Figure 1: Outline of Modified Primers Study.
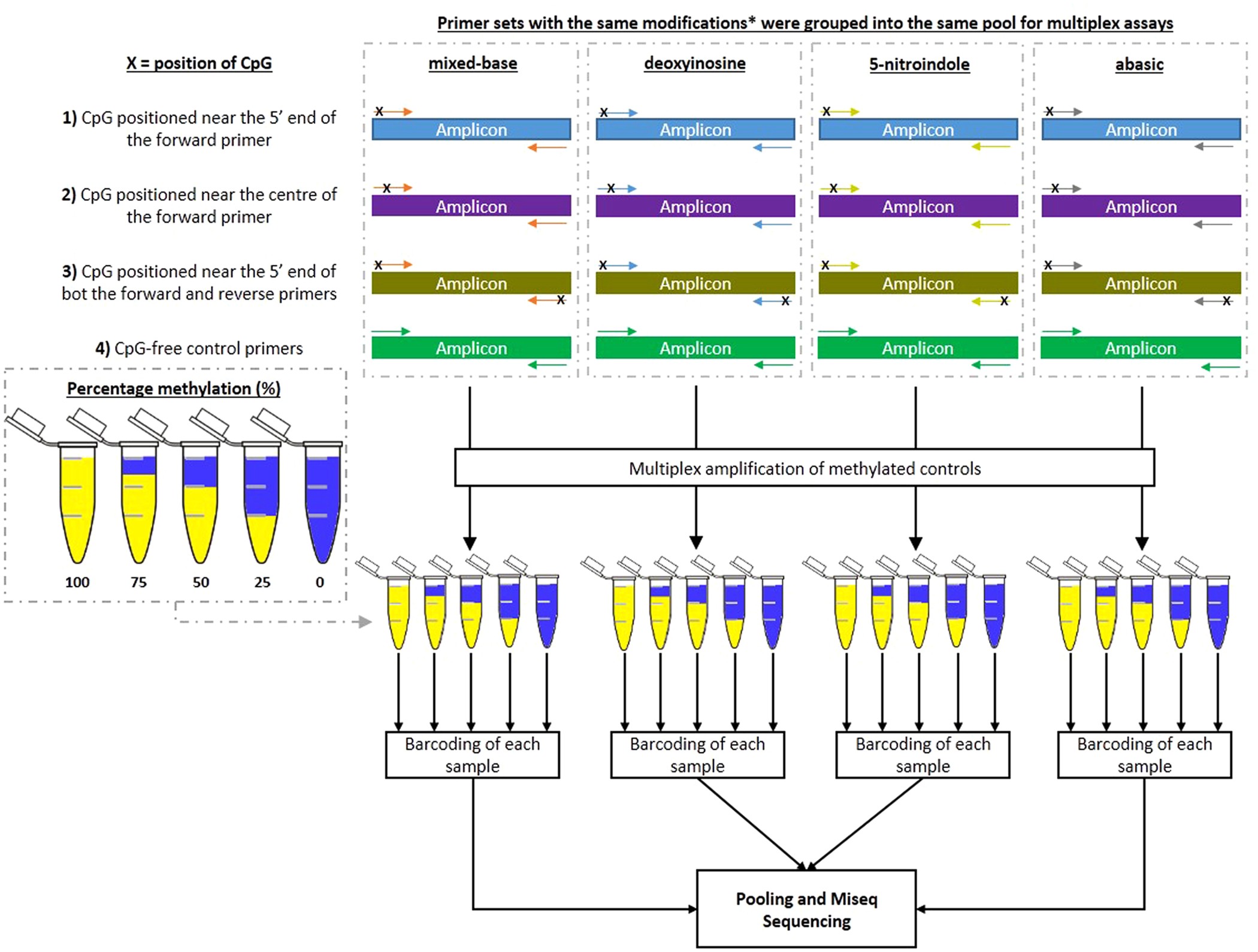
Outline of the molecular structures of the different modified bases used in this study. Four different groups of base modifications were used in this study to compensate for the change in methylation statues of the cytosine site at CpG dinucleotides. Abasic modifications acts as a space in the primer and is commonly used to study the damage and repair mechanism in DNA. For mixed- bases, the modified base R or Y was used depending on the strand orientation of the template (i.e. Y (C and T) was used for sense strand, while R (G and A) was used for the anti-sense strand). The universal bases deoxyinosine and 5-nitroindole has the ability to bind to any of the four DNA bases and therefore was also included in this assay. Each primer pool was screened against a series of graduated methylated controls in order to observe the sensitivity of the primer pairs in reporting the different methylated levels within each control sample. Primers with the same modifications were mixed into the same pool during library preparation and additional CpG-free primer controls were included into each pool as an internal assay control. Each primer pool was used to screen a series of methylated controls (100%, 75%, 50%, 25 and 0% methylation) prior to barcoding of each sample, followed by pooling and sequencing of combined samples.
