Abstract
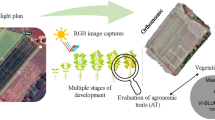
Soybean growth is determined by the interaction of genetic, environmental, and management factors. In the context of future climate and climate extremes, understanding genotype by environment interaction (GxE) will be crucial for selecting resilient breeding lines and optimizing management practices to minimize stress. This requires an in depth elucidation of stressful weather conditions and differing temporal responses of genotypes to those conditions. In field studies, however, the environment is often treated as a static factor, and the specific effects of weather variability on crop growth remain poorly understood. Here, we present a longitudinal dataset comprising 17,247 high-resolution RGB images of soybean breeding lines collected throughout eight years in Eschikon, Switzerland. Top-of-canopy images were acquired throughout the entire growing seasons and complemented by hourly weather data, enabling a comprehensive analysis of soybean growth dynamics under varying field conditions. High spatio-temporal image resolution allows detailed analysis of growth dynamics and GxE, supporting identification of stress-tolerant genotypes to improve yield prediction and yield stability.
Similar content being viewed by others
Code availability
The code is available on: https://gitlab.ethz.ch/crop_phenotyping/fip-soybean-canopycover. Users with similar data can use the implemented workflow to get canopy cover from their experiments.
References
Board, J. E., Kahlon, C. S., Board, J. E. & Kahlon, C. S. Soybean Yield Formation: What Controls It and How It Can Be Improved. In Soybean Physiology and Biochemistry, https://doi.org/10.5772/17596, https://www.intechopen.com/chapters/22761 (IntechOpen, 2011).
Sau, F., Boote, K. J. & Ruíz-Nogueira, B. Evaluation and improvement of CROPGRO-soybean model for a cool environment in Galicia, northwest Spain. Field Crops Research 61, 273–291, https://doi.org/10.1016/S0378-4290(98)00168-3 (1999).
Crestani Mota, M. et al. CROPGRO-soybean model – Validation and application for the southern Amazon, Brazil. Computers and Electronics in Agriculture 216, 108478, https://doi.org/10.1016/j.compag.2023.108478 (2024).
Wu, Y. et al. Combine observational data and modelling to quantify cultivar differences of soybean. European Journal of Agronomy 111, 125940, https://doi.org/10.1016/j.eja.2019.125940 (2019).
Balboa, G. R. et al. A systems-level yield gap assessment of maize-soybean rotation under high- and low-management inputs in the Western US Corn Belt using APSIM. Agricultural Systems 174, 145–154, https://doi.org/10.1016/j.agsy.2019.04.008 (2019).
Battisti, R., Sentelhas, P. C. & Boote, K. J. Inter-comparison of performance of soybean crop simulation models and their ensemble in southern Brazil. Field Crops Research 200, 28–37, https://doi.org/10.1016/j.fcr.2016.10.004 (2017).
Kothari, K. et al. Evaluating differences among crop models in simulating soybean in-season growth. Field Crops Research 309, 109306, https://doi.org/10.1016/j.fcr.2024.109306 (2024).
Xavier, A., Hall, B., Hearst, A. A., Cherkauer, K. A. & Rainey, K. M. Genetic Architecture of Phenomic-Enabled Canopy Coverage in Glycine max. Genetics 206, 1081–1089, https://doi.org/10.1534/genetics.116.198713 (2017).
Roth, L., Aasen, H., Walter, A. & Liebisch, F. Extracting leaf area index using viewing geometry effects—A new perspective on high-resolution unmanned aerial system photography. ISPRS Journal of Photogrammetry and Remote Sensing 141, 161–175, https://doi.org/10.1016/j.isprsjprs.2018.04.012 (2018).
Aasen, H., Kirchgessner, N., Walter, A. & Liebisch, F. PhenoCams for Field Phenotyping: Using Very High Temporal Resolution Digital Repeated Photography to Investigate Interactions of Growth, Phenology, and Harvest Traits. Frontiers in Plant Science 11, 593, https://doi.org/10.3389/fpls.2020.00593 (2020).
Gill, T. et al. A Comprehensive Review of High Throughput Phenotyping and Machine Learning for Plant Stress Phenotyping. Phenomics 2, 156–183, https://doi.org/10.1007/s43657-022-00048-z (2022).
Xu, R. & Li, C. A Review of High-Throughput Field Phenotyping Systems: Focusing on Ground Robots. Plant Phenomics 2022, https://doi.org/10.34133/2022/9760269 (2022).
Rui, Z. et al. High-throughput proximal ground crop phenotyping systems – A comprehensive review. Computers and Electronics in Agriculture 224, 109108, https://doi.org/10.1016/j.compag.2024.109108 (2024).
Kirchgessner, N. et al. The ETH field phenotyping platform FIP: a cable-suspended multi-sensor system. Functional Plant Biology 44, 154–168, https://doi.org/10.1071/FP16165 (2017).
Roth, L. et al. From Neglecting to Including Cultivar-Specific Per Se Temperature Responses: Extending the Concept of Thermal Time in Field Crops. Plant Phenomics 6, 0185, https://doi.org/10.34133/plantphenomics.0185 (2024).
Roth, L. et al. The FIP 1.0 Data Set: Highly Resolved Annotated Image Time Series of 4,000 Wheat Plots Grown in Six Years, https://doi.org/10.1101/2024.10.04.616624 (2024).
Zenkl, R. et al. Outdoor Plant Segmentation With Deep Learning for High-Throughput Field Phenotyping on a Diverse Wheat Dataset. Frontiers in Plant Science 12 https://doi.org/10.3389/fpls.2021.774068 (2022).
