Abstract
The forkhead box M1 (FoxM1) protein, a transcription factor, plays critical roles in regulating tumor growth and drug resistance, while cellular FLICE-inhibitory protein (c-FLIP), an anti-apoptotic regulator, is involved in the ubiquitin–proteasome pathway. In this study, we investigated the effects of c-FLIP on the expression and ubiquitination levels of FoxM1 along with drug susceptibility in non-small-cell lung cancer (NSCLC) cells. We first showed that the expression levels of FoxM1 and c-FLIP were increased and positively correlated (R2 = 0.1106, P < 0.0001) in 90 NSCLC samples. The survival data from prognostic analysis demonstrated that high expression of c-FLIP and/or FoxM1 was related to poor prognosis in NSCLC patients and that the combination of FoxM1 and c-FLIP could be a more precise prognostic biomarker than either alone. Then, we explored the functions of c-FLIP/FoxM1 in drug resistance in NSCLC cell lines and a xenograft mouse model in vivo. We showed that c-FLIP stabilized FoxM1 by inhibiting its ubiquitination, thus upregulated the expression of FoxM1 at post-transcriptional level. In addition, a positive feedback loop composed of FoxM1, β-catenin and p65 also participated in c-FLIP–FoxM1 axis. We revealed that c-FLIP promoted the resistance of NSCLC cells to thiostrepton and osimertinib by upregulating FoxM1. Taken together, these results reveal a new mechanism by which c-FLIP regulates FoxM1 and the function of this interaction in the development of thiostrepton and osimertinib resistance. This study provides experimental evidence for the potential therapeutic benefit of targeting the c-FLIP–FoxM1 axis for lung cancer treatment.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Change history
23 July 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41401-024-01343-3
References
Ros S, Wright AJ, D’Santos P, Hu DE, Hesketh RL, Lubling Y, et al. Metabolic imaging detects resistance to PI3Kalpha inhibition mediated by persistent FOXM1 expression in ER+ breast cancer. Cancer Cell. 2020;38:516–33e9.
Nilsson MB, Sun H, Robichaux J, Pfeifer M, McDermott U, Travers J, et al. A YAP/FOXM1 axis mediates EMT-associated EGFR inhibitor resistance and increased expression of spindle assembly checkpoint components. Sci Transl Med. 2020;12:eaaz4589.
Zhang B, Zhang Y, Zou X, Chan AW, Zhang R, Lee TK, et al. The CCCTC-binding factor (CTCF)-forkhead box protein M1 axis regulates tumour growth and metastasis in hepatocellular carcinoma. J Pathol. 2017;243:418–30.
Shang R, Wang M, Dai B, Du J, Wang J, Liu Z, et al. Long noncoding RNA SLC2A1-AS1 regulates aerobic glycolysis and progression in hepatocellular carcinoma via inhibiting the STAT3/FOXM1/GLUT1 pathway. Mol Oncol. 2020;14:1381–96.
Wang D, Hu G, Du Y, Zhang C, Lu Q, Lv N, et al. Aberrant activation of hedgehog signaling promotes cell proliferation via the transcriptional activation of forkhead Box M1 in colorectal cancer cells. J Exp Clin Cancer Res. 2017;36:23.
Liao GB, Li XZ, Zeng S, Liu C, Yang SM, Yang L, et al. Regulation of the master regulator FOXM1 in cancer. Cell Commun Signal. 2018;16:57.
Zheng S, Zhou C, Wang Y, Li H, Sun Y, Shen Z. TRIM6 promotes colorectal cancer cells proliferation and response to thiostrepton by TIS21/FoxM1. J Exp Clin Cancer Res. 2020;39:23.
Kongsema M, Zona S, Karunarathna U, Cabrera E, Man EP, Yao S, et al. RNF168 cooperates with RNF8 to mediate FOXM1 ubiquitination and degradation in breast cancer epirubicin treatment. Oncogenesis. 2016;5:e252.
Li LQ, Pan D, Chen H, Zhang L, Xie WJ. F-box protein FBXL2 inhibits gastric cancer proliferation by ubiquitin-mediated degradation of forkhead box M1. FEBS Lett. 2016;590:445–52.
