Abstract
Amplification and overexpression of c-MYC is a common event in various neoplasias. Recently, comparative genomic hybridization (CGH) of primary pancreatic adenocarcinomas revealed a distinct high-level amplification of 8q23-qter, suggesting that c-MYC located on 8q24 may be a candidate oncogene. To evaluate the biological significance and prognostic value of c-MYC activation in pancreatic carcinoma, we performed interphase fluorescence in situ hybridization (FISH) and immunohistochemistry on a series of 69 primary pancreatic adenocarcinomas, 19 corresponding lymph node metastases, and 5 pancreatic intraductal lesions. Dual color FISH using a probe for c-MYC (8q24) and a centromeric probe for chromosome 8 revealed amplification of c-MYC in 32.3% and 29.4% of primary and metastatic tumors, respectively. Immunostaining identified c-MYC protein overexpression in 43.5% of primaries and 31.6% of metastases. Low concordance between positive FISH and immunostaining (13.4%) suggests multiple independent regulatory pathways of c-MYC activation. Statistical evaluation revealed significant correlation (α = 0.033) between c-MYC protein overexpression and histopathological tumor grade but absence of correlation with tumor stage or lymph node status. Analysis of pancreatic intraductal lesions showed c-MYC amplification and protein overexpression in two of five cases in which invasive carcinoma exhibited identical aberrations. We conclude that deregulation of c-MYC protein is common in pancreatic cancer and that it may be involved in early neoplastic development and progression rather than in locoregional spread of invasive cancer.
Similar content being viewed by others
INTRODUCTION
Ductal adenocarcinoma of the pancreas represents the fourth most common cause of cancer-related death, and its incidence is increasing (1). Because of the lack of early symptoms and an aggressive tumor phenotype, pancreatic cancer is often diagnosed at an incurable stage of disease. The prognosis, showing a mean 5-year survival rate of <4% (1), is dismal and emphasizes the need for early diagnosis and better treatment modalities. In recent years, considerable insight into the genetic basis of this malignancy has been achieved. The functional inactivation of tumor suppressor genes such as Smad4, TP53, and MTS1/p16, as well as the activation of distinct oncogenes (e.g., KRAS2), seems to be common in pancreatic cancer (1, 2). However, the role of other tumor genes involved in the pathogenesis of pancreatic cancer still needs to be elucidated.
In a recent comparative genomic hybridization (CGH) study on pancreatic cancer, a high–copy number amplification of chromosome 8q23-qter was identified among other chromosomal alterations (3). Amplification of 8q23-qter has been confirmed in pancreatic cell lines (4) and xenografted pancreatic tumors (5). In addition, gain of chromosome 8q in pancreatic cancer was observed earlier by conventional cytogenetics (6) and CGH (4). One important proto-oncogene that may be affected by this amplification is c-MYC, located on chromosome 8q24. c-MYC is involved in the control of cell proliferation and differentiation (7, 8). Amplification and/or overexpression of c-MYC in tumor cells is extremely common, indicating that its activation may be essential during carcinogenesis (9, 10, 11). To date, little is known about genomic and proteomic changes affecting c-MYC in pancreatic cancer. The available data are controversial and show a varying incidence of c-MYC deregulation in the small number of pancreatic cancers analyzed (5, 12, 13, 14). Therefore, further and more detailed investigations to assess the biological significance and prognostic value of c-MYC activation in pancreatic carcinogenesis are demanded. However, gene analysis in pancreatic cancer is hindered by the associated strong desmoplastic reaction, resulting in high contamination of tumor cells with nonneoplastic cells (15). To overcome this problem, we applied interphase fluorescence in situ hybridization (FISH) to validate the amplification status of c-MYC on a cell-by-cell basis. Hybridization was carried out on preparations of interphase nuclei of paraffin embedded tumor samples, thus allowing retrospective analysis of a larger tumor collective. Detection sensitivity was improved by using the tyramide signal amplification system (16). Changes in c-MYC protein expression were analyzed by immunohistochemistry using a monoclonal antibody. Finally, results were statistically correlated with clinicopathological data, including tumor stage, tumor grade, and survival time to assess the prognostic value of c-MYC activation in pancreatic cancer.
