Abstract
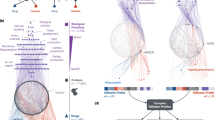
Drug action is inherently multiscale: it connects molecular interactions to emergent properties at cellular and larger scales. Simulation techniques at each of these different scales are already central to drug design and development, but methods capable of connecting across these scales will extend our understanding of complex mechanisms and our ability to predict biological effects. Improved algorithms, ever-more-powerful computing architectures and the accelerating growth of rich data sets are driving advances in multiscale modelling methods capable of bridging chemical and biological complexity from the atom to the cell. Particularly exciting is the development of highly detailed, structure-based physical simulations of biochemical systems, which can now reach experimentally relevant timescales for large systems and, at the same time, achieve unprecedented accuracy. In this Perspective, we discuss how emerging data-rich, physics-based multiscale approaches are on the cusp of realizing their long-promised impact on the discovery, design and development of novel therapeutics. We highlight emerging methods and applications in this growing field and outline how different scales can be combined in practical modelling and simulation strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Jorgensen, W. L. The many roles of computation in drug discovery. Science 303, 1813–1818 (2004).
Vivo, M. De, Masetti, M., Bottegoni, G. & Cavalli, A. Role of molecular dynamics and related methods in drug discovery. J. Med. Chem. 59, 4035–4061 (2016).
Lee, E. H., Hsin, J., Sotomayor, M., Comellas, G. & Schulten, K. Discovery through the computational microscope. Structure 17, 1295–1306 (2009).
Wassman, C. D. et al. Computational identification of a transiently open L1/S3 pocket for reactivation of mutant p53. Nat. Commun. 4, 1407 (2013).
Woods, C. J., Malaisree, M., Long, B., McIntosh-Smith, S. & Mulholland, A. J. Computational assay of H7N9 influenza neuraminidase reveals R292K mutation reduces drug binding affinity. Sci. Rep. 3, 3561 (2013).
Zhao, G. et al. Mature HIV-1 capsid structure by cryo-electron microscopy and all-atom molecular dynamics. Nature 497, 643–646 (2013).
Perilla, J. R. & Schulten, K. Physical properties of the HIV-1 capsid from all-atom molecular dynamics simulations. Nat. Commun. 8, 15959 (2017).
Wang, L. et al. Accurate and reliable prediction of relative ligand binding potency in prospective drug discovery by way of a modern free-energy calculation protocol and force field. J. Am. Chem. Soc. 137, 2695–2703 (2015).
Kulik, H. J., Luehr, N., Ufimtsev, I. S. & Martínez, T. J. Ab initio quantum chemistry for protein structures. J. Phys. Chem. B 116, 12501–12509 (2012).
Yu, I., Mori, T., Ando, T., Harada, R. & Jung, J. Biomolecular interactions modulate macromolecular structure and dynamics in atomistic model of a bacterial cytoplasm. eLife 5, e19274 (2016).
Dye, A. & Kramer, W. 2016 Annual Report Sustained Petascale in Action (NCSA Blue Waters, Champaign, IL, USA, 2016).
Shaw, D. E. Millisecond-long molecular dynamics simulations of proteins on a special-purpose machine. Biophys. J. 104, 45a (2013).
Gray, A. et al. In pursuit of an accurate spatial and temporal models of biomolecules at the atomistic level: a perspective on computer simulation. Acta Crystallogr. D Biol Crystallogr. 71, 162–172 (2015).
The Royal Swedish Academy of Sciences. Scientific background on the Nobel Prize in Chemistry 2013. Development of multiscale models for complex chemical systems (The Nobel Prize, 2013).
Danev, R. & Baumeister, W. Expanding the boundaries of cryo-EM with phase plates. Curr. Opin. Struct. Biol. 46, 87–94 (2017).
Denk, W. & Horstmann, H. Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biol. 2, e329 (2004).
Villa, E., Schaffer, M., Plitzko, J. M. & Baumeister, W. Opening windows into the cell: Focused-ion-beam milling for cryo-electron tomography. Curr. Opin. Struct. Biol. 23, 771–777 (2013).
Mahamid, J. et al. Visualizing the molecular sociology at the HeLa cell nuclear periphery. Science 351, 969–972 (2016).
