Abstract
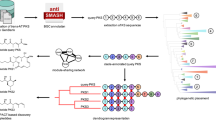
Freshwater dinoflagellates are typically considered non-toxic, and their polyketides and toxin biosynthesis genes are largely unexplored. Here, we generated and analyzed the transcriptome of freshwater dinoflagellate Palatinus apiculatus and compared it with the transcriptome data from Peridinium bipes and Ceratium furcoides to investigate the presence and diversity of polyketide synthases (PKS), fatty acid synthases (FAS), and saxitoxin (STX) biosynthesis genes (sxt). We identified 95, 117, and 39 PKS-related transcripts in P. apiculatus, P. bipes, and C. furcoides, respectively, which include single-domain PKS, multi-domain PKS, and hybrid NRPS/PKS. Phylogenetic analysis revealed a novel clade of ketosynthase (KS) domains unique to freshwater dinoflagellates, suggesting species-specific diversification. Conserved catalytic residues were found in type II FAS genes across both freshwater and marine taxa. Although core STX biosynthesis genes were absent in all analyzed species, several STX-associated transcripts, including sxtA4, sxtU, sxtS, sxtD, sxtH/T, and sxtI, were identified. Phylogenetic analysis of the sxtA4 domain revealed that freshwater dinoflagellate sequences form a distinct clade from those of toxic marine dinoflagellates and cyanobacteria while retaining conserved active sites, suggesting potential functional variation. These findings reveal unique PKS and STX gene features in freshwater dinoflagellates, highlighting their previously unrecognized biosynthetic diversity, ecological roles, and biotechnological potential.
Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are available in the NCBI Sequence Read Archive (SRA) under Bio Project, PRJNA1307768.
References
Bi, R. et al. Responses of marine diatom–dinoflagellate competition to multiple environmental drivers: Abundance, elemental, and biochemical aspects. Front. Microbiol. 12, 731786 (2021).
Taylor, F. J. R., Hoppenrath, M. & Saldarriaga, J. F. Dinoflagellate diversity and distribution. Biodivers. Conserv. 17, 407–418 (2008).
Beedessee, G. et al. Diversified secondary metabolite biosynthesis gene repertoire revealed in symbiotic dinoflagellates. Sci. Rep. 9, 1204 (2019).
Beedessee, G. et al. Integrated omics unveil the secondary metabolic landscape of a basal dinoflagellate. BMC Biol. 18, 16 (2020).
Camacho-Muñoz, D., Praptiwi, R. A., Lawton, L. A. & Edwards, C. High value Phycotoxins from the dinoflagellate Prorocentrum. Front. Mar. Sci. 8, 638739 (2021).
Verma, A. et al. The genetic basis of toxin biosynthesis in dinoflagellates. Microorganisms 7, 222 (2019).
Rein, K. S. & Borrone, J. Polyketides from dinoflagellates: Origins, Pharmacology and biosynthesis. Comp. Biochem. Physiol. B Biochem. Mol. Biol.. 124, 117–131 (1999).
Kellmann, R., Stüken, A., Orr, R. J., Svendsen, H. M. & Jakobsen, K. S. Biosynthesis and molecular genetics of polyketides in marine dinoflagellates. Mar. Drugs. 8, 1011–1048 (2010).
Pawlowiez, R., Morey, J. S., Darius, H. T., Chinain, M. & Van Dolah, F. M. Transcriptome sequencing reveals single domain type I-like polyketide synthases in the toxic dinoflagellate Gambierdiscus polynesiensis. Harmful Algae. 36, 29–37 (2014).
Kohli, G. S. et al. Polyketide synthesis genes associated with toxin production in two species of Gambierdiscus (Dinophyceae). BMC Genom. 16, 410 (2015).
Van Dolah, F. M. et al. Transcriptomic analysis of polyketide synthases in a highly ciguatoxic dinoflagellate, Gambierdiscus polynesiensis and low toxicity Gambierdiscus pacificus, from French Polynesia. PLoS ONE. 15, e0231400 (2020).
Wan, X. et al. Transcriptomic analysis of polyketide synthesis in dinoflagellate, prorocentrum Lima. Harmful Algae. 123, 102391 (2023).
Snyder, R. V. et al. Polyketide synthase genes from marine dinoflagellates. Mar. Biotechnol. 5, 1–12 (2003).