MATERIALS AND METHODS
Tumor Samples and Patient Data
Paraffin-embedded tumor samples of 69 patients with ductal adenocarcinoma of the pancreas were provided by the Institute of Pathology, Mannheim, Germany. Ampullary, duodenal, and biliary cancers were excluded by histopathological review. In 19 of these cases, lymph node metastases were investigated, and in 5 cases, foci of pancreatic intraepithelial neoplasia (PanIN; PanIN-3 in 3 cases, PanIN-2 in 2 cases [17]) were also available for analysis. Histopathological data on tumor stage and grade (UICC classification, 1997) were retrieved from the files of the Institute of Pathology. Survival-time data available for 49 patients were obtained from the Department of Surgery, Mannheim. Survival time was scored as 1 or 2, corresponding to survival of <8 months after surgery (score = 1) or of >8 months after surgery (score = 2). Tumor resection was complete (R0) in all cases.
FISH of Interphase Nuclei
Three to five paraffin sections of 10-μm thickness were stained with hematoxylin and eosin. Under microscopic control, tumor tissue was selectively collected with fine forceps. Extraction of interphase nuclei from microdissected tumor material was performed as described by Liehr et al (18).
Hybridization was carried out using a P1 probe for c-MYC (RMC08P001; provided by the Cancer Center, University of San Francisco, San Francisco, CA) and an α-satellite probe for the centromeric regions of chromosome 8 (Vysis Inc., Downers Grove, IL). Dual-color hybridization was achieved by labeling the c-MYC probe with digoxigenin by nick translation (Roche Diagnostics GmbH, Mannheim, Germany). The α-satellite probe was purchased directly labeled (SpectrumGreen) from Vysis Inc. Approximately 200 ng of the labeled c-MYC probe and 50 ng of the α-satellite probe were ethanol precipitated with 10 μg of unlabeled Cot-1 DNA (Gibco BRL, Gaithersburg, MD) and 2 μg of salmon sperm DNA (Sigma GmbH, Deisenhofen, Germany), and subsequently resuspended in 7 μL of hybridization buffer (Vysis Inc.) and 3 μL of H2O. Before hybridization, slides with interphase nuclei were incubated for 30 minutes in 4 × standard saline citrate, 0.1% Triton X, followed by microwave treatment (5 minutes, 600W) in citrate buffer and a final dehydration step in 70%-90%-100% ethanol. Five μl of hybridization mixture was applied to each slide, sealed with rubber cement, and both probe and target DNA were denaturated simultaneously at 85° C on a heating plate for 10 minutes. Hybridization was carried out for 3 days in a moist chamber at 37° C.
After incubation, coverslips were removed and slides were washed for 15 minutes in 2 × standard saline citrate and 50% formamide at 45° C, followed by 15 minutes in 1 × standard saline citrate at 45° C. For detection of the c-MYC probe, the Tyramide Signal Amplification (TSA) System from NEN Life Science GmbH (Köln, Germany) was applied according to the manufacturer's recommendations. In brief, after a 30-minute blocking step, slides were treated with POD-conjugated anti-digoxigenin antibody (1:50 in TSA blocking solution; Roche Diagnostics GmbH) for 90 minutes at 37° C. For visualization of the probe, slides were incubated for 7 minutes with tyramide-Cy3 (1:50 in TSA amplification solution). Interphase nuclei were counterstained with 0.5 μg/μL of 4,6-diamidino-2-phenylindole (DAPI) in Vectashield mounting medium (Vector Laboratories, Burlingame, CA).
Microscopy and Image Acquisition
At least 100 nonoverlapping interphase nuclei from each tumor sample were evaluated by using a fluorescence microscope (Zeiss Axiophot, C. Zeiss, Göttingen, Germany) fitted with a triple band-pass filter set (Vysis Inc.) allowing the simultaneous visualization of green and red signals. Images were acquired with a cooled charge-coupled device camera (Nu200, Photometrics Inc., Tucson, AZ) and the Quips-XL FISH-imaging software (Vysis, Inc). Each tumor cell was scored for the number of centromeric and c-MYC signals. c-MYC copy gain was defined as more c-MYC signals than centromere 8 signals in >10% of cells. This was based on two controls of normal pancreatic tissue showing more c-MYC signals than centromere 8 signals in ≥5% of cells. Hybridization efficiency was tested in lymphocytes that were always present in interphase nuclei preparations. Only FISH experiments with high hybridization efficiency (showing ≥95% of lymphocyte nuclei with two signals for c-MYC and two signals for centromere 8) were considered adequate for evaluation. Low-level amplification was defined as up to three more c-MYC signals than centromere 8 signals; high-level amplification was scored as a ratio of more than three more c-MYC signals than centromere 8 signals.
