Abstract
Hepatitis C virus (HCV) frequently causes chronic liver disease. The cause of viral persistence might be an inappropriate type I interferon (IFN) induction. To analyze the host's IFN response in chronic hepatitis C, we measured the transcription level of type I IFN genes as well as type I IFN-regulated genes in liver tissue and corresponding blood samples from patients with chronic hepatitis C, nonviral liver diseases, and a suspected but later excluded liver disease. Competitive and real-time RT-PCR assays were used to quantify the messenger RNA (mRNA) levels of all known IFN-α, IFN-β, and IFN-λ genes and those of some IFN-regulated genes. We failed to detect any hepatic type I IFN mRNA induction, although liver tissue of chronic hepatitis C patients contained high numbers of some type I IFN-inducible effector mRNA molecules. Analysis of peripheral blood samples, however, showed a clear type I IFN induction. Parallel experiments employing HCV replicon cell lines revealed that replication of HCV RNA is not sufficient to induce any type I IFN nor to induce directly type I IFN-regulated genes such as MxA. In conclusion, our data provide evidence for the absence of an induction of type I IFN genes by HCV in the human liver and argue for a further development of type I IFN-based therapies.
Similar content being viewed by others
Main
Hepatitis C virus (HCV) infection is a major cause of chronic liver disease, which in many cases proceeds to fatal liver cirrhosis and hepatocellular carcinoma. Worldwide, an estimated 170 million people are persistently infected with the virus,1 indicating that HCV is able to escape innate immune response mechansims. Currently, hepatitis C patients are treated with interferon alpha (IFN-α) given either alone or in combination with the nucleoside analog ribavirin. Owing to improved application schemes, now up to half of the patients benefit from IFN-α-based therapies as evidenced by sustained virus eradication.2
IFNs are divided into type I and type II IFNs. The type I genes encode 13 closely related IFN-α subtypes, IFN-β, and the more distantly related IFN-κ, IFN-ω, and IFN-ɛ.3 The product of the only type II gene, IFN-γ, exhibits no sequence homology to other IFNs but possesses some of their biological properties (see below). Most recently, by genomic screening, a novel cytokine family has been discovered sharing sequence similarities with type I IFNs and the IL-10 family. Accordingly, the members of this family have been named IFN-λ1, IFN-λ2, and IFN-λ34 or IL-29, IL-28A, and IL-28B,5 respectively. Like type I IFNs, they are induced by viral infections and confer antiviral activity.4, 5
A hallmark of all IFNs is their ability to enhance the expression of numerous genes.3 Some of these genes encode effector proteins with antiviral activities. An example is MxA, a cytoplasmic GTPase that inhibits the replication of several RNA viruses.6 Other effector proteins are the double-stranded RNA-dependent protein kinase (PKR) or members of the family of 2′,5′oligoadenylate synthetases.3 In contrast to MxA, their activation requires binding to double-stranded RNA. While the activation of PKR leads to a translational arrest through phosphorylation of an essential translation initiation factor, the activation of 2′,5′oligoadenylate synthetases results in the production of small oligoadenylates. These in turn activate the endogenous ribonuclease RNase L, which subsequently cleaves ssRNAs including mRNAs of viral and host cell origin.
Although both types of IFNs induce the expression of a partially overlapping set of effector proteins, experiments with genetically targeted mice lacking either the type I or type II IFN receptor demonstrate that type I IFNs mediate most of the early innate immune response against viruses.7 In contrast, the contribution of IFN-γ to virus elimination is believed to be primarily systemic, for example, by enhancing the expression of proteins involved in antigen processing and presentation such as proteasome subunits and major histocompatibility complex molecules. Furthermore, IFN-γ regulates the expression of chemokines that support the adaptive immunity. Another difference between the two types of IFNs is their origin. Type I IFNs are produced by many, if not all, virus-infected cells. However, there is evidence that some cell types such as hepatocytes are poor IFN producers,8 whereas a certain blood leukocyte population, termed natural IFN-producing cells, is specialized for the production of high amounts of type I IFNs (reviewed in Colonna et al9). In contrast to type I IFNs, the expression of IFN-γ is restricted to activated T lymphocytes and natural killer cells.
Recently, it has been demonstrated that both types of IFNs efficiently inhibit HCV replication in the human hepatoma cell line Huh-7.10, 11, 12, 13 Several IFN-induced proteins have been analyzed for their ability to interfere with HCV replication, but effector proteins that inhibit HCV replication have not yet been identified.11, 14 A previous approach comparing gene expression in HCV-infected liver tissues with that in noninfected livers by using the technique of suppression subtractive hybridization revealed an enhanced (ie greater than five-fold) expression of some IFN-regulated genes in the HCV-infected livers.15 Among these are the type I IFN-regulated genes MxA, interferon inducible protein 44 (IFI-44), and interferon inducible protein 56 (IFI-56K). On the one hand, these data indicate that especially those genes that are upregulated in chronic hepatitis C might not be those that are capable to inhibit HCV replication sufficiently. On the other hand, these data indicate ongoing type I IFN-regulated processes and suggest that HCV might induce the expression of type I IFNs in liver cells.
Our current study addresses the question whether or not HCV replication triggers the induction of endogenous type I IFNs. We thus analyzed the expression of all known IFN-α, IFN-β, and IFN-λ genes in human liver tissue biopsy samples and matching blood samples from patients with chronic hepatitis C. To study the effect of virus replication on type I IFN and on type I IFN-regulated gene expression more directly in a homogenous hepatocyte-derived cell population, a number of Huh-7 cell lines containing HCV replicons were employed in parallel.
