Abstract
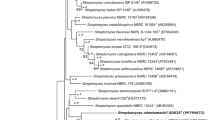
Streptomyces strain TP-A0890T, isolated from a soil sample, is a producer of FR-900452 and A-74863a. The taxonomic status was clarified by a polyphasic approach. Phylogenetic analysis based on 16S rRNA gene sequences showed that the strain was closely related to Streptomyces coriariae, with similarity of 99.7%. Strain TP-A0890T comprised ll-diaminopimelic acid, glutamic acid, glycine and alanine in its peptidoglycan. The predominant menaquinones were MK-9(H8) and MK-9(H6), and major fatty acids were anteiso-C17:0, anteiso-C15:0, iso-C16:0 and iso-C17:0. The chemotaxonomic features matched those described for the genus Streptomyces. The genome size and G + C content were 8.72 Mb and 71.5%, respectively. The results of digital DNA-DNA hybridization along with differences in phenotypic characteristics between the strains suggested strain TP-A0890T to assign to a novel species, for which Streptomyces maremycinicus sp. nov. is proposed; the type strain is TP-A0890T ( = NBRC 110468T). We also show that Streptomyces strain B9173, a producer of FR-900452 and maremycins that was isolated from coastal sediment in Chile, belonged to S. maremycinicus. Twenty-two to 23 secondary metabolite-biosynthetic gene clusters (smBGCs) were present in the genomes of S. maremycinicus strains. Seventeen of them were conserved in the genome of S. coriariae CMB-FBT but the others were not.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Baltz RH. Gifted microbes for genome mining and natural product discovery. J Ind Microbiol Biotechnol. 2017;44:573–88.
Berdy J. Bioactive microbial metabolites. J Antibiot. 2005;58:1–26.
Komaki H. Recent progress of reclassification of the genus Streptomyces. Microorganisms. 2023;11:831.
Komaki H, Ichikawa N, Hosoyama A, Fujita N, Igarashi Y. Draft genome sequence of Streptomyces sp. TP-A0890, a producer of FR-900452 and A-74863a. Genome Announc. 2015;3:e01212–01215.
Duan Y, Liu Y, Huang T, Zou Y, Huang T, Hu K, et al. Divergent biosynthesis of indole alkaloids FR900452 and spiro-maremycins. Org Biomol Chem. 2018;16:5446–51.
Kuroda M, Ogita T, Enokita R, Okazaki T, Kinoshita T. Japan Patent 1995:1995/010875.
Yoon S, Ha S, Kwon S, Lim J, Kim Y, Seo H, et al. Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol. 2017;67:1613–7.
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–25.
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, et al. Clustal W and Clustal X version 2.0. Bioinformatics. 2007;23:2947–8.
Shirling EB, Gottlieb D. Methods for characterization of Streptomyces species. Int J Syst Bacteriol. 1966;16:313–40.
Hayakawa M, Nonomura H. Humic acid-vitamin agar, a new medium for the selective isolation of soil actinomycetes. J Ferment Technol. 1987;65:501–9.
Hasegawa T, Takizawa M, Tanida S. A rapid analysis for chemical grouping of aerobic actinomycetes. J Gen Appl Microbiol. 1983;29:319–22.
Tamura T, Ishida Y, Suzuki K. Descriptions of Actinoplanes ianthinogenes nom. rev. and Actinoplanes octamycinicus corrig. comb. nov., nom. rev. Int J Syst Evol Microbiol. 2011;61:2916–21.
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, et al. An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. Microbiol Methods. 1984;2:233–41.
Hamada M, Iino T, Iwami T, Harayama S, Tamura T, Suzuki K. Mobilicoccus pelagius gen. nov., sp. nov. and Piscicoccus intestinalis gen. nov., sp. nov., two new members of the family Dermatophilaceae, and reclassification of Dermatophilus chelonae (Masters et al. 1995) as Austwickia chelonae gen. nov., comb. nov. J Gen Appl Microbiol. 2010;56:427–36.
Yassin AF, Haggenei B, Budzikiewicz H, Schaal KP. Fatty acid and polar lipid composition of the genus Amycolatopsis: application of fast atom bombardment-mass spectrometry to structure analysis of underivatized phospholipid. Int J Syst Bacteriol. 1993;43:414–20.