Immunohistochemistry
Immunohistochemistry was performed on 2-μm-thick paraffin sections using the anti-c-MYC monoclonal antibody 9E10 (PharMingen, San Diego, CA). Immunostaining was based on an alkaline phosphatase–conjugated streptavidin–biotin detection system (Amersham Pharmacia Biotech Inc., Piscataway, NJ), using Fast Red (Roche Diagnostics GmbH) as a chromogen. Pretreatment of sections for antigen retrieval was not necessary. Incubation of the primary antibody was carried out for 60 minutes at 37° C. As a negative control, sections were treated in the absence of primary antibody.
Immunostaining of c-MYC was scored following a semiquantitative scale (0–3) by evaluating in representative tumor areas the percentage of cells showing significantly higher immunostaining than control cells of normal pancreatic tissue. Nuclear or cytoplasmic immunostaining was considered equally as positive. Scores of 0, 1, 2, and 3 corresponded to no positive staining, <10% positive staining, 10–50% positive staining, and >50% positive staining, respectively.
Statistical Analysis
The relationship between c-MYC gene amplification, c-MYC protein overexpression, and other parameters including tumor size, histological tumor grade, tumor stage (UICC, 1997), and patient survival was assessed by Fisher's exact test using the SAS software package (SAS Institute Inc., Cary, NC).
RESULTS
Amplification of c-MYC Gene
Interphase FISH was performed on cancer nuclei isolated from archival samples of 69 primary pancreatic carcinomas, foci of PanIN in 5 cases, and 19 lymph node metastases.
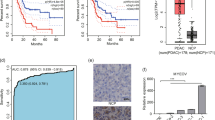
Seven tumor samples (4 primaries; 1 PanIN-3; 2 metastases) were inadequate for evaluation because of weak signals. In the remainder of the tumor preparations, hybridization of the c-MYC and centromere 8 probes resulted in bright and clear signals. Figure 1 illustrates high-level amplification and low-level amplification of c-MYC detected by FISH in two pancreatic tumor samples.
A, high-level amplification of c-MYC identified by fluorescence in situ hybridization by using the Tyramide Signal Amplification system. Hybridization was performed with a c-MYC–specific probe (red signals) and a chromosome 8 centromeric probe (green signals). The picture shows one tumor nucleus with three clusters of c-MYC signals (red) and two green spots representing chromosome 8 (not all signals in focus). Nuclei were counterstained with DAPI. B, low-level gain of c-MYC in another pancreatic carcinoma showing five c-MYC signals (red) in relation to two chromosome 8 signals (green).
FISH analysis of primary pancreatic tumors revealed a relative gain of c-MYC gene copy number in 21/65 (32.3%) cases, of which 18 tumors (27.7%) showed a low-level increase and 3 (4.6%) harbored high-level amplification. Twelve of the 65 pancreatic tumors (18.5%) revealed c-MYC gain in association with an increased chromosome 8 copy number. In these cases, the elevated copy number of c-MYC was suggestive of a global structural aberration of chromosome 8 rather than of a distinct amplification of the c-MYC gene per se. In the remaining 32 tumors, c-MYC gene copy numbers were determined as normal.
FISH analysis of 17 lymph node metastases showed amplification of the c-MYC gene in 5 samples (29.4%). Low-level amplifications were detected in three metastases (17.6%). Two lymph node metastases (11.8%) harbored a high–copy number gain of c-MYC. In two lymph node metastases (11.8%), amplification of c-MYC was identified as part of chromosome 8 gain. Ten pancreatic metastases (58.8%) showed normal copy numbers of c-MYC. Results of the FISH experiments are summarized in Table 1.