Plitzko, M., Schuler, B. & Selenko, P. Structural Biology outside the box — inside the cell. Curr. Opin. Struct. Biol. 46, 110–121 (2017).
Larabell, C. A. & Nugent, K. A. Imaging cellular architecture with X-rays. Curr. Opin. Struct. Biol. 20, 623–631 (2010).
Wall, M. E., Adams, P. D., Fraser, J. S. & Sauter, N. K. Diffuse x-ray scattering to model protein motions. Structure 22, 182–184 (2014).
Neutze, R., Branden, G. & Schertler, G. F. X. Membrane protein structural biology using X-ray free electron lasers. Curr. Opin. Struct. Biol. 33, 115–125 (2015).
Levantino, M., Yorke, B. A., Monteiro, D. C. F., Cammarata, M. & Pearson, A. R. Using synchrotrons and XFELs for time-resolved X-ray crystallography and solution scattering experiments on biomolecules. Curr. Opin. Struct. Biol. 35, 41–48 (2015).
Casadei, C. M. et al. Neutron cryo-crystallography captures the protonation state of ferryl heme in a peroxidase. Science 345, 193–197 (2014).
Shah, N. H. & Muir, T. W. Inteins: nature's gift to protein chemists. Chem. Sci. 5, 446–461 (2014).
Konermann, L., Pan, J. & Liu, Y.-H. Hydrogen exchange mass spectrometry for studying protein structure and dynamics. Chem. Soc. Rev. 40, 1224–1234 (2011).
The National Strategic Computing Initiative Executive Council. National Strategic Computing Initiative Strategic Plan (NSCI, 2016).
Pérez, F. & Granger, B. E. IPython: a system for interactive scientific computing. Comput. Sci. Eng. 9, 21–29 (2007).
Ieong, P. U. et al. Progress towards automated Kepler scientific workflows for computer-aided drug discovery and molecular simulations. Procedia Comput. Sci. 29, 1745–1755 (2014).
Purawat, S. et al. A Kepler workflow tool for reproducible AMBER GPU molecular dynamics. Biophys. J. 112, 2469–2474 (2017).
Gathiaka, S. et al. D3R grand challenge 2015: evaluation of protein-ligand pose and affinity predictions. J. Comput. Aided. Mol. Des. 30, 651–668 (2016).
Gaieb, Z. et al. D3R grand challenge 2: blind prediction of protein–ligand poses, affinity rankings, and relative binding free energies. J. Comput. Aided Mol. Des. 32, 1–20 (2018).
Patwardhan, A. et al. Building bridges between cellular and molecular structural biology. eLife 6, e25835 (2017).
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Woods, C. J. & Mulholland, A. J. Multiscale modelling of biological systems. Chem. Model. 5, 13–50 (2008).
Votapka, L. W. & Amaro, R. E. Multiscale estimation of binding kinetics using Brownian dynamics, molecular dynamics and milestoning. PLoS Comput. Biol. 11, e1004381 (2015).
Chudyk, E. I. et al. QM/MM simulations as an assay for carbapenemase activity in class A β-lactamases. Chem. Commun. 50, 14736–14739 (2014).
Boras, B. W. et al. Bridging scales through multiscale modeling: A case study on protein kinase A. Front. Physiol. 6, 250 (2015).
Mih, N., Brunk, E., Bordbar, A. & Palsson, B. O. A. Multi-scale computational platform to mechanistically assess the effect of genetic variation on drug responses in human erythrocyte metabolism. PLoS Comput. Biol. 12, e1005039 (2016).
Yu, J. S. & Bagheri, N. Multi-class and multi-scale models of complex biological phenomena. Curr. Opin. Biotechnol. 39, 167–173 (2016).
Zhou, G., Pantelopulos, G. A., Mukherjee, S. & Voelz, V. A. Bridging microscopic and macroscopic mechanisms of p53-MDM2 binding with kinetic network models. Biophys. J. 113, 785–793 (2017).
Muzic, M. Le, Autin, L., Parulek, J. & Viola, I. in VCBM 15: Eurographics Workshop on Visual Computing for Biology and Medicinehttps://dx.doi.org/10.2312/vcbm.20151209 (Chester, UK, 2015).