Meier-Kolthoff JP, Carbasse JS, Peinado-Olarte RL, Goker M. TYGS and LPSN: a database tandem for fast and reliable genome-based classification and nomenclature of prokaryotes. Nucleic Acids Res. 2022;50:D801–D807.
Richter M, Rossello-Mora R, Oliver Glockner F, Peplies J. JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics. 2016;32:929–31.
Blin K, Shaw S, Kloosterman AM, Charlop-Powers Z, van Wezel GP, Medema MH, et al. antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021;49:W29–W35.
Komaki H, Ichikawa N, Hosoyama A, Takahashi-Nakaguchi A, Matsuzawa T, Suzuki K, et al. Genome based analysis of type-I polyketide synthase and nonribosomal peptide synthetase gene clusters in seven strains of five representative Nocardia species. BMC Genom. 2014;15:323.
Komaki H, Ichikawa N, Oguchi A, Hanamaki T, Fujita N. Genome-wide survey of polyketide synthase and nonribosomal peptide synthetase gene clusters in Streptomyces turgidiscabies NBRC 16081. J Gen Appl Microbiol. 2012;58:363–72.
Meier-Kolthoff JP, Goker M, Sproer C, Klenk HP. When should a DDH experiment be mandatory in microbial taxonomy?. Arch Microbiol. 2013;195:413–8.
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, et al. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol. 1987;37:463–4.
Luo F, Zou Y, Huang T, Lin S. Draft genome sequence of Streptomyces sp. B9173, a producer of indole diketopiperazine maremycins. Genome Announc. 2017;5:e00447–00417.
Kodani S, Komaki H, Suzuki M, Hemmi H, Ohnishi-Kameyama M. Isolation and structure determination of new siderophore albachelin from Amycolatopsis alba. Biometals. 2015;28:381–9.
Liu Z, Huang T, Shi Q, Deng Z, Lin S. Catechol siderophores framed on 2,3-dihydroxybenzoyl-L-serine from Streptomyces varsoviensis. Front Microbiol. 2023;14:1182449.
Raymond KN, Dertz EA, Kim SS. Enterobactin: an archetype for microbial iron transport. Proc Natl Acad Sci USA. 2003;100:3584–8.
Maruyama C, Niikura H, Izumikawa M, Hashimoto J, Shin-ya K, Komatsu M, et al. tRNA-dependent aminoacylation of an amino sugar intermediate in the biosynthesis of a streptothricin-related antibiotic. Appl Environ Microbiol. 2016;82:3640–8.
Hoshino S, Ijichi S, Asamizu S, Onaka H. Insights into arsenic secondary metabolism in actinomycetes from the structure and biosynthesis of bisenarsan. J Am Chem Soc. 2023;145:17863–71.
Wang B, Guo F, Huang C, Zhao H. Unraveling the iterative type I polyketide synthases hidden in Streptomyces. Proc Natl Acad Sci USA. 2020;117:8449–54.
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, da Costa MS, et al. Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol. 2018;68:461–6.
Berckx F, Bandong CM, Wibberg D, Kalinowski J, Willemse J, Brachmann A, et al. Streptomyces coriariae sp. nov., a novel streptomycete isolated from actinorhizal nodules of Coriaria intermedia. Int J Syst Evol Microbiol. 2022;72:005603.
Vela Gurovic MS, Diaz ML, Gallo CA, Dietrich J. Phylogenomics, CAZyome and core secondary metabolome of Streptomyces albus species. Mol Genet Genom. 2021;296:1299–311.
Komaki H, Tamura T. Profile of PKS and NRPS gene clusters in the genome of Streptomyces cellostaticus NBRC 12849T. Fermentation. 2023;9:924.
Acknowledgements
We are grateful to Ms. Satomi Saitou for assistance of taxonomic experiments. We also thank Ms. Yuko Kitahashi and Ms. Aya Uohara for finishing the genome sequences and registering the genome sequences in the DDBJ, respectively. We thank Dr. Aharon Oren for reviewing specific epithets and the etymologies.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Komaki, H., Igarashi, Y. & Tamura, T. Streptomyces maremycinicus sp. nov. and its secondary metabolite-biosynthetic gene clusters. J Antibiot 78, 560–568 (2025). https://doi.org/10.1038/s41429-025-00844-5
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41429-025-00844-5