A concordant gain in c-MYC copy numbers in primary tumor and lymph node metastasis was found in two cases with low-level amplification. In two lymph node metastases, one exhibiting a low- and one a high-level increase of c-MYC, the detected aberration was restricted to the metastatic tumor. Enhancement from low- to high-level amplification was identified in one lymph node metastasis compared to its primary. Chromosome 8 polyploidy, detected in two metastases, was absent in the corresponding primaries. Interestingly, in one case, the lymph node metastasis seemed to lack c-MYC amplification, whereas the corresponding primary tumor harbored low-level amplification (data not shown).
FISH analysis of PanIN lesions demonstrated low-level amplification in high-grade foci (PanIN-3) of two cases in which adjacent invasive adenocarcinoma also showed low-level amplification. In two cases, both low-grade PanIN (PanIN-2) and invasive carcinoma harbored normal c-MYC copy numbers.
Expression of c-MYC Protein
Immunostaining for c-MYC protein was performed on the entire tumor collective (69 primary tumors, 19 lymph node metastases, and 5 PanIN lesions).
The majority of tumor cells exhibited strong nuclear staining accompanied by equally intense cytoplasmic staining for c-MYC protein. However, in some tumors, either nuclear (9 cases) or cytoplasmic (13 cases) staining predominated. There was no significant correlation between the nuclear or cytoplasmic staining pattern and gene amplification status (low or high level). Adjacent stromal cells were uniformly negative. Normal exocrine pancreatic tissue, in other words, ductal epithelium or acinar cells, revealed only very weak cytoplasmic staining. Protein overexpression was defined as strong positive immunostaining in 10–50% (score, 2) or >50% (score, 3) of tumor cells. No immunostaining (score, 0) or staining in <10% of neoplastic cells (score, 1) was interpreted as absence of protein overexpression. Figure 2 demonstrates a pancreatic carcinoma with intense cytoplasmic c-MYC staining.
Thirty of 69 primary tumors (43.5%) showed positive immunostaining in either 10–50% (27.5%) or in >50% (16%) of the tumor cells. Thirty cases (43.5%) showed c-MYC positivity in <10% of tumor cells. C-MYC staining was absent in 9 (13%) primary tumors.
Immunohistological evaluation of corresponding lymph node metastases revealed protein expression in 10–50% of the tumor cells in 6 of 19 cases (31.6%). None of the metastases revealed c-MYC positivity in >50% of the tumor cells. In the remainder of the metastases, c-MYC staining was detected in <10% of cells (7/19; 36.8%) or was completely absent (6/19; 31.6%; Table 2).
Immunostaining of intraductal lesions was positive in foci of PanIN-3 in three cases, and absence of immunostaining was seen in foci of PanIN-2 in two cases.
Correlation between Gene and Protein Expression Status of c-MYC
Gene amplification was compared with protein overexpression in 65 primary pancreatic tumors and 17 lymph node metastases (Table 3). In 11 (13.4%) tumor samples (9 primary and 2 lymph node metastases), low-level amplification of the c-MYC gene was associated with protein overexpression. All cases with high-level amplification (3 primary and 2 metastases) showed only mild up-regulation of the c-MYC protein (score, 1). In cases with chromosome 8 polyploidy, 7 of 12 primary pancreatic tumors and 1 of 2 lymph node metastases revealed c-MYC protein overexpression, whereas in the remaining 5 primaries and one metastasis, immunostaining was absent or was <10%. No concordance was seen in 12 primary and 3 metastatic tumors exhibiting gene amplification but no protein overexpression. Furthermore, 14 primary tumors and 3 metastases showed protein overexpression in the absence of gene amplification. In 25 tumors (18 primaries and 7 metastases), neither c-MYC amplification nor enhanced protein expression were identified. Although c-MYC amplification seemed to be associated with protein overexpression in PanIN lesions, numbers were considered too small to allow definitive conclusions.
Statistical Analysis
To assess the prognostic significance of c-MYC deregulation in pancreatic carcinomas, results were correlated with tumor size, grade, tumor stage, and survival time using Fisher's exact test. c-MYC activation by gene amplification did not reveal any significant correlation with clinicopathological parameters, nor was there a correlation between c-MYC protein overexpression and tumor size, tumor stage, or survival time. However, correlation of c-MYC protein expression with tumor grade was significant (α = 0.033).