Chen Z, Li L, Xu S, Liu Z, Zhou C, Li Z, et al. A Cdh1-FoxM1-Apc axis controls muscle development and regeneration. Cell Death Dis. 2020;11:180.
Choi YE, Madhi H, Kim H, Lee JS, Kim MH, Kim YN, et al. FAM188B expression is critical for cell growth via FOXM1 regulation in lung cancer. Biomedicines. 2020;8:465.
Ning Z, Wang A, Liang J, Xie Y, Liu J, Feng L, et al. USP22 promotes the G1/S phase transition by upregulating FoxM1 expression via beta-catenin nuclear localization and is associated with poor prognosis in stage II pancreatic ductal adenocarcinoma. Int J Oncol. 2014;45:1594–608.
Li XY, Wu HY, Mao XF, Jiang LX, Wang YX. USP5 promotes tumorigenesis and progression of pancreatic cancer by stabilizing FoxM1 protein. Biochem Biophys Res Commun. 2017;492:48–54.
Song Z, Li J, Zhang L, Deng J, Fang Z, Xiang X, et al. UCHL3 promotes pancreatic cancer progression and chemo-resistance through FOXM1 stabilization. Am J Cancer Res. 2019;9:1970–81.
Ishioka T, Katayama R, Kikuchi R, Nishimoto M, Takada S, Takada R, et al. Impairment of the ubiquitin-proteasome system by cellular FLIP. Genes Cells. 2007;12:735–44.
Lei S, Yang J, Chen C, Sun J, Yang L, Tang H, et al. FLIP(L) is critical for aerobic glycolysis in hepatocellular carcinoma. J Exp Clin Cancer Res. 2016;35:79.
Naito M, Katayama R, Ishioka T, Suga A, Takubo K, Nanjo M, et al. Cellular FLIP inhibits beta-catenin ubiquitylation and enhances Wnt signaling. Mol Cell Biol. 2004;24:8418–27.
Nakagiri S, Murakami A, Takada S, Akiyama T, Yonehara S. Viral FLIP enhances Wnt signaling downstream of stabilized beta-catenin, leading to control of cell growth. Mol Cell Biol. 2005;25:9249–58.
Shi P, Oh YT, Deng L, Zhang G, Qian G, Zhang S, et al. Overcoming acquired resistance to AZD9291, a third-generation EGFR inhibitor, through modulation of MEK/ERK-dependent Bim and Mcl-1 degradation. Clin Cancer Res. 2017;23:6567–79.
Wang WD, Shang Y, Li Y, Chen SZ. Honokiol inhibits breast cancer cell metastasis by blocking EMT through modulation of Snail/Slug protein translation. Acta Pharmacol Sin. 2019;40:1219–27.
Ni J, Wang X, Shang Y, Li Y, Chen S. CD13 inhibition augments DR4-induced tumor cell death in a p-ERK1/2-independent manner. Cancer Biol Med. 2021;18:569–86.
Li X, Qiu W, Liu B, Yao R, Liu S, Yao Y, et al. Forkhead box transcription factor 1 expression in gastric cancer: FOXM1 is a poor prognostic factor and mediates resistance to docetaxel. J Transl Med. 2013;11:204.
Consolaro F, Basso G, Ghaem-Magami S, Lam EW, Viola G. FOXM1 is overexpressed in B-acute lymphoblastic leukemia (B-ALL) and its inhibition sensitizes B-ALL cells to chemotherapeutic drugs. Int J Oncol. 2015;47:1230–40.
Hegde NS, Sanders DA, Rodriguez R, Balasubramanian S. The transcription factor FOXM1 is a cellular target of the natural product thiostrepton. Nat Chem. 2011;3:725–31.
Jin B, Wang C, Li J, Du X, Ding K, Pan J. Anthelmintic niclosamide disrupts the interplay of p65 and FOXM1/beta-catenin and eradicates leukemia stem cells in chronic myelogenous leukemia. Clin Cancer Res. 2017;23:789–803.
Hardy LR, Pergande MR, Esparza K, Heath KN, Onyuksel H, Cologna SM, et al. Proteomic analysis reveals a role for PAX8 in peritoneal colonization of high grade serous ovarian cancer that can be targeted with micelle encapsulated thiostrepton. Oncogene. 2019;38:6003–16.