Materials and methods
Patients
As part of a routine clinical evaluation, liver biopsy procedures from a total of 60 consecutive outpatients with chronic hepatitis C (n=39; 19 female, 20 male; mean age 44.6 years, range 18–67), and nonviral liver diseases (n=12; six female, six male; mean age 45 years, range 27–63) were performed. Additionally, liver biopsy specimens were obtained from patients with a suspected but later excluded liver disease that thus can be referred to as ‘healthy liver controls’ (n=9; four female, five male; mean age 52 years, range 29–67). Chronic HCV infection was diagnosed by the detection of HCV-specific antibodies and HCV RNAs in the patient's serum. Liver disease was confirmed histopathologically as described.16, 17 Inflammatory activity was estimated to be mild in 15 patients, moderate in 19, and severe in 4. Histopathological findings and diagnoses for patients with nonviral liver diseases comprised primary biliary cirrhosis (n=2), steatosis (n=4), cryptogenic hypodense liver foci (n=1), hydropic swelling of hepatocytes (n=1), cryptogenic cirrhosis (n=2), primary sclerosing cholangitis (n=1), and Stauffer's syndrome (n=1). Diagnoses of patients with the absence of any hepatic histopathological condition included non-Hodgkin's lymphoma (n=1), dermatomyositis (n=1), gastric cancer (n=1), and mesenterial cyst (n=1). Additional samples were taken to rule out focal nodular hyperplasia (n=1), and other liver diseases in patients who showed slightly elevated serum transaminase activities (n=4). Patients with concomitant non-C viral infections and those with continued alcohol or other drug abuse were excluded. Informed consent was obtained from each patient. The study was approved by the local ethical committee of the Georg-August-University, Göttingen and conformed to the ethical guidelines of the 1975 Declaration of Helsinki.
Cells and Viruses
The human hepatoma cell line Huh-7 has been originally described by Nakabayashi et al.18 The Huh-7 cell clones 5-15 and 9-13 (containing the subgenomic HCV replicons I389/NS3-3′ and I377/NS3-3′, respectively) and the cell clones 20-1 and 21-5 (containing the genomic replicon I389/Core-3′/5.1) were previously described.19, 20, 21, 22 Maps showing the genetic organization of the replicons used in this study have already been published.13 The Huh-7 cell clones Neo 3/1 and Neo 3/2 have been generated after transfection of naive Huh-7 cells with pcDNA3 (Invitrogen, Karlsruhe, Germany). Human 293 cells were kindly provided by Friedemann Weber (University of Freiburg, Germany). All cells were maintained in Dulbecco's modified Eagle's medium supplemented with 10% fetal calf serum, 200 U/ml of penicillin G, and 200 μg/ml of streptomycin. For cells that have been transfected with HCV replicons or pcDNA3, the culture medium was additionally supplemented with 0.25–1 mg/ml of G418.
Newcastle disease virus (NDV) strain H53 (kindly provided by Peter Staeheli, University of Freiburg, Germany) was propagated by inoculating the allantoic cavity of 10-day-old embryonated chicken eggs. Stock virus contained 8.8 × 108 50% tissue culture infective doses per ml as determined on CEC32 cells (kindly provided by P Staeheli).
Preparation of Peripheral Blood Mononuclear Cells
Heparinized peripheral blood samples were taken on the same day when liver biopsy procedures were performed. Peripheral blood mononuclear cells (PBMCs) obtained from approximately 30 ml of peripheral blood were isolated by Ficoll density centrifugation and dextran sedimentation according to Boyum.23 Cell preparations were routinely assessed for viability (>95%) by trypan blue exclusion. In a typical experiment, a PBMC preparation consisted of more than 98% lymphocytes/monocytes and less than 2% granulocytes as determined by the morphology of cells stained according to Pappenheim.
Isolation of Total Cellular RNA
Total cellular RNA was prepared by CsCl density-gradient ultracentrifugation using guanidinium isothiocyanate24, 25 essentially as described.26 Starting material was homogenized liver tissue, PBMCs, or Huh-7 cells containing HCV replicons. Finally, RNA was dissolved in distilled water and the RNA concentration was determined photometrically.
Quantification of Gene Expression by Competitive RT-PCR
An amount of 2 μg of total cellular RNA was reverse transcribed as described previously.26 Quantification of cDNAs corresponding to transcripts of interest were performed by using internal cDNA standards (IS).26 In brief, IS were constructed to be complementary to and to compete with oligonucleotide primers and for amplification of target sequences. Target cDNAs were amplified in the presence of 10- and two-fold serial dilutions of the IS. The amount of target transcripts was then calculated on the basis of the known molecular quantity of the IS and related to the amount of a reference mRNA (albumin, β-actin, or glycerinaldehydephosphate dehydrogenase (GAPDH)), which had been quantified in parallel (Figure 1). Primer sequences, annealing temperature, the number of cycles carried out, the size of the target amplification product, and the size of the IS amplification product are given in Table 1.
Analysis of transcript expression by quantitative RT-PCR. cDNAs corresponding to 32 ng of total cellular RNA were coamplified with decreasing amounts of an IS in 10-fold dilution series. Concentrations of the IS (amol/reaction) are given above the lanes for IFN-α and β-actin (upper left and lower left panels, respectively). Based on the amount of the IS that yielded a stronger amplification signal than the corresponding target cDNA (arrows in the left panels), a second round of reactions was carried out in which the IS was titrated by two-fold dilutions. IS concentrations that yielded amplification products of comparable intensities as determined by visualization, or values in-between (arrows in the right panels) were taken as a basis for subsequent calculations. In the example shown, the ratio of IFN-α (0.0000935 amol) to β-actin transcripts (0.75 amol) is 0.12 × 10−3.