Oppliger, C., Zenkl, R., Walter, A. & Keller, B. Investigating pea (Pisum sativum L.) flowering with high throughput field phenotyping and object detection. Smart Agricultural Technology 11, 100942, https://doi.org/10.1016/j.atech.2025.100942 (2025).
Liu, X. et al. Genetic variation of world soybean maturity date and geographic distribution of maturity groups. Breeding Science 67, 221–232, https://doi.org/10.1270/jsbbs.16167 (2017).
Roth, L., Barendregt, C., Bétrix, C.-A., Hund, A. & Walter, A. High-throughput field phenotyping of soybean: Spotting an ideotype. Remote Sensing of Environment 269, 112797, https://doi.org/10.1016/j.rse.2021.112797 (2022).
Kronenberg, L., Yu, K., Walter, A. & Hund, A. Monitoring the dynamics of wheat stem elongation: genotypes differ at critical stages. Euphytica 213, 157, https://doi.org/10.1007/s10681-017-1940-2 (2017).
Easlon, H. M. & Bloom, A. J. Easy Leaf Area: Automated digital image analysis for rapid and accurate measurement of leaf area. Applications in Plant Sciences 2, 1400033, https://doi.org/10.3732/apps.1400033 (2014).
Rodríguez-Álvarez, M. X., Boer, M. P., van Eeuwijk, F. A. & Eilers, P. H. Correcting for spatial heterogeneity in plant breeding experiments with P-splines. Spatial Statistics 23, 52–71, https://doi.org/10.1016/j.spasta.2017.10.003 (2018).
Keller, B. et al. Genomic Prediction of Agronomic Traits in Common Bean (Phaseolus vulgaris L.) Under Environmental Stress. Frontiers in Plant Science 11, https://doi.org/10.3389/fpls.2020.01001 (2020).
Aparicio, J. et al. Mr.Bean: a comprehensive statistical and visualization application for modeling agricultural field trials data. Frontiers in Plant Science 14, https://doi.org/10.3389/fpls.2023.1290078 (2024).
Keller, B. et al. Data repository: FIP 1.0 Soybean data set: Insights on soybean growth dynamics from eight years of high-throughput image field phenotyping, https://doi.org/10.3929/ethz-b-000742401 (2025). Geographic location: Eschikon, Lindau, Switzerland (47.449 N, 8.682 E). Data collected 2015–2022.
Boss, M. & Keller, B. FIP1SOY: High-throughput soybean field phenotyping dataset, https://doi.org/10.57967/hf/6052 (2025).
Meek, D. W., Hatfield, J. L., Howell, T. A., Idso, S. B. & Reginato, R. J. A Generalized Relationship between Photosynthetically Active Radiation and Solar Radiation. Agronomy Journal 76, 939–945, https://doi.org/10.2134/agronj1984.00021962007600060018x (1984).
Papoutsoglou, E. A. et al. Enabling reusability of plant phenomic datasets with MIAPPE 1.1. New Phytologist 227, 260–273, https://doi.org/10.1111/nph.16544 (2020).
Wood, S. N.Generalized Additive Models: An Introduction with R, Second Edition (Chapman and Hall/CRC, New York, 2017), 2 edn. https://doi.org/10.1201/9781315370279
Mourtzinis, S., Gaspar, A. P., Naeve, S. L. & Conley, S. P. Planting Date, Maturity, and Temperature Effects on Soybean Seed Yield and Composition. Agronomy Journal 109, 2040–2049, https://doi.org/10.2134/agronj2017.05.0247 (2017).
Salmerón, M. et al. Regional analysis of planting date and cultivar maturity recommendations that improve soybean oil yield and meal protein concentration. Frontiers in Plant Science 13, https://doi.org/10.3389/fpls.2022.954111 (2022).
Anderegg, J., Zenkl, R., Walter, A., Hund, A. & McDonald, B. A. Combining High-Resolution Imaging, Deep Learning, and Dynamic Modeling to Separate Disease and Senescence in Wheat Canopies. Plant Phenomics 5, 0053, https://doi.org/10.34133/plantphenomics.0053 (2023).
Zhao, J. et al. Improved Field-Based Soybean Seed Counting and Localization with Feature Level Considered. Plant Phenomics 5, 0026, https://doi.org/10.34133/plantphenomics.0026 (2023).
Zhou, S., Sun, Q., Zhang, N., Chai, X. & Sun, T. PodNet: Pod real-time instance segmentation in pre-harvest soybean fields. Plant Phenomics 7, 100052, https://doi.org/10.1016/j.plaphe.2025.100052 (2025).
Acknowledgements
We thank Agroscope Soybean Breeding and Delley seeds and plants Ltd. (DSP) for providing the seeds. We would also like to thank Brigitta Herzog for seed preparation and inoculation. Special thanks go to Mike Boss for making the data set available on hugging face. We would like to thank the entire Crop Science Group at ETH Zürich for their support in and out of the field and for their engagement in discussions about the manuscript. Large language model (ChatGPT/Claude) were used in part to assist with coding and phrasing. All outputs were checked, verified and finalized by the authors. This work was supported by the Swiss National Science Foundation (Grant Nos. 200756 and 169542).
Author information
Authors and Affiliations
Contributions
B.K.: Developed algorithm, analyzed data and drafted manuscript; N.K., L.R., A.H., A.M.: FIP development, B.K., N.K., C.O., L.K., L.R., O.Z., S.C., F.L., H.A., N.S., F.T., H.Z., C.A.B., C.B., A.H.: Collected and prepared data; Experimental design: B.K., L.K., L.R., A.H.; all authors improved and approved the manuscript
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Keller, B., Kirchgessner, N., Oppliger, C. et al. FIP 1.0 soybean data: Insights on soybean growth from eight years of high-throughput image field phenotyping. Sci Data (2026). https://doi.org/10.1038/s41597-026-06663-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-06663-z