Kerr, R. A. et al. Fast Monte Carlo simulation methods for biological reaction-diffusion systems in solution and on surfaces. SIAM J.Sci. Comput. 30, 3126–3149 (2009).
Hake, J., Kekenes-Huskey, P. M. & McCulloch, A. D. Computational modeling of subcellular transport and signaling. Curr. Opin. Struct. Biol. 25, 92–97 (2014).
Roberts, E. Cellular and molecular structure as a unifying framework for whole-cell modeling. Curr. Opin. Struct. Biol. 25, 86–91 (2014).
Roberts, E., Stone, J. E. & Luthey-Schulten, Z. Lattice microbes: High-performance stochastic simulation method for the reaction-diffusion master equation. J. Comput. Chem. 34, 245–255 (2013).
Earnest, T. M. et al. Challenges of integrating stochastic dynamics and cryo-electron tomograms in whole-cell simulations. J. Phys. Chem. B 121, 3871–3881 (2017).
Noe, F., Biedermann, J., Ullrich, A. & Scho, J. ReaDDyMM: fast interacting particle reaction-diffusion simulations using graphical processing units. Biophys. J. 108, 457–461 (2015).
Chiricotto, M., Sterpone, F., Derreumaux, P. & Melchionna, S. Multiscale simulation of molecular processes in cellular environments. Phil. Trans. R. Soc. A Math. Phys. Eng. Sci. 374, 20160225 (2016).
Tozzini, V. Coarse-grained models for proteins. Curr. Opin. Struct. Biol. 15, 144–150 (2005).
Oliver, R., Read, D. J., Harlen, O. G. & Harris, S. A. A. Stochastic finite element model for the dynamics of globular macromolecules. J. Comput. Phys. 239, 147–165 (2013).
McGuffee, S. R. & Elcock, A. H. Diffusion, crowding and protein stability in a dynamic molecular model of the bacterial cytoplasm. PLoS Comput. Biol. 6, e1000694 (2010).
Freddolino, P. L., Arkhipov, A. S., Larson, S. B., McPherson, A. & Schulten, K. Molecular dynamics simulations of the complete satellite tobacco mosaic virus. Structure 14, 437–449 (2006).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Ou, H. D. et al. ChromEMT: visualizing 3D chromatin structure and compaction in interphase and mitotic cells. Science 357, eaag0025 (2017).
Go, H., Dans, P. D. & Orozco, M. Multiscale simulation of DNA. Curr. Opin. Struct. Biol. 37, 29–45 (2016).
Skolnick, J. Perspective: On the importance of hydrodynamic interactions in the subcellular dynamics of macromolecules. J. Chem. Phys. 145, 100901 (2016).
Trovato, F. & Fumagalli, G. Molecular simulations of cellular processes. Biophys. Rev. 9, 941–958 (2017).
Rydzewski, J. & Nowak, W. Machine learning based dimensionality reduction facilitates ligand diffusion paths assessment: a case of cytochrome P450cam. J. Chem. Theory Comput. 12, 2110–2120 (2016).
Stone, E. Chemistry to go boldly into virtual world. Chem. World (2017).
Russel, D. et al. Putting the pieces together: integrative modeling platform software for structure determination of macromolecular assemblies. PLoS Biol. 10, e1001244 (2012).
Villa, E. & Lasker, K. Finding the right fit: Chiseling structures out of cryo-electron microscopy maps. Curr. Opin. Struct. Biol. 25, 118–125 (2014).
Demir, O., Ieong, P. U. & Amaro, R. E. Full-length p53 tetramer bound to DNA and its quaternary dynamics. Oncogene 36, 1451–1460 (2017).
The Royal Swedish Academy of Sciences. Scientific background on the Nobel Prize in Chemistry 2017. The development of cryo-electron microscopy (The Nobel Prize, 2017).
Willis, J. R. et al. Long antibody HCDR3s from HIV-naïve donors presented on a PG9 neutralizing antibody background mediate HIV neutralization. Proc. Natl Acad. Sci. USA 113, 4446–4451 (2016).
Bale, J. B. et al. Accurate design of megadalton-scale two-component icosahedral protein complexes. Science 353, 389–394 (2016).
Fleishman, S. J. et al. Computational design of proteins targeting the conserved stem region of influenza hemagglutinin. Science 332, 816–821 (2011).
