomics Personalized Med. 2017;10:197–204.
Voskuil MD, Bangma A, Weersma RK, Festen EAM. Predicting (side) effects for patients with inflammatory bowel disease: the promise of pharmacogenetics. World J Gastroenterol. 2019;25:2539–48.
Weinshilboum RM, Wang L. Pharmacogenomics: precision medicine and drug response. Mayo Clin Proc. 2017;92:1711–22.
Luzum JA, Petry N, Taylor AK, Van Driest SL, Dunnenberger HM, Cavallari LH. Moving pharmacogenetics into practice: it’s all about the evidence! Clin Pharmacol Ther. 2021;110:649–61.
Relling MV, Klein TE, Gammal RS, Whirl-Carrillo M, Hoffman JM, Caudle KE. The clinical pharmacogenetics implementation consortium: 10 years later. Clin Pharmacol Ther. 2020;107:171–5.
Gaedigk A, Ingelman-Sundberg M, Miller NA, Leeder JS, Whirl-Carrillo M, Klein TE. The pharmacogene variation (PharmVar) consortium: incorporation of the human cytochrome P450 (CYP) allele nomenclature database. Clin Pharmacol Ther. 2018;103:399–401.
Statistics. “The Danish Register of Medical Product Statistics,.” 2025. www.medstat.dk.
McInnes G, Lavertu A, Sangkuhl K, Klein TE, Whirl-Carrillo M, Altman RB. Pharmacogenetics at scale: an analysis of the UK biobank. Clin Pharmacol Ther. 2021;109:1528–37.
Bangma A, Voskuil MD, Uniken Venema WTC, Brugge H, Hu S, Lanting P, et al. Predicted efficacy of a pharmacogenetic passport for inflammatory bowel disease. Aliment Pharmacol Ther. 2020;51:1105–15.
Sazonovs A, Kennedy NA, Moutsianas L, Heap GA, Rice DL, Reppell M, et al. HLA-DQA1*05 carriage associated with development of anti-drug antibodies to infliximab and adalimumab in patients with crohn’s disease. Gastroenterology. 2020;158:189–99.
Chanchlani N, Lin S, Bewshea C, Hamilton B, Thomas A, Smith R, et al. Mechanisms and management of loss of response to anti-TNF therapy for patients with Crohn’s disease: 3-year data from the prospective, multicentre PANTS cohort study. Lancet Gastroenterol Hepatol. 2024;9:521–38.
Aterido A, Palau N, Domènech E, Nos Mateu P, Gutiérrez A, Gomollón F, et al. Genetic association between CD96 locus and immunogenicity to anti-TNF therapy in Crohn’s disease. Pharmacogenomics J. 2019;19:547–55.
Oray M, Abu Samra K, Ebrahimiadib N, Meese H, Foster CS. Long-term side effects of glucocorticoids. Expert Opin Drug Saf. 2016;15:457–65.
Skrzypczak-Zielinska M, Gabryel M, Marszalek D, Dobrowolska A, Slomski R. NGS study of glucocorticoid response genes in inflammatory bowel disease patients. Arch Med Sci. 2021;17:417–33.
Chen HL, Li LR. Glucocorticoid receptor gene polymorphisms and glucocorticoid resistance in inflammatory bowel disease: a meta-analysis. Digestive Dis Sci. 2012;57:3065–75.
Gammal RS, Pirmohamed M, Somogyi AA, Morris SA, Formea CM, Elchynski AL, et al. Expanded clinical pharmacogenetics implementation consortium (CPIC) guideline for medication use in the context of G6PD genotype. www.cpicpgx.org.
Yee J, Kim SM, Han JM, Lee N, Yoon HY, Gwak HS The association between NAT2 acetylator status and adverse drug reactions of sulfasalazine: a systematic review and meta-analysis. Sci Rep. 2020;10.
Hou ZD, Xiao ZY, Gong Y, Zhang YP, Zeng QY. Arylamine N-acetyltransferase polymorphisms in Han Chinese patients with ankylosing spondylitis and their correlation to the adverse drug reactions to sulfasalazine. BMC Pharmacol Toxicol. 2014;15:64.
Heap GA, So K, Weedon M, Edney N, Bewshea C, Singh A, et al. Clinical features and HLA association of 5-aminosalicylate (5-ASA)-induced nephrotoxicity in inflammatory bowel disease. J Crohns Colitis. 2016;10:149–58.