Kohli, G. S., John, U., Van Dolah, F. M. & Murray, S. A. Evolutionary distinctiveness of fatty acid and polyketide synthesis in eukaryotes. ISME J. 10, 1877–1890 (2016).
Wang, H., Kim, H. & Ki, J. S. Transcriptome survey and toxin measurements reveal evolutionary modification and loss of saxitoxin biosynthesis genes in the dinoflagellates Amphidinium Carterae and Prorocentrum micans. Ecotoxicol. Environ Saf. 195, 110474 (2020).
Kellmann, R. et al. Biosynthetic intermediate analysis and functional homology reveal a saxitoxin gene cluster in cyanobacteria. Appl. Environ. Microbiol. 74, 4044–4053 (2008).
Stüken, A. et al. Discovery of nuclear-encoded genes for the neurotoxin saxitoxin in dinoflagellates. PLoS ONE. 6, e20096 (2011).
Camacho, F. G. et al. Biotechnological significance of toxic marine dinoflagellates. Biotechnol. Adv. 25, 176–194 (2007).
Assunção, J., Guedes, A. C. & Malcata, F. X. Biotechnological and Pharmacological applications of biotoxins and other bioactive molecules from dinoflagellates. Mar. Drugs. 15, 393 (2017).
Morales-Amador, A., Souto, M. L., Hertweck, C., Fernández, J. J. & García-Altares, M. Rapid screening of polyol polyketides from marine dinoflagellates. Anal. Chem. 94, 14205–14213 (2022).
Pradhan, B. & Ki, J. S. Phytoplankton toxins and their potential therapeutic applications: A journey toward the quest for potent pharmaceuticals. Mar. Drugs. 20, 271 (2022).
Beedessee, G., Hisata, K., Roy, M. C., Satoh, N. & Shoguchi, E. Multifunctional polyketide synthase genes identified by genomic survey of the symbiotic dinoflagellate, Symbiodinium Minutum. BMC Genom. 16, 941 (2015).
Jenke-Kodama, H., Sandmann, A., Müller, R. & Dittmann, E. Evolutionary implications of bacterial polyketide synthases. Mol. Biol. Evol. 22, 2027–2039 (2005).
Shen, B. et al. Prerequisites for combinatorial biosynthesis: evolution of hybrid NRPS/PKS gene clusters. Biocombinatorial Approaches Drug Finding, 107–126 (2005).
Keatinge-Clay, A. T. The structures of type I polyketide synthases. Nat. Prod. Rep. 29, 1050–1073 (2012).
Rasmussen, S. A. et al. Chemical diversity, origin, and analysis of Phycotoxins. J. Nat. Prod. 79, 662–673 (2016).
Meyer, J. M. et al. Transcriptomic characterisation and genomic glimpse into the toxigenic dinoflagellate Azadinium spinosum, with emphasis on polyketide synthase genes. BMC Genom. 16, 27 (2015).
Kohli, G. S. et al. Role of modular polyketide synthases in the production of polyether ladder compounds in ciguatoxin-producing Gambierdiscus polynesiensis and G. excentricus (Dinophyceae). J. Eukaryot. Microbiol. 64, 691–706 (2017).
Van Dolah, F. M., Kohli, G. S., Morey, J. S. & Murray, S. A. Both modular and single-domain type I polyketide synthases are expressed in the brevetoxin‐producing dinoflagellate, Karenia brevis (Dinophyceae). J. Phycol. 53, 1325–1339 (2017).
Wang, H., Guo, R., Lim, W. A., Allen, A. E. & Ki, J. S. Comparative transcriptomics of toxin synthesis genes between the non-toxin producing dinoflagellate Cochlodinium Polykrikoides and toxigenic Alexandrium Pacificum. Harmful Algae. 93, 101777 (2020).
Paguigan, N. D., Raja, H. A., El-Elimat, T. & Oberlies, N. H. New polyketides from a freshwater Lindgomycetaceae Sp. Planta Med. 80, PC33 (2014).
Ebada, S. S. & Ebrahim, W. A new antioxidant decalin polyketide from freshwater-sediment‐derived fungus Penicillium sp. strain S1a1. Chem. Select. 4, 9814–9816 (2019).