Quantification of Gene Expression by Real-Time PCR
A measure of 2 μg of total cellular RNA was used for reverse transcription as described above except that random hexamers (6 μM) were used for priming. Complementary DNAs (cDNAs) derived from liver tissue (corresponding to 32 ng of RNA) or PBMC or Huh-7 cell preparations (corresponding to 8 ng of RNA) were amplified using the ABI Prism 7000 sequence detection system. Quantification of IFN-α2, IFN-α8, IFN-α21, IFN-β, MxA, and GAPDH cDNAs was performed using the 5′ nuclease assay-based Assays on Demand (Applied Biosystems, Darmstadt, Germany) according to the supplier's instructions. The amount of albumin, IFN-λ1, and IFN-λ2/λ3 cDNAs was determined by using primer and 6-carboxy-fluorescein-amedit (FAM)-labeled Minor Groove Binder (MGB) probes that had been designed by the PrimerExpress software (Applied Biosystems) (albumin: forward 5′-AAC ACA AAG ATG ACA ACC CAA ACC-3′, reverse 5′-GCA GTG CAC ATC ACA TCA ACC T-3′, probe 5′-CCC CCG ATT GGT G-3′; IFN-λ1: forward 5′-CAC GCG AGA CCT CAA ATA TGT G-3′, reverse 5′-AGG GTG GGT TGA CGT TCT CA-3′, probe 5′-CCG ATG GGA ACC TG-3′; IFN-λ2/λ3: forward 5′-GCC ACA TAG CCC AGT TCA AGT C-3′; reverse 5′-GGC ATC TTT GGC CCT CTT AAA-3′, probe 5′-CTC CAC AGG AGC TGC-3′). The amount of HCV RNA in relation to a cellular reference transcript was determined using a primer pair from the 5′ UTR and a mixture of two FAM-labeled TAMRA TaqMan probes that recognize most of the viral subtypes with comparable amplification efficiency (forward 5′-CAG AAA GCG TCT AGC CAT GG-3′, reverse 5′-CGC AGA CCA CTA TGG CTC TC-3′, probe 1a/3 5′-TAG TAY GAG TGT CGT GCA GCC TCC AGG-3′, probe 1b 5′-TTA GTA TGA GTG TTG TGC AGC CTC CAG G-3′).27 Amplification of β-actin cDNA was monitored in the presence of SYBR Green I dye (Applied Biosystems) (forward 5′-AAC ACA AAG ATG ACA ACC CAA ACC-3′, reverse 5′-GCA GTG CAC ATC ACA TCA ACC T-3′). Fractional threshold cycles (CT) expressing the initial concentration of target sequence were determined according to the supplier's guidelines. Relative mRNA quantification was calculated by using the arithmetic formula 2−ΔCT. Note that ΔCT is the difference between the CT of a given target cDNA and an endogenous reference cDNA. Thus, this value yields the amount of the target normalized to an endogenous reference. For instance, a ΔCT of −10 means that the ratio of target to reference mRNAs is 2−10 or 1:1024.
IFN Promoter Analysis
Cells were seeded into six-well plates and the next day, monolayers of about 80% confluency were cotransfected with 1 μg of plasmid pαLuc or p-125Luc (kindly provided by Takashi Fujita, The Tokyo Metropolitan Institute of Medical Science, Tokyo, Japan) and 0.1 μg of pRL-SV40 (Promega, Madison, WI, USA) by using Optimem (Life Technologies, Gaithersburg, MD, USA) and the DAC-30 transfection reagent (Eurogentec, Seraing, Belgium). Note that the reporter plasmids pαLuc and p-125Luc contain the firefly luciferase gene under control of an IFN-α and IFN-β promoter enhancer element, respectively,28 whereas pRL-SV40 contains the Renilla luciferase gene under control of the constitutive simian virus 40 (SV40) promoter. At 12 h after transfection, cells were either incubated with NDV (multiplicity of infection=2) or left untreated. After 18 h of further cultivation, cells were lysed and luciferase activities were measured using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA). Finally, firefly luciferase readings were divided by the corresponding Renilla luciferase activities to correct for differences in transfection efficiency.
Statistical Analysis
Statistics were calculated by using the PC-STATISTIK software package version 4.0 (Hoffmann-Software, Giessen, Germany). Gaussian distributed data were analyzed by parametric t-tests for independent samples or by analysis of variance (ANOVA). P-values of less than 0.05 were considered statistically significant.
Results
Expression of MxA, IFN-α, IFN-β, and IFN-λ in the Liver of Hepatitis C Patients
MxA protein expression is a sensitive biological marker for ongoing virus replication and/or the presence of type I IFNs.29 It has been shown previously that liver biopsy samples of hepatitis C patients contain elevated MxA mRNA levels along with those of other type I IFN-regulated genes.15, 30, 31 To find out whether the observed expression of type I IFN-regulated genes in HCV-infected livers might be induced by intrahepatically produced IFNs, we took liver specimens from patients with chronic hepatitis C, liver disorders unrelated to viral infections, and a suspected but later excluded liver disease and analyzed the samples for the expression of type I IFN genes and that of type I IFN-regulated genes by using quantitative RT-PCR techniques (for details, see the Materials and methods section). In line with previous findings, we found that MxA is highly expressed in the liver of hepatitis C patients. The median amount of MxA mRNA in samples of hepatitis C patients was 11- and 15-fold higher than that in patients with nonviral liver diseases and healthy livers, respectively (Figure 2a). Next, we quantified the type I IFN expression by using degenerated primers that amplified mRNAs of all 13 IFN-α subtypes (subsequently referred to as IFN-αn) and specific primers that amplified IFN-β mRNAs. The median IFN/albumin mRNA ratio in the liver of chronic hepatitis C patients was 0.2 × 10−3 for IFN-αn (Figure 2b) and 0.006 × 10−3 for IFN-β (Figure 2c), respectively. These levels did not differ significantly from those in liver specimens derived from patients with liver disorders unrelated to viral infections or from those in patients with healthy livers indicating that HCV replication does not trigger the production of type I IFNs in the liver.
