Abstract
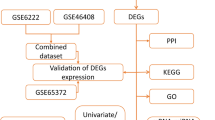
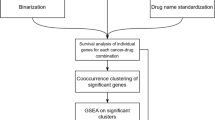
Identification of the functional impact of mutated and altered genes in cancer is critical for implementing precision oncology and drug repurposing. In recent years, the emergence of multiomics data from large, well-characterized patient cohorts has provided us with an unprecedented opportunity to address this problem. In this study, we investigated survival-associated genes across 26 cancer types and found that these genes tended to be hub genes and had higher K-core values in biological networks. Moreover, the genes associated with adverse outcomes were mainly enriched in pathways related to genetic information processing and cellular processes, while the genes with favorable outcomes were enriched in metabolism and immune regulation pathways. We proposed using the number of survival-related neighbors to assess the impact of mutations. In addition, by integrating other databases including the Human Protein Atlas and the DrugBank database, we predicted novel targets and anticancer drugs using the drug repurposing strategy. Our results illustrated the significance of multidimensional analysis of clinical data in important gene identification and drug development.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA. Cancer Genome Atlas Research Network et al. The cancer genome atlas pan-cancer analysis project. Nat Genet. 2013;45:1113–20.
Chen JC, Alvarez MJ, Talos F, Dhruv H, Rieckhof GE, Iyer A, et al. Identification of causal genetic drivers of human disease through systems-level analysis of regulatory networks. Cell. 2014;159:402–14.
McDonald ER III, de Weck A, Schlabach MR, Billy E, Mavrakis KJ, Hoffman GR, et al. Project DRIVE: a compendium of cancer dependencies and synthetic lethal relationships uncovered by large-Scale, deep RNAi screening. Cell. 2017;170:577–92 e10.
Tsherniak A, Vazquez F, Montgomery PG, Weir BA, Kryukov G, Cowley GS, et al. Defining a cancer dependency map. Cell. 2017;170:564–76 e16.
Aguirre AJ, Meyers RM, Weir BA, Vazquez F, Zhang CZ, Ben-David U, et al. Genomic copy number dictates a gene-independent cell response to CRISPR/Cas9 targeting. Cancer Discov. 2016;6:914–29.
Ng PK, Li J, Jeong KJ, Shao S, Chen H, Tsang YH, et al. Systematic functional annotation of somatic mutations in cancer. Cancer Cell. 2018;33:450–62 e10.
Leiserson MD, Vandin F, Wu HT, Dobson JR, Eldridge JV, Thomas JL, et al. Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes. Nat Genet. 2015;47:106–14.
Porta-Pardo E, Kamburov A, Tamborero D, Pons T, Grases D, Valencia A, et al. Comparison of algorithms for the detection of cancer drivers at subgene resolution. Nat Methods. 2017;14:782.
Gao JJ, Chang MT, Johnsen HC, Gao SP, Sylvester BE, Sumer SO, et al. 3D clusters of somatic mutations in cancer reveal numerous rare mutations as functional targets. Genome Med. 2017;9:4.
Niu B, Scott AD, Sengupta S, Bailey MH, Batra P, Ning J, et al. Protein-structure-guided discovery of functional mutations across 19 cancer types. Nat Genet. 2016;48:827–37.
Tokheim C, Bhattacharya R, Niknafs N, Gygax DM, Kim R, Ryan M, et al. Exome-scale discovery of hotspot mutation regions in human cancer using 3D protein structure. Cancer Res. 2016;76:3719–31.
Jin GX, Wong STC. Toward better drug repositioning: prioritizing and integrating existing methods into efficient pipelines. Drug Discov Today. 2014;19:637–44.
Ashburn TT, Thor KB. Drug repositioning: identifying and developing new uses for existing drugs. Nat Rev Drug Discov. 2004;3:673–83.
Ben Sahra I, Le Marchand-Brustel Y, Tanti JF, Bost F. Metformin in cancer therapy: a new perspective for an old antidiabetic drug? Mol Cancer Ther. 2010;9:1092–9.
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, et al. Tissue-based map of the human proteome. Science. 2015;347:1260419.
Law V, Knox C, Djoumbou Y, Jewison T, Guo AC, Liu Y, et al. DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res. 2014;42:D1091–7.
Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 2011;12:R41.
Keshava Prasad TS, Goel R, Kandasamy K, Keerthikumar S, Kumar S, Mathivanan S, et al. Human protein reference database–2009 update. Nucleic Acids Res. 2009;37:D767–72.
Rolland T, Tasan M, Charloteaux B, Pevzner SJ, Zhong Q, Sahni N, et al. A proteome-scale map of the human interactome network. Cell. 2014;159:1212–26.
Yu GC, Wang LG, Han YY, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics. 2012;16:284–7.