Hsia, Y. et al. Design of a hyperstable 60-subunit protein icosahedron. Nature 535, 136–139 (2016).
Jansen, J. M. et al. Inhibition of prenylated KRAS in a lipid environment. PLOS ONE 12, e0174706 (2017).
Mouchlis, V. D., Bucher, D., McCammon, J. A. & Dennis, E. A. Membranes serve as allosteric activators of phospholipase A2, enabling it to extract, bind, and hydrolyze phospholipid substrates. Proc. Natl Acad. Sci. USA 112, E516–E525 (2015).
Hocker, H. J. et al. Andrographolide derivatives inhibit guanine nucleotide exchange and abrogate oncogenic Ras function. Proc. Natl Acad. Sci. USA 110, 10201–10206 (2013).
Singhal, N., Snow, C. D. & Pande, V. S. Using path sampling to build better Markovian state models: predicting the folding rate and mechanism of a tryptophan zipper beta hairpin. J. Chem. Phys. 121, 415–425 (2004).
Swope, W. C. et al. Describing protein folding kinetics by molecular dynamics simulations. 2. Example applications to alanine dipeptide and a beta-hairpin peptide. J. Phys. Chem. B 108, 6582–6594 (2004).
Swope, W. C., Pitera, J. W. & Suits, F. Describing protein folding kinetics by molecular dynamics simulations. 1. Theory. J. Phys. Chem. B 108, 6571–6581 (2004).
Meng, Y., Shukla, D., Pande, V. S. & Roux, B. Transition path theory analysis of c-Src kinase activation. Proc. Natl Acad. Sci. USA 113, 9193–9198 (2016).
Malmstrom, R. D., Kornev, A. P., Taylor, S. S. & Amaro, R. E. Allostery through the computational microscope: cAMP activation of a canonical signalling domain. Nat. Commun. 6, 7588 (2015).
Plattner, N., Doerr, S., De Fabritiis, G. & Noé, F. Complete protein–protein association kinetics in atomic detail revealed by molecular dynamics simulations and Markov modelling. Nat. Chem. 9, 1005–1011 (2017).
Kohlhoff, K. J. et al. Cloud-based simulations on Google Exacycle reveal ligand modulation of GPCR activation pathways. Nat. Chem. 6, 15–21 (2013).
Folmer, R. H. A. Drug target residence time: a misleading concept. Drug Discov. Today 23, 12–16 (2018).
Schuetz, D. A. et al. Kinetics for Drug Discovery: an industry-driven effort to target drug residence time. Drug Discov. Today 22, 896–911 (2017).
Zheng, W., Gallicchio, E., Deng, N., Andrec, M. & Levy, R. M. Kinetic network study of the diversity and temperature dependence of Trp-Cage folding pathways: Combining transition path theory with stochastic simulations. J. Phys. Chem. B 115, 1512–1523 (2011).
Plattner, N. & Noé, F. Protein conformational plasticity and complex ligand-binding kinetics explored by atomistic simulations and Markov models. Nat. Commun. 6, 7653 (2015).
Segala, E. et al. Controlling the dissociation of ligands from the adenosine A. J. Med. Chem. 59, 7167–7176 (2016).
Mollica, L. et al. Kinetics of protein-ligand unbinding via smoothed potential molecular dynamics simulations. Sci. Rep. 5, 11539 (2015).
Tiwary, P., Limongelli, V., Salvalaglio, M. & Parrinello, M. Kinetics of protein-ligand unbinding: predicting pathways, rates, and rate-limiting steps. Proc. Natl Acad. Sci. USA 112, E386–E391 (2015).
Juraszek, J., Saladino, G., Van Erp, T. S. & Gervasio, F. L. Efficient numerical reconstruction of protein folding kinetics with partial path sampling and pathlike variables. Phys. Rev. Lett. 110, 108106 (2013).
Votapka, L. W., Jagger, B. R., Heyneman, A. L. & Amaro, R. E. SEEKR: simulation enabled estimation of kinetic rates, a computational tool to estimate molecular kinetics and its application to trypsin-benzamidine binding. J. Phys. Chem. B 121, 3597–3606 (2017).
Faradjian, A. K. & Elber, R. Computing time scales from reaction coordinates by milestoning. J. Chem. Phys. 120, 10880–10889 (2004).
