Abstract
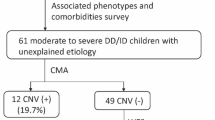
Next generation sequencing provides an important opportunity for improved diagnosis in epilepsy. To date, the majority of diagnostic genetic testing is conducted in the paediatric arena, while the utility of such testing is less well understood in adults with epilepsy. We conducted whole exome sequencing (WES) and copy number variant analyses in an Irish cohort of 101 people with epilepsy and co-morbid intellectual disability to compare the diagnostic yield of genomic testing between adult and paediatric patients. Variant interpretation followed American College of Medical Genetics and Genomics (ACMG) guidelines. We demonstrate that WES, in combination with array-comparative genomic hybridisation, provides a diagnostic rate of 27% in unrelated adult epilepsy patients and 42% in unrelated paediatric patients. We observe a 2.7% rate of ACMG-defined incidental findings. Our findings indicate that WES has similar utility in both adult and paediatric cohorts and is appropriate for diagnostic testing in both epilepsy patient groups.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Notes
One of these CNVs (patient 84) awaits confirmation and segregation testing from accredited laboratory.
References
Covanis A, Guekht A, Li S, Secco M, Shakir R, Perucca E. From global campaign to global commitment: The World Health Assembly’s Resolution on epilepsy. Epilepsia. 2015;56:1651–7.
Myers CT, Mefford HC. Advancing epilepsy genetics in the genomic era. Genome Med BioMed Cent. 2015;7:91.
Allen AS, Berkovic SF, Cossette P, Delanty N, Dlugos D, Eichler EE, et al. De novo mutations in epileptic encephalopathies. Nature. 2013;501:217–21.
Pagnamenta AT, Murakami Y, Taylor JM, Anzilotti C, Howard MF, Miller V, et al. Analysis of exome data for 4293 trios suggests GPI-anchor biogenesis defects are a rare cause of developmental disorders. Eur J Hum Genet. 2017;25:669–79.
Wright CF, Fitzgerald TW, Jones WD, Clayton S, McRae JF, Van Kogelenberg M, et al. Genetic diagnosis of developmental disorders in the DDD study: a scalable analysis of genome-wide research data. Lancet. 2015;385:1305–14.
Hamdan FF, Myers CT, Cossette P, Lemay P, Spiegelman D, Laporte AD, et al. High rate of recurrent de novo mutations in developmental and epileptic encephalopathies. Am J Hum Genet. 2017;101:664–85.
Ortega-Moreno L, Giráldez BG, Soto-Insuga V, Pozo RL Del, Rodrigo-Moreno M, Alarcón-Morcillo C. et al. Molecular diagnosis of patients with epilepsy and developmental delay using a customized panel of epilepsy genes. PLoS ONE. 2017;12:e0188978.
Allen NM, Conroy J, Shahwan A, Lynch B, Correa RG, Pena SDJ, et al. Unexplained early onset epileptic encephalopathy: exome screening and phenotype expansion. Epilepsia. 2016;57:e12–7.
Snoeijen-Schouwenaars FM, van Ool JS, Verhoeven JS, van Mierlo P, Braakman HMH, Smeets EE, et al. Diagnostic exome sequencing in 100 consecutive patients with both epilepsy and intellectual disability. Epilepsia. 2019;60:155–64.
Thevenon J, Duffourd Y, Masurel-Paulet A, Lefebvre M, Feillet F, El Chehadeh-Djebbar S, et al. Diagnostic odyssey in severe neurodevelopmental disorders: toward clinical whole-exome sequencing as a first-line diagnostic test. Clin Genet. 2016;89:700–7.
Helbig KL, Farwell Hagman KD, Shinde DN, Mroske C, Powis Z, Li S, et al. Diagnostic exome sequencing provides a molecular diagnosis for a significant proportion of patients with epilepsy. Genet Med. 2016;18:898–905.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Olson H, Shen Y, Avallone J, Sheidley BR, Pinsky R, Bergin AM, et al. Copy number variation plays an important role in clinical epilepsy. Ann Neurol. 2014;75:943–58.
Borlot F, Regan BM, Bassett AS, Stavropoulos DJ, Andrade DM. Prevalence of pathogenic copy number variation in adults with pediatric-onset epilepsy and intellectual disability. JAMA Neurol. 2017;74:1301–11.