Barabasi AL, Oltvai ZN. Network biology: understanding the cell’s functional organization. Nat Rev Genet. 2004;5:101–13.
Yang Y, Han L, Yuan Y, Li J, Hei N, Liang H. Gene co-expression network analysis reveals common system-level properties of prognostic genes across cancer types. Nat Commun. 2014;5:3231.
Rooney MS, Shukla SA, Wu CJ, Getz G, Hacohen N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell. 2015;160:48–61.
Bullock TNJ. TNF-receptor superfamily agonists as molecular adjuvants for cancer vaccines. Curr Opin Immunol. 2017;47:70–7.
Lawrence MS, Stojanov P, Polak P, Kryukov GV, Cibulskis K, Sivachenko A, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. 2013;499:214–8.
Hollstein M, Sidransky D, Vogelstein B, Harris CC. p53 mutations in human cancers. Science. 1991;253:49–53.
Hellmann MD, Callahan MK, Awad MM, Calvo E, Ascierto PA, Atmaca A, et al. Tumor mutational burden and efficacy of nivolumab monotherapy and in combination with ipilimumab in small-cell lung cancer. Cancer Cell. 2018;33:853–61 e4.
Hellmann MD, Ciuleanu TE, Pluzanski A, Lee JS, Otterson GA, Audigier-Valette C, et al. Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N Engl J Med. 2018;378:2093–104.
Erson-Omay EZ, Caglayan AO, Schultz N, Weinhold N, Omay SB, Ozduman K, et al. Somatic POLE mutations cause an ultramutated giant cell high-grade glioma subtype with better prognosis. Neuro Oncol. 2015;17:1356–64.
Cancer Genome Atlas Research N, Kandoth C, Schultz N, Cherniack AD, Akbani R, Liu Y, et al. Integrated genomic characterization of endometrial carcinoma. Nature. 2013;497:67–73.
Campos-Arroyo D, Maldonado V, Bahena I, Quintanar V, Patino N, Carlos Martinez-Lazcano J, et al. Probenecid sensitizes neuroblastoma cancer stem cells to cisplatin. Cancer Invest. 2016;34:155–66.
Jones MR, Schrader KA, Shen Y, Pleasance E, Ch’ng C, Dar N, et al. Response to angiotensin blockade with irbesartan in a patient with metastatic colorectal cancer. Ann Oncol. 2016;27:801–6.
Segura-Pacheco B, Perez-Cardenas E, Taja-Chayeb L, Chavez-Blanco A, Revilla-Vazquez A, Benitez-Bribiesca L, et al. Global DNA hypermethylation-associated cancer chemotherapy resistance and its reversion with the demethylating agent hydralazine. J Transl Med. 2006;4:32.
Altieri DC. Survivin, cancer networks and pathway-directed drug discovery. Nat Rev Cancer. 2008;8:61–70.
Consortium I T P-C A o W G. Pan-cancer analysis of whole genomes. Nature. 2020;578:82–93.
Mitra R, Adams CM, Jiang W, Greenawalt E, Eischen CM. Pan-cancer analysis reveals cooperativity of both strands of microRNA that regulate tumorigenesis and patient survival. Nat Commun. 2020;11:968.
Wong HS, Chang CM, Liu X, Huang WC, Chang WC. Characterization of cytokinome landscape for clinical responses in human cancers. Oncoimmunology. 2016;5:e1214789.
Wong HS, Chang WC. Losses of cytokines and chemokines are common genetic features of human cancers: the somatic copy number alterations are correlated with patient prognoses and therapeutic resistance. Oncoimmunology. 2018;7:e1468951.
Carli F, Chiellini EE, Bellich B, Macchiavelli S, Cadelli G. Ubidecarenone nanoemulsified composite systems. Int J Pharm. 2005;291:113–8.
Acknowledgements
This study was funded by the National Natural Science Foundation of China (81602620, 81502086, 81572896, 8172203 and 91859205). National Major Scientific and Technological Special Project for “Significant New Drugs Development” (2018ZX09101-002), the Scientific Research Foundation of Shanghai Municipal Commission of Health and Family Planning (No. 20154Y0140) and the Shanghai Pujiang Program (18PJD060).
Author information
Authors and Affiliations
Contributions
YL and YPD performed the centrality analysis, drafted and revised the manuscript. YWQ, LXY and WW participated in the statistical analysis and manuscript writing. XLC and HYW conceived and supervised this study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Li, Y., Dong, Yp., Qian, Yw. et al. Identification of important genes and drug repurposing based on clinical-centered analysis across human cancers. Acta Pharmacol Sin 42, 282–289 (2021). https://doi.org/10.1038/s41401-020-0451-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41401-020-0451-1
Keywords
This article is cited by
-
Pan-cancer and multiomics: advanced strategies for diagnosis, prognosis, and therapy in the complex genetic and molecular universe of cancer
Clinical and Translational Oncology (2024)