Intrahepatic expression of MxA, IFN-α, IFN-β, and IFN-λ in hepatitis C patients. Liver biopsy specimens taken from patients with chronic hepatitis C, nonviral liver diseases, and from those with a suspected but later excluded liver disease were quantified with respect to (a) MxA mRNAs, (b) all subtypes of IFN-α mRNAs (IFN-αn), (c) IFN-β mRNAs, (d) IFN-λ1 mRNAs, and (e) IFN-λ2/λ3 mRNAs in relation to albumin mRNAs by using quantitative RT-PCR assays. Medians are indicated by horizontal bars. Levels of significance are indicated (ns, not significant). Similar results were obtained when data were related to β-actin or GAPDH (data not shown).
To exclude that members of the IFN-λ family substitute for type I IFNs in HCV-infected livers, a randomly chosen subset of the previously used biopsy samples was analyzed for the presence of IFN-λ mRNAs. We used primers that either specifically bind to IFN-λ1 transcripts or those that recognize an evolutionary conserved sequence present in IFN-λ2 and IFN-λ3 mRNAs. Figure 2d and e shows that similar amounts of IFN-λ mRNAs were detected in samples of diseased livers irrespective of whether HCV had caused the pathological changes. However, a slight but statistically significant higher number of IFN-λ2/λ3 mRNAs was found in HCV-infected liver samples compared to healthy control livers (Figure 2e). Nevertheless, it is unlikely that the slight increase in the expression of IFN-λ2 or IFN-λ3 accounts for the expression of MxA in HCV-infected tissues. First, the livers of patients with nonviral liver diseases and those infected with HCV both contain elevated levels of IFN-λ2/λ3 mRNAs, but only the latter have elevated MxA mRNA levels also. Second, data from a previous experiment in which we used suppression subtractive hybridization to compare the gene expression in HCV-infected liver tissue to that in noninfected tissues of a comparable histopathological status, revealed no evidence for an enhanced intrahepatic expression of any of the IFN-λ genes in hepatitis C patients (R Patzwahl and S Mihm, unpublished observation). Taken together, the results suggest that the activation of IFN-regulated genes in hepatitis C patients is not associated with an intrahepatic production of type I IFNs.
Expression of MxA, IFN-α, IFN-β, and IFN-λ in PBMCs of Hepatitis C Patients
PBMCs have been reported to comprise a specialized population of leukocytes that possess the ability to produce high amounts of type I IFNs.9 This knowledge prompted us to investigate whether an infection with HCV activates the type I IFN expression in peripheral blood cells. For this analysis, we chose all available hepatitis C patients' blood samples that were obtained at the time the liver biopsy was taken (n=12). As controls, blood samples were taken from healthy volunteers instead of patients with healthy livers because the latter suffered from diseases other than hepatitis C and, therefore, might have an abnormal PBMC gene expression profile. In accordance with a previous report,32 we found that MxA mRNA was moderately but significantly elevated in PBMCs of chronic hepatitis C patients (Figure 3a). Furthermore, RNA preparations of PBMC from hepatitis C patients contained 23-fold higher levels of IFN-αn than those from healthy individuals (Figure 3b). This difference was significant irrespective of whether IFN-αn mRNA levels were related to β-actin or GAPDH as reference transcripts. Similar to IFN-αn, PBMCs of chronic hepatitis C patients also contained elevated mRNA levels of IFN-β (Figure 3c), IFN-λ1 (Figure 3d), IFN-λ2/λ3 (Figure 3e), and the IFN regulatory factor 7 (IRF-7) (data not shown). The data demonstrate that chronic HCV infection activates the expression of type I IFNs and type I IFN-regulated genes in blood cells indicating that the patients' capacity to activate type I IFNs is not generally impaired.
Expression of MxA, IFN-α, IFN-β, and IFN-λ in PBMCs of hepatitis C patients. PBMCs were isolated from blood samples taken from patients with hepatitis C at the time of liver biopsy and from healthy volunteers. RNA was prepared and analyzed for the amount of (a) MxA mRNAs, (b) all subtypes of IFN-αn mRNAs, (c) IFN-β mRNAs, (d) IFN-λ1 mRNAs, and (e) IFN-λ2/λ3 mRNAs in relation to β-actin as a reference by quantitative RT-PCR assays. Medians are indicated by horizontal bars. Levels of significance are given (ns, not significant).