Kaluzhnaya, O. V. & Itskovich, V. B. Features of diversity of polyketide synthase genes in the community of freshwater sponge Baikalospongia fungiformis. Russ J. Genet. 58, 336–346 (2022).
Scesa, P. et al. Defensive polyketides produced by an abundant gastropod are candidate keystone molecules in estuarine ecology. Sci. Adv. 10, eadp8643 (2024).
Roset, J. et al. Mortality of rainbow trout (Oncorynchus Mykiss (Walbaum)) associated with freshwater dinoflagellate bloom (Peridinium polonicum (Woloszynska)) in a fish farm. Aquac Res. 33, 159–164 (2002).
Oshima, Y., Minami, H., Takano, Y. & Yasumoto, T. Ichthyotoxins in a freshwater dinoflagellate Peridinium polonicum. In Red Tides: Biology, Environmental Science and Toxicology. Proceedings of the First International Symposium on Red Tides 375–377 (Elsevier, 1989).
Rengefors, K. & Legrand, C. Toxicity in Peridinium aciculiferum—an adaptive strategy to outcompete other winter phytoplankton? Limnol. Oceanogr. 46, 1990–1997 (2001).
Wu, J. T., Kuo-Huang, L. L. & Lee, J. Algicidal effect of Peridinium bipes on Microcystis aeruginosa. Curr. Microbiol. 37, 257–261 (1998).
Krahmalniy, A. F. Mass development of Palatinus apiculatus (Dinoflagellata) in the verbne lake (Kyiv, Ukraine). Hydrobiol. J. 54, 41–47 (2018).
Kim, T. & Ki, J. S. New record of the cold freshwater dinoflagellate Palatinus apiculatus (Dinophyceae) from the Paldang Reservoir, Korea. J. Species Res. 11, 162–168 (2022).
Corrêa, R. F. et al. First report of the invasive Ceratium furcoides (dinoflagellate) in Paracambi Reservoir, Rio de janeiro: risks to the world’s largest domestic water treatment plant. Lakes Reserv. Res. Manag. 27, e12400 (2022).
Lin, S. A decade of dinoflagellate genomics illuminating an enigmatic eukaryote cell. BMC Genom. 25, 932 (2024).
Wang, H. et al. Nuclear genome of dinoflagellates: size variation and insights into evolutionary mechanisms. Eur. J. Protistol.. 93, 126061 (2024).
Van Dolah, F. M. et al. Transcriptomic analysis of polyketide synthases in a highly ciguatoxic dinoflagellate, Gambierdiscus polynesiensis and low toxicity Gambierdiscus pacificus, from French Polynesia. PLoS ONE. 15, e0231400 (2020).
Bui, Q. T. N., Pradhan, B., Kim, H. S. & Ki, J. S. Environmental factors modulate saxitoxins (STXs) production in toxic dinoflagellate Alexandrium: an updated review of STXs and synthesis gene aspects. Toxins 16, 210 (2024).
Muhammad, B. L., Kim, H. S., Bui, Q. T. N. & Ki, J. S. Transcriptomic comparison unveils saxitoxin biosynthesis genes in the marine dinoflagellate Gymnodinium catenatum. Harmful Algae. 137, 102872 (2025).
Kimura, B. & Ishida, Y. Photophagotrophy in Uroglena americana, Chrysophyceae. Jpn J. Limnol. 46, 315–318 (1985).
Guillard, R. R. L. Culture of phytoplankton for feeding marine invertebrates. In Culture of Marine Invertebrate Animals: Proceedings—1st Conference on Culture of Marine Invertebrate Animals, Greenport 29–60Springer, (1975).
Andrews, S. & FastQC A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Bolger, A. M., Lohse, M., Usadel, B. & Trimmomatic A flexible trimmer for illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Li, W. & Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Haas, B. J. et al. De Novo transcript sequence reconstruction from RNA-seq using the trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Kanehisa, M. Toward pathway engineering: a new database of genetic and molecular pathways. Sci. Technol. Japan. No. 59, 34–38 (1996).
Jeanmougin, F., Thompson, J. D., Gouy, M., Higgins, D. G. & Gibson, T. J. Multiple sequence alignment with clustal X. Trends Biochem. Sci. 23, 403–405 (1998).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Verma, A., Kohli, G. S., Harwood, D. T., Ralph, P. J. & Murray, S. A. Transcriptomic investigation into polyketide toxin synthesis in Ostreopsis (Dinophyceae) species. Environ. Microbiol. 21, 4196–4211 (2019).