Shi P, Zhang S, Zhu L, Qian G, Ren H, Ramalingam SS, et al. The yhird-generation EGFR inhibitor, osimertinib, promotes c-FLIP degradation, enhancing apoptosis including TRAIL-induced apoptosis in NSCLC cells with activating EGFR mutations. Transl Oncol. 2019;12:705–13.
Nandi D, Cheema PS, Jaiswal N, Nag A. FoxM1: Repurposing an oncogene as a biomarker. Semin Cancer Biol. 2018;52:74–84.
Wang J, Li W, Zhao Y, Kang FW, Zheng X, et al. Members of FOX family could be drug targets of cancers. Pharmacol Ther. 2018;181:183–96.
Luebke T, Schwarz L, Beer YY, Schumann S, Misterek M, Sander FE, et al. c-FLIP and CD95 signaling are essential for survival of renal cell carcinoma. Cell Death Dis. 2019;10:384.
Bivona TG, Hieronymus H, Parker J, Chang K, Taron M, Rosell R, et al. FAS and NF-kappaB signalling modulate dependence of lung cancers on mutant EGFR. Nature. 2011;471:523–6.
Shirley S, Micheau O. Targeting c-FLIP in cancer. Cancer Lett. 2013;332:141–50.
McCourt C, Maxwell P, Mazzucchelli R, Montironi R, Scarpelli M, Salto-Tellez M, et al. Elevation of c-FLIP in castrate-resistant prostate cancer antagonizes therapeutic response to androgen receptor-targeted therapy. Clin Cancer Res. 2012;18:3822–33.
Weiler SME, Pinna F, Wolf T, Lutz T, Geldiyev A, Sticht C, et al. Induction of chromosome instability by activation of yes-associated protein and Forkhead box M1 in liver cancer. Gastroenterology. 2017;152:2037–-51e22.
Hu G, Su Y, Kang BH, Fan Z, Dong T, Brown DR, et al. High-throughput phenotypic screen and transcriptional analysis identify new compounds and targets for macrophage reprogramming. Nat Commun. 2021;12:773.
Wang Y, Xie W, Humeau J, Chen G, Liu P, Pol J, et al. Autophagy induction by thiostrepton improves the efficacy of immunogenic chemotherapy. J Immunother Cancer 2020;8:e000462.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (81702934), CAMS Innovation Fund for Medical Sciences (CIFMS, 2019-I2M-1-003, 2021-I2M-1-030), and Beijing Natural Science Foundation (7202132, 7192041).
Author information
Authors and Affiliations
Contributions
SZC designed the project. WDW and SZC designed the experiments and wrote the paper. WDW performed the research. YS and JN assisted with the murine experiments. CW, AMW, and GJL assisted with the cell-based high-throughput drug screen. LS offered the stable H157-Lac-Z and c-FLIP cells.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
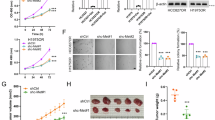
The original online version of this article was revised: The ‘HCC827’ in the sentence of Results section ‘Among these cells, only HCC827 cells showed significantly high mRNA levels of c-FLIP and FoxM1, while only H1975 cells showed significantly low mRNA levels of c-FLIP and FoxM1 (Fig. 6a)’ should be changed to ‘A549’ according to the original data. For Fig6c, the color of the columns in c-FLIP should be marked in orange. The reason for this error is that when editing Fig.6c in AI software, the color of the columns in c-FLIP was omitted.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Wd., Shang, Y., Wang, C. et al. c-FLIP promotes drug resistance in non-small-cell lung cancer cells via upregulating FoxM1 expression. Acta Pharmacol Sin 43, 2956–2966 (2022). https://doi.org/10.1038/s41401-022-00905-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41401-022-00905-7
Keywords
This article is cited by
-
Beyond Death: Unmasking the Intricacies of Apoptosis Escape
Molecular Diagnosis & Therapy (2024)
-
Comprehensive clinicopathological significance and putative transcriptional mechanisms of Forkhead box M1 factor in hepatocellular carcinoma
World Journal of Surgical Oncology (2023)
-
MiR-3677-3p promotes development and sorafenib resistance of hepatitis B-related hepatocellular carcinoma by inhibiting FOXM1 ubiquitination
Human Cell (2023)