Expression of MxA, IFN-α, IFN-β, and IFN-λ in Huh-7 Cells Containing HCV Replicons
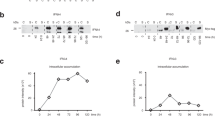
To corroborate the supposition that ongoing HCV replication does not activate type I IFN genes in infected hepatocytes, we analyzed the effect of HCV RNA replication on the innate immune response of Huh-7 cells. In contrast to the situation in vivo, in which only a proportion of hepatocytes is virus infected, the Huh-7 cell culture system for HCV replication consists of a cell population in which virtually all cells contain HCV RNAs.19 In a first experiment, we addressed the question whether HCV RNA replication induces the expression type I IFN mRNAs in cell culture. Total RNA from cells containing the subgenomic HCV replicons I389/NS3-3′ and I377/NS3-3′ (cell clones 5-15 and 9-13, respectively), the genomic replicon I389/Core-3′/5.1 (cell clones 20-1 and 21-5), and from control cells without HCV replicons (naive cells and the neomycin-resistant cell clones Neo 3/1 and Neo 3/2) was used to determine the mRNA levels of several type I IFNs by real-time RT-PCR. Figure 4 shows that cells with and without HCV replicons contained similar amounts of IFN-α2 (Figure 4a), IFN-α8 (Figure 4b), IFN-α21 (Figure 4c), and IFN-β (Figure 4d). In addition, we found that the presence of HCV replicons does not increase the amount of IFN-λ transcripts (data not shown).
Expression of type I IFN genes in Huh-7 cells with and without HCV replicons. Naive Huh-7 cells, those that carry a neomycin resistance gene (cell clones Neo 3/1 and Neo 3/2), those that contain subgenomic replicons (cell clones 5-15 and 9-13), and those that contain genomic HCV replicons (cell clones 20-1 and 21-5) were seeded into multiple cell culture dishes and cultivated for 72 h. Cells were harvested, total RNA was prepared and analyzed for the amount of (a) IFN-α2 mRNAs, (b) IFN-α8 mRNAs, (c) IFN-α21 mRNAs, (d) IFN-β mRNAs, (e) MxA mRNAs, and (f) HCV RNA in relation to β-actin mRNAs using quantitative RT-PCR assays in triplicate. Mean values and standard errors of the mean are given. The results of one representative experiment out of three are shown.
For some IFN-regulated genes, evidence has been found that their expression is induced by virus replication without a preceding type I IFN production.33, 34, 35, 36 These findings prompted us to investigate whether HCV replication in Huh-7 cells leads to the activation of IFN-regulated genes even if we could not detect an increase in the number of type I IFN mRNAs. As shown in Figure 4e, the presence of HCV replicons did not cause a general increase in the amount of MxA mRNAs although we observed that mRNA levels slightly varied between different cell clones. To exclude the possibility that different amounts of HCV RNAs account for the clonal differences in MxA expression, the number of viral nucleic acids was determined, too. Under the experimental conditions chosen, the number of viral nucleic acids was quite comparable in all Huh-7 cell clones (Figure 4f) suggesting that the presence of HCV RNAs does not enhance MxA gene expression. Furthermore, we quantified mRNA levels of additional IFN-regulated genes including those that encode IFI-44, IFI-56K, and PKR. In no case did we detect an enhanced expression in cells with HCV replicons (data not shown). Thus, we conclude that HCV replication has no direct effect on the activation of type I IFN-regulated genes.
As an alternative readout for the activation of type I IFN genes, we determined the type I IFN promoter activity in the presence or absence of HCV RNAs. Naive Huh-7 cells, several Huh-7 cell clones with or without HCV replicons, and 293 cells were transiently transfected with luciferase reporter constructs containing IFN-α or IFN-β promoter sequences. At 12 h after transfection, the cells were infected with NDV or left untreated. After an additional incubation period of 18 h, luciferase activities were determined and normalized for transfection efficiencies (see Materials and methods for details). Figure 5 shows that untreated Huh-7 cells exhibited much lower IFN-α and IFN-β baseline promoter activities than 293 cells. Furthermore, IFN promoter activities in Huh-7 cells did not increase in response to the treatment with NDV. In contrast, 293 cells showed a 38- and 25-fold enhancement of their IFN-α and IFN-β promoter activities, respectively, in response to the NDV treatment. These findings indicate that Huh-7 cells in general are rather poor IFN producers. Interestingly, the presence of HCV replicons had a weak stimulatory effect on the expression of both type I IFN promoter constructs. However, this effect was only observed with untreated cells and not with NDV-infected cells. The biological relevance of the observed slight increase in the IFN promoter activities is not clear since we did not detect increased amounts of type I IFN mRNAs by real-time RT-PCR (Figure 4a–d). Taken together, the data obtained by the HCV replicon system are in line with the weak intrahepatic type I IFN expression observed in HCV-infected individuals.
IFN-α and IFN-β promoter activity in Huh-7 cells with and without HCV replicons. Naive Huh-7 cells, those that carry a neomycin resistance gene (cell clone Neo 3/1), those that contain subgenomic HCV replicons (cell clone 9-13), those that contain genomic HCV replicons (cell clones 20-1 and 21-5), and 293 cells were cotransfected with two luciferase reporter plasmids. One plasmid encodes the firefly luciferase either under control of an (a) IFN-α or (b) IFN-β promoter. The other plasmid encodes the Renilla luciferase under control of the constitutive SV40 promoter (see Materials and methods for details). At 12 h after transfection, cells were incubated with NDV or left untreated. After 18 h, cells were lyzed and luciferase activities were determined (measurements were taken in duplicate). Columns represent the quotient of the firefly and Renilla luciferase readings. The figure shows the result of a single representative experiment.