Mefford HC. Copy number matters in epilepsy. Epilepsy Curr. 2015;15:180–2.
Galizia EC, Srikantha M, Palmer R, Waters JJ, Lench N, Ogilvie CM, et al. Array comparative genomic hybridization: results from an adult population with drug-resistant epilepsy and co-morbidities. Eur J Med Genet. 2012;55:342–8.
Coppola A, Cellini E, Stamberger H, Saarentaus E, Cetica V, Lal D, et al. Diagnostic implications of genetic copy number variation in epilepsy plus. Epilepsia. 2019;60:689–706.
Kearney HM, Thorland EC, Brown KK, Quintero-Rivera F, South ST. American College of Medical Genetics standards and guidelines for interpretation and reporting of postnatal constitutional copy number variants. Genet Med. 2011;13:680–5.
Striano P, Coppola A, Paravidino R, Malacarne M, Gimelli S, Robbiano A, et al. Clinical significance of rare copy number variations in epilepsy: a case-control survey using microarray-based comparative genomic hybridization. Arch Neurol. 2012;69:322–30.
Church DM, Schneider VA, Graves T, Auger K, Cunningham F, Bouk N, et al. Modernizing reference genome assemblies. PLoS Biol. 2011;9:e1001091.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(Jul):1754–60.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–91.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4:1073–81.
Schwarz JM, Cooper DN, Schuelke M, Seelow D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods. 2014;11:361–2.
McKusick-Nathans Institute of Genetic Medicine, Johns Hopkins University (Baltimore M. Online Mendelian Inheritance in Man, OMIM®. https://omim.org/.
Kalia SS, Adelman K, Bale SJ, Chung WK, Eng C, Evans JP, et al. Recommendations for reporting of secondary findings in clinical exome and genome sequencing, 2016 update (ACMG SF v2.0): a policy statement of the American College of Medical Genetics and Genomics. Genet Med. 2017;19:249–55.
MacDonald JR, Ziman R, Yuen RKC, Feuk L, Scherer SW. The database of genomic variants: a curated collection of structural variation in the human genome. Nucleic Acids Res. 2014;42:D986–92.
Delanty N, White M, Benson K, McCormack M, Heavin S, Comerford E, et al. Development of a genomics module within an epilepsy-specific electronic health record: toward genomic medicine in epilepsy care. Epilepsia. 2019;epi.16278.
Palmer EE, Schofield D, Shrestha R, Kandula T, Macintosh R, Lawson JA, et al. Integrating exome sequencing into a diagnostic pathway for epileptic encephalopathy: evidence of clinical utility and cost effectiveness. Mol Genet Genomic Med. 2018;6:186–99.
Green RC, Berg JS, Grody WW, Kalia SS, Korf BR, Martin CL, et al. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med. 2013;15:565–74.
Saha S, Barnett AG, Foldi C, Burne TH, Eyles DW, Buka SL. et al. Advanced paternal age is associated with impaired neurocognitive outcomes during infancy and childhood. PLoS Med. 2009;6:e1000040.
Sanders SJ, Murtha MT, Gupta AR, Murdoch JD, Raubeson MJ, Willsey AJ, et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature. 2012;485:237–41.
Carvill GL, Heavin SB, Yendle SC, McMahon JM, O’Roak BJ, Cook J, et al. Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat Genet. 2013;45:825–30.
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS, Church DM, et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014;42:D980–5.
Samueli S, Abraham K, Dressler A, Gröppel G, Mühlebner-Fahrngruber A, Scholl T, et al. Efficacy and safety of Everolimus in children with TSC—associated epilepsy—Pilot data from an open single-center prospective study. Orphanet J Rare Dis. 2016;11:145.
French JA, Lawson JA, Yapici Z, Ikeda H, Polster T, Nabbout R, et al. Adjunctive everolimus therapy for treatment-resistant focal-onset seizures associated with tuberous sclerosis (EXIST-3): a phase 3, randomised, double-blind, placebo-controlled study. Lancet. 2016;388:2153–63.
EpiPM Consortium. A roadmap for precision medicine in the epilepsies. Lancet Neurol. 2015;14:1219–28.
