Fig. 1: RP mode reveals two distinct TF classes: short-range and long-range.
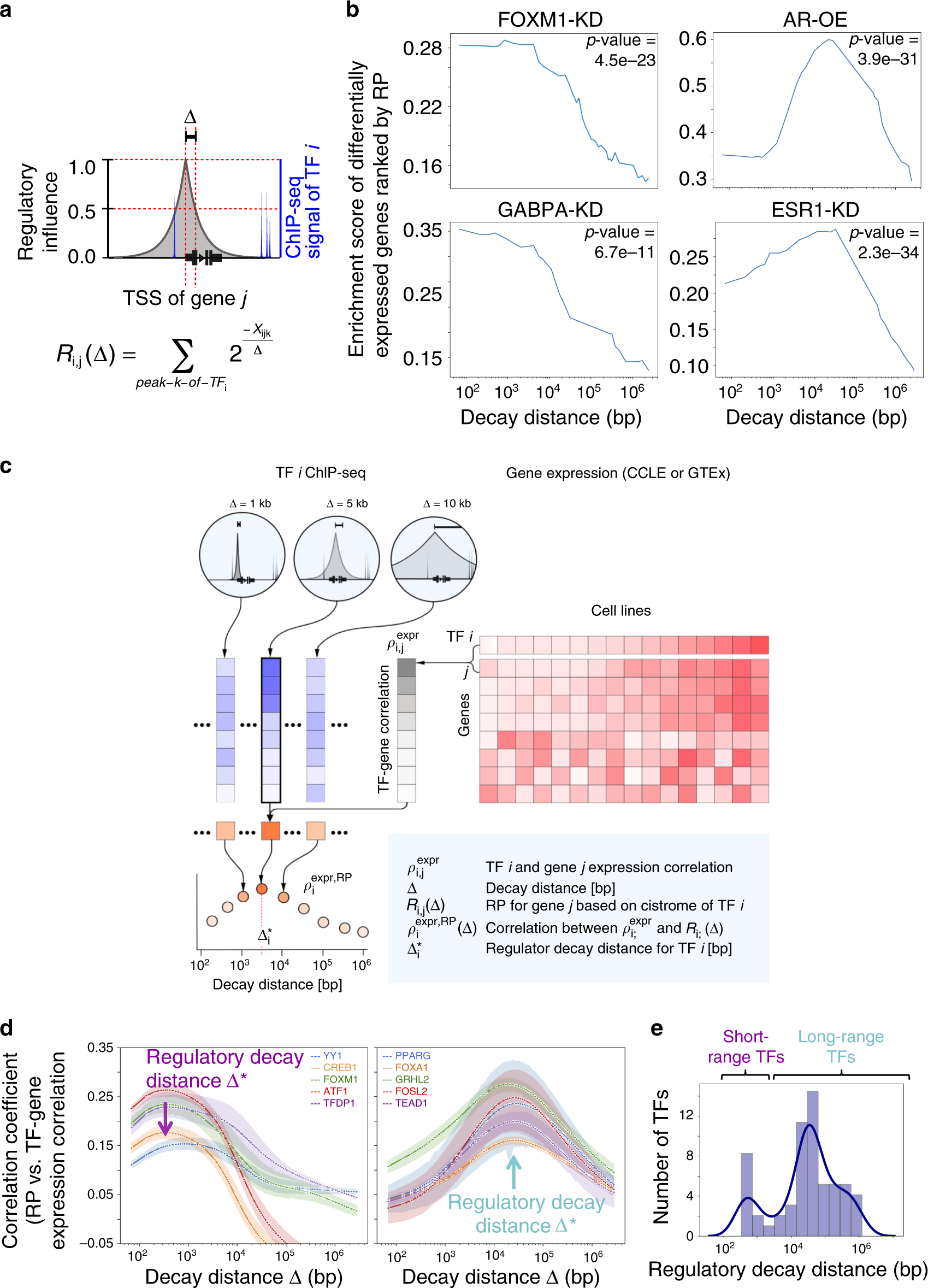
a Schematic of the regulatory potential (RP) model. The regulatory effect of TF i on gene j is modeled as the RP, Ri, j(Δ), which sums up all TF i ChIP-seq binding effects on the gene j. The effect of a single binding site k of TF i on gene j decays exponentially with increasing \(x_{ijk}\), the genomic distance between TSS of gene \(j\) and TF \(i\) binding site \(k\). The exponential decay function (\(2^{\frac{{ - x_{ijk}}}{\Delta }}\)) is parameterized by the decay distance (Δ), the distance at which the TF regulatory effects are halved. b TF \(i\)-specific regulatory decay distances (\({\mathrm{\Delta }}_i^ \ast\)) can be inferred as the Δ that best separates TF \(i\) perturbation-induced differentially expressed (DE) genes from other genes. \(R_{i,j}\left( \Delta \right)\) with short-range (<1 kb) best separates FOXM1-knockdown or GABPA-knockdown DE gene sets (left). AR overexpression or ESR1-knockdown DE gene sets are best separated by \(R_{i,j}\left( \Delta \right)\) with long-range Δ (>10 kb). The two-sided Kolmogorov–Smirnov two-sample test is used to estimate the degree of separation of DE genes from other genes. c \({\mathrm{\Delta }}_i^ \ast\) can also be inferred as the \(\Delta\) that leads to the best concordance between TF \(i\) regulatory effects estimated by TF \(i\) ChIP-seq (\(R_{i,j}\left( \Delta \right)\)) and expression cohorts (\(\rho _{i,j}^{{\mathrm{expr}}}\): TF i-gene \(j\) expression correlations), respectively. A second correlation coefficient \(\rho _i^{{\mathrm{expr}},{\mathrm{RP(}}\Delta {\mathrm{)}}}\) was calculated to measure the concordance between \(\rho _{i,j}^{{\mathrm{expr}}}\) and \(R_{i,j}\left( \Delta \right)\) (see the main text for the rationale and Methods for statistical details). d TFs with short-range \({\Delta}_i^ \ast\) (100bp-3 kb) include YY1, CREB1, FOXM1, ATF1, and TFDP1 (left). TFs with long-range \({\Delta}_i^ \ast\) (3 kb–100 kb) include PPARG, FOXA1, GRHL2, FOSL2, and TEAD1 (right). Colored shaded regions depict the 95% confidence intervals derived from all ChIP-seq samples that passed QC for each TF. Dots along the line are Δ values being tried. e Distribution of regulatory decay distances (\({\Delta}_i^ \ast\)) of 11 short-range TFs (left) and 49 long-range TFs (right). Source data are provided as a Source Data file.