Discussion
The present study demonstrates the absence of detectable amounts of type I IFN mRNAs in a number of individual liver samples of chronic hepatitis C patients in spite of an enhanced expression of type I IFN-regulated mRNAs (Figure 2). The expression of all known type I IFNs was monitored including those of the recently described IFN-λ family. Our findings are in line with earlier observations made on selected single tissue samples or small pools of samples that had been compared by DNA chip arrays or by related techniques. For instance, the analysis of a subtracted library (transcriptome of an HCV-infected liver tissue minus that of noninfected tissues with similiar histopathological changes) revealed an enhanced expression of some IFN-regulated genes but not that of type I IFN genes themselves.15 Moreover, when HCV-associated liver cirrhosis was compared to nondiseased liver tissue, a limited gene expression analysis using a cDNA microarray of 874 genes revealed the induction of the type I IFN-inducible peptide 6–16 in the absence of IFN-α2.37 A more comprehensive oligonucleotide microarray analysis of more than 9000 genes documented a higher expression of four IFN-inducible genes in HCV-associated hepatocellular carcinoma specimens than in noninfected nontumorous liver, but not that of type I IFN genes.38 A microarray analysis performed by Honda et al,39 in contrast, revealed an increased intrahepatic expression of IFN-α genes in chronic hepatitis C patients. The deviant findings of this analysis might be due to the fact that Honda et al compared HCV-infected liver biopsy samples to uninfected tissue samples surgically obtained from noncancerous liver parts of liver tumor patients.
Experimentally infected chimpanzees served as an animal model in several studies on the innate immune response against HCV. Oligonucleotide microarrays revealed that liver biopsy samples of acute and persistently HCV-infected animals contain high amounts of type I IFN-induced mRNAs.40, 41 Elevated type I IFN mRNAs, however, were described only in one report so far.42
In our experiments, we employed a number of HCV replicons to analyze the effect of HCV replication on the IFN gene expression in cell culture. We found that the presence of HCV replicons does not enhance the transcription of type I IFN genes and type I IFN-regulated genes (Figure 4). Furthermore, ongoing HCV replication had only minor effects on the expression of a luciferase reporter that is controlled by a type I IFN promoter (Figure 5). These findings are in line with data of a most recent transcriptome analysis, in which we used an oligonucleotide microarray that contains probe sets for all known human type I IFN genes to analyze the gene expression in Huh-7 cells with and without HCV replicons (M Trippler and M Frese, unpublished results). Our finding that the incubation of Huh-7 cells with NDV is not sufficient to induce type I IFN mRNAs is in complete agreement with earlier findings made by Keskinen et al8 who analyzed IFN production in several human hepatoma lines after stimulation with various agents and viruses. In HepG2 and Huh-7 cells, for example, they found no detectable IFN-α or IFN-β production in response to the stimuli applied. Thus, the authors suggested that hepatoma cells may have an intrinsically poor ability to produce type I IFNs, which may contribute to their inability to efficiently resist viral infections. Furthermore, our results are in line with a recent observation of Lanford et al43 who failed to detect IFN-β mRNA induction in naive Huh-7 cells after stimulation with poly-IC or in the presence of HCV replicons. The authors speculated that the lack of IFN-β inducibility in Huh-7 cells is the result of a defect in the signaling pathway for dsRNA. However, the view of poor type I IFN inducibility in hepatoma cells has recently been challenged by Foy et al44 and Fredericksen et al45, who both used Sendai virus (SENV) to successfully trigger the IFN system in Huh-7 cells as evidenced by an increased production of IFN-β. Thus, it remains an open question whether hepatocytes/hepatoma cells are generally impaired in sensing ongoing viral replication.
Interestingly, it has recently been noted that ongoing HCV replication interferes with several levels of the IFN system as do many other viruses.3 For example, Foy et al reported that the HCV protease NS3/4A inhibits the phosphorylation of IRF-3, a transcription factor required for the activation of type I IFN genes. These findings are compatible with our results in the replicon system and with the situation in the HCV-infected liver as evidenced by the present study.
The observation that some type I-regulated effector genes are activated in chronic hepatitis C might be due to secondary inflammatory events. This process, however, is obviously not able to terminate the infection because the induction is either not high enough for an efficient virus elimination or the expressed effector proteins do not affect HCV replication. In fact, the effector protein that is most frequently found in HCV infection, MxA, has been shown not to inhibit HCV RNA replication in human hepatoma cells.11
The finding that type I IFN mRNA molecules are easily detectable in peripheral blood samples but not in the liver of chronic hepatitis C patients might be due to the selective recruitment of peripheral cells. The liver of hepatitis C patients expresses interferon-inducible protein 10 (IP-10), a chemokine that attracts monocytes and activated T-helper cells.15, 46, 47, 48 The main producers of type I IFNs, however, are the so-called natural IFN producing cells (IPCs) or plasmacytoid dendritic cells (pDCs).9 Despite expression of very high levels of the IP-10 receptor CXCR3, natural IFN producing cells do not respond efficiently to CXCR3 ligands.49 Upon activation, the CXCR3 appears to be nonfunctional. On the other hand, activated natural IFN-producing cells do have a propensity to migrate to secondary lymphoid organs, rather than to sites of inflammation.50 This behavior might be due to the expression of the lymph node homing molecule 1-selectin.50 The absence of detectable hepatic type I IFN mRNA molecules substantiates the rationale for applying type I IFNs. Although being not yet as effective as desired, improvements of application schedules in the past clearly led to improvements of therapy response. Thus, our data argue for further development of type I IFN-based therapies.
References
WHO. Hepatitis C. Fact Sheet 164, 2000. http://www.who.int.