Swift, R. V. & Amaro, R. E. Back to the Future: Can physical models of passive membrane permeability help reduce drug candidate attrition and move us beyond QSPR? Chem. Biol. Drug Des. 81, 61–71 (2013).
Lee, C. T. et al. Simulation-based approaches for determining membrane permeability of small compounds. J. Chem. Inf. Model. 56, 721–733 (2016).
Parkin, J., Chavent, M. & Khalid, S. Molecular simulations of Gram-negative bacterial membranes: a vignette of some recent successes. Biophys. J. 109, 461–468 (2015).
Berglund, N. A. et al. Interaction of the antimicrobial peptide polymyxin B1 with both membranes of E. coli : a molecular dynamics study. PLoS Comput. Biol. 11, e1004180 (2015).
Pavlova, A., Hwang, H., Lundquist, K., Balusek, C. & Gumbart, J. Living on the edge: Simulations of bacterial outer-membrane proteins. Biochim. Biophys. Acta 1858, 1753–1759 (2016).
Orsi, M., Noro, M. G. & Essex, J. W. Dual-resolution molecular dynamics simulation of antimicrobials in biomembranes. J. R. Soc. Interface 8, 826–841 (2011).
Dickson, C. J., Hornak, V., Pearlstein, R. A. & Duca, J. S. Structure–kinetic relationships of passive membrane permeation from multiscale modeling. J. Am. Chem. Soc. 139, 442–452 (2017).
Cardenas, A. E. et al. Unassisted transport of N-acetyl-l-tryptophanamide through membrane: experiment and simulation of kinetics. J. Phys. Chem. B 116, 2739–2750 (2012).
Votapka, L. W., Lee, C. T. & Amaro, R. E. Two relations to estimate membrane permeability using milestoning. J. Phys. Chem. B 120, 8606–8616 (2016).
Wu, H., Paul, F., Wehmeyer, C. & Noé, F. Multiensemble Markov models of molecular thermodynamics and kinetics. Proc. Natl Acad. Sci. USA 114, E3221–E3230 (2017).
Boninsegna, L. et al. A data-driven perspective on the hierarchical assembly of molecular structures. J. Chem. Theory Comput. 14, 453–460 (2018).
Dama, J. F. et al. The theory of ultra-coarse-graining. 1. General principles. J. Chem. Theory Comput. 9, 2466–2480 (2013).
Kmiecik, S. et al. Coarse-grained protein models and their applications. Chem. Rev. 116, 7898–7936 (2016).
Graham, J. A., Essex, J. W. & Khalid, S. PyCGTOOL: automated generation of coarse-grained molecular dynamics models from atomistic trajectories. J. Chem. Inf. Model. 57, 650–656 (2017).
Sharma, S. et al. A coarse-grained protein model in a water-like solvent. Sci. Rep. 3, 1841 (2013).
Altae-tran, H., Ramsundar, B., Pappu, A. S. & Pande, V. Low data drug discovery with one-shot learning. ACS Cent. Sci. 3, 283–293 (2017).
Cui, Q. Perspective: Quantum mechanical methods in biochemistry and biophysics. J. Chem. Phys. 145, 140901 (2016).
Ranaghan, K. E. et al. A catalytic role for methionine revealed by a combination of computation and experiments on phosphite dehydrogenase. Chem. Sci. 5, 2191–2199 (2014).
Lodola, A. et al. Identification of productive inhibitor binding orientation in fatty acid amide hydrolase (FAAH) by QM/MM mechanistic modelling. Chem. Commun., 214–216 (2008).
Lonsdale, R., Rouse, S. L., Sansom, M. S. P. & Mulholland, A. J. A. Multiscale approach to modelling drug metabolism by membrane-bound cytochrome P450 enzymes. PLoS Comput. Biol. 10, e1003714 (2014).
Lonsdale, R. et al. Quantum mechanics/molecular mechanics modeling of regioselectivity of drug metabolism in cytochrome P450 2C9. J. Am. Chem. Soc. 135, 8001–8015 (2013).
Tyzack, J. D., Hunt, P. A. & Segall, M. D. Predicting regioselectivity and lability of cytochrome P450 metabolism using quantum mechanical simulations. J. Chem. Inf. Model. 56, 2180–2193 (2016).