Riley SA, Mani V, Goodman MJ, Herd ME, Dutt S, Turnberg LA. Comparison of delayed release 5 aminosalicylic acid (mesalazine) and sulphasalazine in the treatment of mild to moderate ulcerative colitis relapse. Gut. 1988;29:669–74.
Masuda H, Takahashi Y, Nishida Y, Asai S. Comparison of the effect of mesalazine and sulfasalazine on laboratory parameters: A retrospective observational study. Eur J Clin Pharmacol. 2012;68:1549–55.
Yasutomi E, Hiraoka S, Yamamoto S, Oka S, Hirai M, Yamasaki Y, et al. Switching between three types of mesalazine formulation and sulfasalazine in patients with active ulcerative colitis who have already received high-dose treatment with these agents. J Clin Med. 2019;8:2109.
Raine T, Bonovas S, Burisch J, Kucharzik T, Adamina M, Annese V, et al. ECCO guidelines on therapeutics in ulcerative colitis: medical treatment. J Crohns Colitis. 2022;16:2–17.
Deben DS, Wong DR, van Bodegraven AA. Current status and future perspectives on the use of therapeutic drug monitoring of thiopurine metabolites in patients with inflammatory bowel disease. Expert Opin Drug Metab Toxicol. 2021;17:1433–44.
Relling MV, Schwab M, Whirl-Carrillo M, Suarez-Kurtz G, Pui CH, Stein CM, et al. Clinical pharmacogenetics implementation consortium guideline for thiopurine dosing based on TPMT and NUDT15 Genotypes: 2018 Update. Clin Pharmacol Ther. 2019;105:1095–105.
Feuerstein JD, Nguyen GC, Kupfer SS, Falck-Ytter Y, Singh S, Gerson L, et al. American gastroenterological association institute guideline on therapeutic drug monitoring in inflammatory bowel disease. Gastroenterology. 2017;153:827–34.
Moyer AM. NUDT15: A bench to bedside success story. Clin Biochem. 2021;92:1–8.
Coenen MJH, De Jong DJ, Van Marrewijk CJ, Derijks LJJ, Vermeulen SH, Wong DR, et al. Identification of Patients With Variants in TPMT and dose reduction reduces hematologic events during thiopurine treatment of inflammatory bowel disease. Gastroenterology. 2015;149:907–917.e7.
Šmid A, Urbančič D, Žlajpah JV, Stollarova N, Prelog T, Kavčič M, et al. Genetic profiling of NUDT15 in the Slovenian population. Pharmacogenomics. 2024;25:515–25.
Campbell JM, Bateman E, Stephenson MD, Bowen JM, Keefe DM, Peters MDJ. Methotrexate-induced toxicity pharmacogenetics: an umbrella review of systematic reviews and meta-analyses. Cancer Chemother Pharmacol. 2016;78:27–39.
Nielsen OH, Steenholdt C, Juhl CB, Rogler G. Efficacy and safety of methotrexate in the management of inflammatory bowel disease: a systematic review and meta-analysis of randomized, controlled trials. EClinicalMedicine. 2020;20:100271.
Wang Y, Li Y, Liu Y, Zhang Y, Ke Z, Zhang Y, et al. Patients With IBD receiving methotrexate are at higher risk of liver injury compared with patients with Non-IBD Diseases: a meta-analysis and systematic review. Front Med. 2021;8:774824.
Giletti A, Esperon P. Genetic markers in methotrexate treatments. Pharmacogenomics J. 2018;18:689–703.
Cao M, Guo M, Wu DQ, Meng L. Pharmacogenomics of methotrexate: current status and future outlook. Curr Drug Metab. 2018;19:1182–7.
Treviño LR, Shimasaki N, Yang W, Panetta JC, Cheng C, Pei D, et al. Germline genetic variation in an organic anion transporter polypeptide associated with methotrexate pharmacokinetics and clinical effects. J Clin Oncol. 2009;27:5972–8.
Ramsey LB, Panetta JC, Smith C, Yang W, Fan Y, Winick NJ, et al. Genome-wide study of methotrexate clearance replicates SLCO1B1. Blood. 2013;121:898–904.
Mehta RS, Taylor ZL, Martin LJ, Rosen MJ, Ramsey LB. SLCO1B1 *15 allele is associated with methotrexate-induced nausea in pediatric patients with inflammatory bowel disease. Clin Transl Sci. 2022;15:63–9.
Han JM, Choi KH, Lee HH, Gwak HS. Association between SLCO1B1 polymorphism and methotrexate-induced hepatotoxicity: a systematic review and meta-analysis. Anticancer Drugs. 2022;33:75–9.