Chander G, Sulkowski MS, Jenckes MW, et al. Treatment of chronic hepatitis C: a systematic review. Hepatology 2002;36:S135–S144.
Goodbourn S, Didcock L, Randall RE . Interferons: cell signalling, immune modulation, antiviral response and virus countermeasures. J Gen Virol 2000;81:2341–2364.
Kotenko SV, Gallagher G, Baurin VV, et al. IFN-lambdas mediate antiviral protection through a distinct class II cytokine receptor complex. Nat Immunol 2003;4:69–77.
Sheppard P, Kindsvogel W, Xu W, et al. IL-28, IL-29 and their class II cytokine receptor IL-28R. Nat Immunol 2003;4:63–68.
Haller O, Frese M, Kochs G . Mx proteins: mediators of innate resistance to RNA viruses. Rev Sci Technol 1998;17:220–230.
Müller U, Steinhoff U, Reis LF, et al. Functional role of type I and type II interferons in antiviral defense. Science 1994;264:1918–1921.
Keskinen P, Nyqvist M, Sareneva T, et al. Impaired antiviral response in human hepatoma cells. Virology 1999;263:364–375.
Colonna M, Krug A, Cella M . Interferon-producing cells: on the front line in immune responses against pathogens. Curr Opin Immunol 2002;14:373–379.
Blight KJ, Kolykhalov AA, Rice CM . Efficient initiation of HCV RNA replication in cell culture. Science 2000;290:1972–1974.
Frese M, Pietschmann T, Moradpour D, et al. Interferon-alpha inhibits hepatitis C virus subgenomic RNA replication by an MxA-independent pathway. J Gen Virol 2001;82:723–733.
Guo JT, Bichko VV, Seeger C . Effect of alpha interferon on the hepatitis C virus replicon. J Virol 2001;75:8516–8523.
Frese M, Schwärzle V, Barth K, et al. Interferon-gamma inhibits replication of subgenomic and genomic hepatitis C virus RNAs. Hepatology 2002;35:694–703.
Zhu H, Zhao H, Collins CD, et al. Gene expression associated with interferon alfa antiviral activity in an HCV replicon cell line. Hepatology 2003;37:1180–1188.
Patzwahl R, Meier V, Ramadori G, et al. Enhanced expression of interferon-regulated genes in the liver of patients with chronic hepatitis C virus infection: detection by suppression-subtractive hybridization. J Virol 2001;75:1332–1338.
Mihm S, Fayyazi A, Hartmann H, et al. Analysis of histopathological manifestations of chronic hepatitis C virus infection with respect to virus genotype. Hepatology 1997;25:735–739.
Desmet VJ, Gerber M, Hoofnagle JH, et al. Classification of chronic hepatitis: diagnosis, grading and staging. Hepatology 1994;19:1513–1520.
Nakabayashi H, Taketa K, Miyano K, et al. Growth of human hepatoma cells lines with differentiated functions in chemically defined medium. Cancer Res 1982;42:3858–3863.
Lohmann V, Korner F, Koch J, et al. Replication of subgenomic hepatitis C virus RNAs in a hepatoma cell line. Science 1999;285:110–113.
Lohmann V, Korner F, Dobierzewska A, et al. Mutations in hepatitis C virus RNAs conferring cell culture adaptation. J Virol 2001;75:1437–1449.
Pietschmann T, Lohmann V, Rutter G, et al. Characterization of cell lines carrying self-replicating hepatitis C virus RNAs. J Virol 2001;75:1252–1264.
Pietschmann T, Lohmann V, Kaul A, et al. Persistent and transient replication of full-length hepatitis C virus genomes in cell culture. J Virol 2002;76:4008–4021.
Boyum A . Separation of lymphocytes, granulocytes, and monocytes from human blood using iodinated density gradient media. Methods Enzymol 1984;108:88–102.
Glisin V, Crkvenjakov R, Byus C . Ribonucleic acid isolated by cesium chloride centrifugation. Biochemistry 1974;13:2633–2637.
Chirgwin JM, Przybyla AE, MacDonald RJ, et al. Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry 1979;18:5294–5299.
Mihm S, Hutschenreiter A, Fayyazi A, et al. High inflammatory activity is associated with an increased amount of IFN-gamma transcripts in peripheral blood cells of patients with chronic hepatitis C virus infection. Med Microbiol Immunol (Berl) 1996;185:95–102.
Hennig H, Luhm J, Hartwig D, et al. A novel RT-PCR for reliable and rapid HCV RNA screening of blood donations. Transfusion 2001;41:1100–1106.
Yoneyama M, Suhara W, Fukuhara Y, et al. Autocrine amplification of type I interferon gene expression mediated by interferon stimulated gene factor 3 (ISGF3). J Biochem (Tokyo) 1996;120:160–169.
Roers A, Hochkeppel HK, Horisberger MA, et al. MxA gene expression after live virus vaccination: a sensitive marker for endogenous type I interferon. J Infect Dis 1994;169:807–813.
Leifeld L, Ramakers J, Schneiders AM, et al. Intrahepatic MxA expression is correlated with interferon-alpha expression in chronic and fulminant hepatitis. J Pathol 2001;194:478–483.
MacQuillan GC, Mamotte C, Reed WD, et al. Upregulation of endogenous intrahepatic interferon stimulated genes during chronic hepatitis C virus infection. J Med Virol 2003;70:219–227.
Meier V, Mihm S, Ramadori G . MxA gene expression in peripheral blood mononuclear cells from patients infected chronically with hepatitis C virus treated with interferon-alpha. J Med Virol 2000;62:318–326.