John, U. et al. Novel insights into evolution of Protistan polyketide synthases through phylogenomic analysis. Protist 159, 21–30 (2008).
Kimura, K. et al. RNA sequencing revealed numerous polyketide synthase genes in the harmful dinoflagellate Karenia Mikimotoi. PLoS ONE. 10, e0142731 (2015).
Nivina, A., Yuet, K. P., Hsu, J. & Khosla, C. Evolution and diversity of assembly-line polyketide synthases: focus review. Chem. Rev. 119, 12524–12547 (2019).
Murray, S. A., Diwan, R., Orr, R. J., Kohli, G. S. & John, U. Gene duplication, loss and selection in the evolution of saxitoxin biosynthesis in alveolates. Mol. Phylogenet Evol. 92, 165–180 (2015).
Vingiani, G. M. et al. De Novo transcriptome of the non-saxitoxin producing Alexandrium tamutum reveals new insights on harmful dinoflagellates. Mar. Drugs. 18, 386 (2020).
Shelest, E., Heimerl, N., Fichtner, M. & Sasso, S. Multimodular type I polyketide synthases in algae evolve by module duplications and displacement of AT domains in trans. BMC Genom. 16, 1015 (2015).
Van Dolah, F. M. et al. Subcellular localization of dinoflagellate polyketide synthases and fatty acid synthase activity. J. Phycol. 49, 1118–1127 (2013).
Cho, Y. et al. Intracellular abundance, localization, and enzymatic activity of a saxitoxin biosynthesis enzyme, SxtG, in two sister subclones of the dinoflagellate Alexandrium catenella with extremely different levels of paralytic shellfish toxins. Harmful Algae. 139, 102723 (2024).
Orr, R. J., Stüken, A., Murray, S. A. & Jakobsen, K. S. Evolution and distribution of saxitoxin biosynthesis in dinoflagellates. Mar. Drugs. 11, 2814–2828 (2013).
Hackett, J. D. et al. Evolution of saxitoxin synthesis in cyanobacteria and dinoflagellates. Mol. Biol. Evol. 30, 70–78 (2013).
Akbar, M. A. et al. Biosynthesis of saxitoxin in marine dinoflagellates: an omics perspective. Mar. Drugs. 18, 103 (2020).
Van Donk, E. & Ianora, A. Induced defences in marine and freshwater phytoplankton: a review. Hydrobiologia 668, 3–19 (2011).
Kilham, P. & Hecky, R. E. Comparative ecology of marine and freshwater phytoplankton. Limnol. Oceanogr. 33, 776–795 (1988).
Hertweck, C. The biosynthetic logic of polyketide diversity. Angew Chem. Int. Ed. 48, 4688–4716 (2009).
Li, C., Nitka, M. V., Gloer, J. B., Campbell, J. & Shearer, C. A. Annularins A–H: new polyketide metabolites from the freshwater aquatic fungus Annulatascus triseptatus. J. Nat. Prod. 66, 1302–1306 (2003).
Kaluzhnaya, O. V. & Itskovich, V. B. Features of diversity of polyketide synthase genes in the community of freshwater sponge Baikalospongia fungiformis. Russ J. Genet. 58, 336–346 (2022).
Acknowledgements
We thank Mr. T Kim for cell culture and valuable comments to the early version of our manuscript.
Funding
This work was supported by Korea Environment Industry & Technology Institute (KEITI) through Aquatic Ecosystem Conservation Research Program funded by Korea Ministry of Environment (MOE) (2022003050002 or RS-2022-KE002119211530) and by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (RS-2024-00354842).
Author information
Authors and Affiliations
Contributions
B.L.M designed the research, analyzed the data, wrote the original draft, and reviewed and edited the paper. H.S.K performed the experiment, reviewed and edited the paper. Q.T.N.B. reviewed and edited the paper. J.S.K. designed the research, supervised the research, coordinated with co-authors, and provided extensive feedback on the text.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Muhammad, B.L., Bui, Q.T.N., Kim, HS. et al. Transcriptomic insights into polyketides and toxin biosynthesis genes in freshwater dinoflagellates. Sci Rep (2026). https://doi.org/10.1038/s41598-026-40315-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-40315-x