Nabbout R, Chemaly N, Chipaux M, Barcia G, Bouis C, Dubouch C, et al. Encephalopathy in children with Dravet syndrome is not a pure consequence of epilepsy. Orphanet J Rare Dis. 2013;8:176.
Heyne HO, Singh T, Stamberger H, Abou Jamra R, Caglayan H, Craiu D, et al. De novo variants in neurodevelopmental disorders with epilepsy. Nat Genet. 2018;50:1048–53.
EuroEPINOMICS-RES Consortium, Epilepsy Phenome/Genome Project and EC, Balling R, Barisic N, Baulac S, Caglayan H, Craiu D, et al. De novo mutations in synaptic transmission genes including DNM1 cause epileptic encephalopathies. Am J Hum Genet. 2014;95:360–70.
Smith LA, Douglas J, Braxton AA, Kramer K. Reporting incidental findings in clinical whole exome sequencing: incorporation of the 2013 ACMG recommendations into current practices of genetic counseling. J Genet Couns. 2015;24:654–62.
Berkovic SF. Genetics of epilepsy in clinical practice. Epilepsy Curr. 2015;15:192–6.
Mefford HC. Clinical genetic testing in epilepsy. Epilepsy Curr. 2015;15:197–201.
Forman EB, Gorman KM, Conroy J, Arthur N, Grant C, Ennis S, et al. Cost of exome sequencing in epileptic encephalopathy: is it “worth it”? Arch Dis Child. 2018;103:304.
Brunklaus A, Ellis R, Reavey E, Forbes GH, Zuberi SM. Prognostic, clinical and demographic features in SCN1A mutation-positive Dravet syndrome. Brain. 2012;135:2329–36.
Poduri A. When should genetic testing be performed in epilepsy patients? Epilepsy Curr. 2017;17:16–22.
Poduri A, Sheidley BR, Shostak S, Ottman R. Genetic testing in the epilepsies-developments and dilemmas. Nat Rev Neurol. 2014;10:293–9.
Ding L-E, Burnett L, Chesher D. The impact of reporting incidental findings from exome and whole-genome sequencing: predicted frequencies based on modeling. Genet Med. 2015;17:197–204.
Delanty N, Goldstein DB. Neuron NeuroView diagnostic exome sequencing: a new paradigm in neurology. 2013;80:841–3.
Acknowledgements
This publication has emanated from research supported in part by eHealth Ireland, Health Service Executive (HSE), Grant Number 13/CDA/2223 and a research grant from Science Foundation Ireland (SFI) under Grant Number 16/RC/3948 and co-funded under the European Regional Development Fund and by FutureNeuro industry partners. We thank Prof. Brendan Loftus and Catherine Moss of University College Dublin for access to sequencing platforms. We also thank the patients, their families, carers and clinical staff involved in the care of these patients for making this research possible.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
NJL, ASR and CAS are employed by Congenica Ltd.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Benson, K.A., White, M., Allen, N.M. et al. A comparison of genomic diagnostics in adults and children with epilepsy and comorbid intellectual disability. Eur J Hum Genet 28, 1066–1077 (2020). https://doi.org/10.1038/s41431-020-0610-3
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41431-020-0610-3
This article is cited by
-
Monogenic Epilepsies in Adult Epilepsy Clinics and Gene-Driven Approaches to Treatment
Current Neurology and Neuroscience Reports (2025)
-
Insights into the ANKRD11 variants and short-stature phenotype through literature review and ClinVar database search
Orphanet Journal of Rare Diseases (2024)
-
Experience of the first adult-focussed undiagnosed disease program in Australia (AHA-UDP): solving rare and puzzling genetic disorders is ageless
Orphanet Journal of Rare Diseases (2024)
-
Genetische Diagnostik der Epilepsien: Empfehlung der Kommission Epilepsie und Genetik der Deutschen Gesellschaft für Epileptologie (DGfE)
Clinical Epileptology (2023)
-
The association between early childhood onset epilepsy and attention-deficit hyperactivity disorder (ADHD) in 3237 children and adolescents with Autism Spectrum Disorder (ASD): a historical longitudinal cohort data linkage study
European Child & Adolescent Psychiatry (2023)