Guo J, Peters KL, Sen GC . Induction of the human protein P56 by interferon, double-stranded RNA, or virus infection. Virology 2000;267:209–219.
Goetschy JF, Zeller H, Content J, et al. Regulation of the interferon-inducible IFI-78K gene, the human equivalent of the murine Mx gene, by interferons, double-stranded RNA, certain cytokines, and viruses. J Virol 1989;63:2616–2622.
Memet S, Besancon F, Bourgeade MF, et al. Direct induction of interferon-gamma- and interferon-alpha/beta-inducible genes by double-stranded RNA. J Interferon Res 1991;11:131–141.
Wu C, Ohmori Y, Bandyopadhyay S, et al. Interferon-stimulated response element and NF kappa B sites cooperate to regulate double-stranded RNA-induced transcription of the IP-10 gene. J Interferon Res 1994;14:357–363.
Shackel NA, McGuinness PH, Abbott CA, et al. Insights into the pathobiology of hepatitis C virus-associated cirrhosis: analysis of intrahepatic differential gene expression. Am J Pathol 2002;160:641–654.
Iizuka N, Oka M, Yamada-Okabe H, et al. Comparison of gene expression profiles between hepatitis B virus- and hepatitis C virus-infected hepatocellular carcinoma by oligonucleotide microarray data on the basis of a supervised learning method. Cancer Res 2002;62:3939–3944.
Honda M, Kaneko S, Kawai H, et al. Differential gene expression between chronic hepatitis B and C hepatic lesion. Gastroenterology 2001;120:955–966.
Bigger CB, Brasky KM, Lanford RE . DNA microarray analysis of chimpanzee liver during acute resolving hepatitis C virus infection. J Virol 2001;75:7059–7066.
Su AI, Pezacki JP, Wodicka L, et al. Genomic analysis of the host response to hepatitis C virus infection. Proc Natl Acad Sci USA 2002;99:15669–15674.
Thomson M, Nascimbeni M, Havert MB, et al. The clearance of hepatitis C virus infection in chimpanzees may not necessarily correlate with the appearance of acquired immunity. J Virol 2003;77:862–870.
Lanford RE, Guerra B, Lee H, et al. Antiviral effect and virus–host interactions in response to alpha interferon, gamma interferon, poly(i)-poly(c), tumor necrosis factor alpha, and ribavirin in hepatitis C virus subgenomic replicons. J Virol 2003;77:1092–1104.
Foy E, Li K, Wang C, et al. Regulation of interferon regulatory factor-3 by the hepatitis C virus serine protease. Science 2003;300:M45–M48.
Fredericksen B, Akkaraju GR, Foy E, et al. Activation of the interferon-beta promoter during hepatitis C virus RNA replication. Viral Immunol 2002;15:29–40.
Mihm S, Schweyer S, Ramadori G . Expression of the chemokine IP-10 correlates with the accumulation of hepatic IFN-gamma and IL-18 mRNA in chronic hepatitis C but not in hepatitis B. J Med Virol 2003;70:562–570.
Harvey CE, Post JJ, Palladinetti P, et al. Expression of the chemokine IP-10 (CXCL10) by hepatocytes in chronic hepatitis C virus infection correlates with histological severity and lobular inflammation. J Leukoc Biol 2003;74:360–369.
Shields PL, Morland CM, Salmon M, et al. Chemokine and chemokine receptor interactions provide a mechanism for selective T cell recruitment to specific liver compartments within hepatitis C-infected liver. J Immunol 1999;163:6236–6243.
Vanbervliet B, Bendriss-Vermare N, Massacrier C, et al. The inducible CXCR3 ligands control plasmacytoid dendritic cell responsiveness to the constitutive chemokine stromal cell-derived factor 1 (SDF-1)/CXCL12. J Exp Med 2003;198:823–830.
Penna G, Vulcano M, Sozzani S, et al. Differential migration behavior and chemokine production by myeloid and plasmacytoid dendritic cells. Hum Immunol 2002;63:1164–1171.
Acknowledgements
We thank all physicians of the Division of Gastroenterology who were involved in liver biopsy procedures and patients' care and control for their kind cooperation, Friedemann Weber, Peter Staeheli, and Takashi Fujita for reagents and helpful discussions, Waltraut Kopp and Ulrike Herian for expert technical assistance, and Kerry Mills for critical reading of the manuscript. Supporting grants were obtained from the Deutsche Forschungsgemeinschaft (SFB402/C1 and SFB 402/C6) and the Bundesministerium für Bildung und Forschung (Kompetenznetz Hepatitis, Contract No. 01KI0102).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mihm, S., Frese, M., Meier, V. et al. Interferon type I gene expression in chronic hepatitis C. Lab Invest 84, 1148–1159 (2004). https://doi.org/10.1038/labinvest.3700135
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/labinvest.3700135
Keywords
This article is cited by
-
Evaluation of the Relationship between Alopecia Areata and Viral Antigen Exposure
American Journal of Clinical Dermatology (2018)
-
Gene–disease association with human IFNL locus polymorphisms extends beyond hepatitis C virus infections
Genes & Immunity (2016)
-
IFNL3/4 genotype is associated with altered immune cell populations in peripheral blood in chronic hepatitis C infection
Genes & Immunity (2016)
-
The impact of the interferon-lambda family on the innate and adaptive immune response to viral infections
Emerging Microbes & Infections (2014)
-
IL28B in hepatitis C virus infection: translating pharmacogenomics into clinical practice
Journal of Gastroenterology (2010)