Goodpaster, J. D., Barnes, T. A., Manby, F. R. & Miller, T. F. Accurate and systematically improvable density functional theory embedding for correlated wavefunctions. J. Chem. Phys. 140, 18A507 (2014).
Barnes, T. A., Goodpaster, J. D., Manby, F. R. & Miller, T. F. Accurate basis set truncation for wavefunction embedding. J. Chem. Phys. 139, 024103 (2013).
Zhang, X. et al. Multiscale analysis of enantioselectivity in enzyme-catalysed ‘lethal synthesis’ using projector-based embedding. R. Soc. Open Sci. 5, 171390 (2018).
Ryde, U. & Soderhjelm, P. Ligand-binding affinity estimates supported by quantum-mechanical methods. Chem. Rev. 116, 5520–5566 (2016).
Woods, C. J., Shaw, K. E. & Mulholland, A. J. Combined quantum mechanics/molecular mechanics (QM/MM) simulations for protein-ligand complexes: free energies of binding of water molecules in influenza neuraminidase. J. Phys. Chem. B 119, 997–1001 (2015).
Genheden, S., Cabedo Martinez, A. I., Criddle, M. P. & Essex, J. W. Extensive all-atom Monte Carlo sampling and QM/MM corrections in the SAMPL4 hydration free energy challenge. J. Comput. Aided. Mol. Des. 28, 187–200 (2014).
Konig, G., Pickard IV, F. C., Mei, Y. & Brooks, B. R. Predicting hydration free energies with a hybrid QM/MM approach: an evaluation of implicit and explicit solvation models in SAMPL4. J. Comput. Aided. Mol. Des. 28, 245–257 (2014).
Ko, G. et al. Computation of hydration free energies using the multiple environment single system quantum mechanical / molecular mechanical method. J. Chem. Theory Comput. 12, 332–344 (2016).
Ho, M. H., De Vivo, M., Peraro, M. D. & Klein, M. L. Unraveling the catalytic pathway of metalloenzyme farnesyltransferase through QM/MM computation. J. Chem. Theory Comput. 5, 1657–1666 (2009).
Woods, C. J., Shaw, K. E., M. A. Combined quantum mechanics/molecular mechanics (QM/MM) simulations for protein-ligand complexes: free energies of binding of water molecules in influenza neuraminidase. J. Phys. Chem. B 119, 997–1001 (2015).
Smith, J. S., Isayev, O. & Roitberg, A. E. ANI-1: an extensible neural network potential with DFT accuracy at force field computational cost. Chem. Sci. 8, 3192–3203 (2017).
Doemer, M., Maurer, P., Campomanes, P., Tavernelli, I. & Rothlisberger, U. Generalized QM/MM force matching approach applied to the 11-cis protonated Schiff base chromophore of rhodopsin. J. Chem. Theory Comput. 10, 412–422 (2014).
Singh, J., Petter, R. C., Baillie, T. A. & Whitty, A. The resurgence of covalent drugs. Nat. Rev. Drug Discov. 10, 307–317 (2011).
Callegari, D. et al. L718Q mutant EGFR escapes covalent inhibition by stabilizing a non-reactive conformation of the lung cancer drug osimertinib. Chem. Sci. 9, 2740–2749 (2018).
Kamerlin, S. C. L. & Warshel, A. The empirical valence bond model: theory and applications. Wiley Interdiscip. Rev. Comput. Mol. Sci. 1, 30–45 (2011).
Kazemi, M., Himo, F. & Aqvist, J. Enzyme catalysis by entropy without Circe effect. Proc. Natl Acad. Sci. USA 113, 2406–2411 (2016).
Ryde, U. QM/MM calculations on proteins. Methods Enzymol. 577, 119–158 (2016).
Karelina, M. & Kulik, H. J. Systematic quantum mechanical region determination in QM/MM simulation. J. Chem. Theory Comput. 13, 563–576 (2017).
Mones, L. et al. The adaptive buffered force QM/MM method in the CP2K and amber software packages. J. Comput. Chem. 36, 633–648 (2015).
Sokkar, P., Boulanger, E., Thiel, W. & Sanchez-Garcia, E. Hybrid quantum mechanics/molecular mechanics/coarse grained modeling: a triple-resolution approach for biomolecular systems. J. Chem. Theory Comput. 11, 1809–1818 (2015).