Steenholdt C, Bendtzen K, Brynskov J, Ainsworth MA. Optimizing Treatment with TNF Inhibitors in inflammatory bowel disease by monitoring drug levels and antidrug antibodies. Inflamm Bowel Dis. 2016;22:1999–2015.
Suh K, Kyei I, Hage DS. Approaches for the detection and analysis of antidrug antibodies to biopharmaceuticals: a review. J Sep Sci. 2022;45:2077–92.
Papamichael K, Vogelzang EH, Lambert J, Wolbink G, Cheifetz AS. Therapeutic drug monitoring with biologic agents in immune mediated inflammatory diseases. Expert Rev Clin Immunol. 2019;15:837–48.
Abraham NS, Hlatky MA, Antman EM, Bhatt DL, Bjorkman DJ, Clark CB, et al. ACCF/ACG/AHA 2010 expert consensus document on the concomitant use of proton pump inhibitors and thienopyridines: a focused update of the ACCF/ACG/AHA 2008 expert consensus document on reducing the gastrointestinal risks of antiplatelet therapy and NSAID use. J Am Coll Cardiology. 2010;56:2051–66.
Abdullah-Koolmees H, van Keulen AM, Nijenhuis M, Deneer VHM. Pharmacogenetics guidelines: overview and comparison of the DPWG, CPIC, CPNDS, and RNPGx Guidelines. Front Pharmacology. 2021;11:595219.
van Schaik RHN. CYP450 pharmacogenetics for personalizing cancer therapy. Drug Resist Updat. 2008;11:77–98.
Deneer VHM, van Schaik RHN. Evidence based drug dosing and pharmacotherapeutic recommendations per genotype. Methods Mol Biol. 2013;1015:345–54.
Danese S, Solitano V, Jairath V, Peyrin-Biroulet L. The future of drug development for inflammatory bowel disease: the need to ACT (advanced combination treatment). Gut. 2022;71:2380–7.
Stalgis C, Deepak P, Mehandru S, Colombel JF. Rational combination therapy to overcome the plateau of drug efficacy in inflammatory bowel disease. Gastroenterology. 2021;161:394–9.
Thomsen SB, Ungaro RC, Allin KH, Elmahdi R, Poulsen G, Andersson M, et al. Impact of thiopurine discontinuation at anti-tumour necrosis factor initiation in inflammatory bowel disease treatment: a nationwide Danish cohort study. Aliment Pharmacol Ther. 2022;55:1128–38.
Lauro R, Mannino F, Irrera N, Squadrito F, Altavilla D, Squadrito G, et al. Pharmacogenetics of biological agents used in inflammatory bowel disease: a systematic review. Biomedicines. 2021;9:1748.
Hindorf U, Appell ML. Genotyping should be cSonsidered the primary choice for pre-treatment evaluation of thiopurine methyltransferase function. J Crohns Colitis. 2012;6:655–9.
Chang JY, Park SJ, Jung ES, Jung SA, Moon CM, Chun J, et al. Genotype-based treatment with thiopurine reduces incidence of myelosuppression in patients with inflammatory bowel diseases. Clin Gastroenterol Hepatol. 2020;18:2010–2018.e2.
Higgs JE, Payne K, Roberts C, Newman WG. Are patients with intermediate TPMT activity at increased risk of myelosuppression when taking thiopurine medications? Pharmacogenomics. 2010;11:177–88.
Yang SK, Hong M, Baek J, Choi H, Zhao W, Jung Y, et al. A common missense variant in NUDT15 confers susceptibility to thiopurine-induced leukopenia. Nat Genet. 2014;46:1017–20.
Hoffmann P, Lamerz D, Hill P, Kirchner M, Gauss A. Gene Polymorphisms of NOD2, IL23R, PTPN2 and ATG16L1 in patients with crohn’s disease: on the way to personalized medicine? Genes. 2021;12:866.
Author information
Authors and Affiliations
Contributions
TA, HN and VA designed the study, conceived the work, revised the manuscript, approved the final version, and agreed on the aspects of the work. TA performed data analysis. JAGA revised the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
TA, JAGA and HN have no competing financial interests in relation to the work described. VA is supported by Sundhedsdonationer (2024-0379).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Andresen, T., Agúndez, J.A.G., Nazari, H. et al. Potential of pharmacogenetics in treatment of chronic inflammatory diseases – a danish report overview from 2000 – 2024. Pharmacogenomics J 26, 5 (2026). https://doi.org/10.1038/s41397-026-00400-w
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41397-026-00400-w