Weidle, U. H., Tiefenthaler, G. & Georges, G. Proteases as activators for cytotoxic prodrugs in antitumor therapy. Cancer Genom. Proteom. 11, 67–80 (2014).
Schellmann, N., Deckert, P. M., Bachran, D., Fuchs, H. & Bachran, C. Targeted enzyme prodrug therapies. Mini Rev. Med. Chem. 10, 887–904 (2010).
Bown, S. G. Photodynamic therapy for photochemists. Phil. Trans. R. Soc. A Math. Phys. Eng. Sci. 371, 20120371 (2013).
Davis, B. et al. Selective protein degradation by ligand-targeted enzymes: towards the creation of catalytic antagonists. Chembiochem 4, 533–537 (2003).
Johnson, G. T. et al. cellPACK: a virtual mesoscope to model and visualize structural systems biology. Nat. Methods 12, 85–91 (2014).
Durrant, J. D. & Amaro, R. E. LipidWrapper: an algorithm for generating large-scale membrane models of arbitrary geometry. PLoS Comput. Biol. 10, e1003720 (2014).
Lomize, M. A., Lomize, A. L., Pogozheva, I. D. & Mosberg, H. I. OPM: orientations of proteins in membranes database. Bioinformatics 22, 623–625 (2006).
Bennie, S. J. et al. A projector-embedding approach for multiscale coupled-cluster calculations applied to citrate synthase. J. Chem. Theory Comput. 12, 2689–2697 (2016).
Cavalli, A., Spitaleri, A., Saladino, G. & Gervasio, F. L. Investigating drug-target association and dissociation mechanisms using metadynamics-based algorithms. Acc. Chem. Res. 48, 277–285 (2015).
Melo, M. C. R. et al. NAMD goes quantum: an integrative suite for hybrid simulations. Nat. Methods https://doi.org/10.1038/nmeth.4638 (2018).
Fowler, P. W. et al. Robust prediction of resistance to trimethoprim in Staphylococcus aureus. Cell Chem. Biol. 25, 339–349 (2018).
Callegari, D. et al. Metadynamics simulations distinguish short- and long-residence-time inhibitors of cylcin-dependent kinase 8. J. Chem. Inf. Model. 57, 159–169 (2017).
Acknowledgements
Support from NIH DP2 OD007237 and NIH P41 GM103426 to R.E.A. is gratefully acknowledged. R.E.A. thanks Pek Ieong, Ludovic Autin and Benjamin Jagger for assistance in figure preparation. A.J.M. thanks EPSRC for support (EP/M022609/1 (for CCP-BioSim see www.ccpbiosim.ac.uk) and EP/M015378/1) and BBSRC (BB/M000354/1). A.J.M. thanks co-workers, including Christopher Woods, Christine Bathelt, Sarah Rouse, Richard Lonsdale and Mark Sansom, for help in the preparation of the figures.
Author information
Authors and Affiliations
Contributions
All authors contributed equally to the preparation of this manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.E.A. is a co-founder, a Scientific Advisory Board member and has equity interest in Actavalon, Inc. A.J.M has no competing interests.
PowerPoint slides
Rights and permissions
About this article
Cite this article
Amaro, R., Mulholland, A. Multiscale methods in drug design bridge chemical and biological complexity in the search for cures. Nat Rev Chem 2, 0148 (2018). https://doi.org/10.1038/s41570-018-0148
Published:
Version of record:
DOI: https://doi.org/10.1038/s41570-018-0148
This article is cited by
-
Morphological and phase behaviors of symmetric diblock copolymers: insights from coarse-grained molecular dynamics simulations
npj Soft Matter (2025)
-
Cannabis sativa Phytochemicals as Potent Regulators of Aquaporin-4 in Alzheimer’s Disease: A Cheminformatics and Machine Learning Approach
Chemistry Africa (2025)
-
SQM2.20: Semiempirical quantum-mechanical scoring function yields DFT-quality protein–ligand binding affinity predictions in minutes
Nature Communications (2024)
-
Drug design on quantum computers
Nature Physics (2024)
-
Extrapolative prediction of small-data molecular property using quantum mechanics-assisted machine learning
npj Computational Materials (2024)