DISCUSSION
In a previous CGH study on ductal adenocarcinoma of the pancreas, high–copy number amplification was found on chromosome 8q23-ter, including the locus (8q24) of c-MYC proto-oncogene (3). Activation of c-MYC by amplification has been identified in a variety of neoplasias, where it is presumed to play an oncogenic role during tumor development and/or progression (7, 8). Hence, the current study was undertaken to assess the biological role and prognostic value of c-MYC activation in pancreatic cancer.
To date, only few and controversal data on c-MYC alterations in pancreatic carcinoma exist. Yamada et al. (12) observed a 50-fold amplification of the c-MYC oncogene in one primary pancreatic carcinoma and its metastasis. In contrast, an immunohistochemical study failed to detect overexpression of c-MYC in 20 pancreatic cancers (13). Recently, amplification of the c-MYC oncogene was identified by FISH in 2 of 10 metastatic pancreatic effusion cells (14) and in 4 of 7 human xenografted pancreatic tumors (5). However, all studies pertaining to c-MYC in pancreatic cancer are based on small tumor numbers and appear therefore of limited validity.
In this study, we were able to evaluate the gene and protein expression status of c-MYC in a collective of 69 primary and 19 metastatic pancreatic tumor samples and in 5 PanIN. FISH analysis revealed a relative gain of c-MYC copy numbers in 32.3% of primary and 29.4% of metastatic tumors. Interestingly, the amplification rate of the c-MYC gene in cancers from other organs (breast, colon, prostate, stomach) is comparable with our current FISH observations in pancreatic carcinoma (11).
Gene alterations detected in this study consisted predominantly of low-level amplifications showing one to three extra c-MYC gene copies per cell (27.7% primary tumors, 17.6% lymph node metastases). A preponderance of low-level amplification has not only been confirmed in xenografted pancreatic tumors (5) but also seems to be a common phenomenon in various other tumor entities (11, 19, 20). High-level amplifications were identified in only 4.6% of primary pancreatic tumors and 11.8% of lymph node metastases. Hence, there seems to be a tendency towards the acquisition of low–copy number gains of c-MYC in pancreatic cancer.
Immunostaining detected nuclear and cytoplasmic overexpression of c-MYC protein in 43.5% of primary and 31.6% of metastatic pancreatic tumors. In contrast, a previous immunohistochemical study revealed only weak nuclear staining in a small number (2/20) of pancreatic cancers (13).
Only 11 (13.4%) tumor samples (9 primaries, 2 metastases) with low-level increase of c-MYC and 8 cases with chromosome 8 polyploidy revealed concordant protein up-regulation in 10–50% or >50% of tumor cells. Surprisingly, all cases with high-level amplification (3 primaries and 2 metastases) showed only low-level protein expression. In 38 (46.3%) tumors (31 primaries, 7 metastases), there was no association between gene amplification and protein expression level. The remaining 25 (30.5%) tumor samples (18 primaries, 7 metastases) showed neither gene amplification nor protein overexpression. Out of a total of 36 tumor samples with c-MYC protein overexpression, only 11 harbored c-MYC gene amplification. Thus, in less than one third of the analyzed pancreatic tumors, gene amplification may account for subsequent protein overexpression. The reason for this discrepancy may lie in the complex intracellular regulation mechanisms of gene and protein expression. As is the case for c-MYC, regulation mechanisms involve gene amplification, transcriptional activation, transcriptional attenuation, and mRNA/or protein stability (21, 22). Therefore, c-MYC protein expression may underlie epigenetic events that are independent of intragenetic alterations (10, 21). Alternatively, amplified gene copies may still be under control of transcriptional regulation such as promoter activation or attenuation (22). Discrepancy between c-MYC gene amplification and protein overexpression has also been described in bladder and colon cancers. It was therefore concluded that c-MYC amplification represents only one of multiple possible mechanisms leading to enhanced protein expression in various neoplasias (20, 23, 24, 25). Overall, the functional aspects of c-MYC gene amplification and protein overexpression remain unknown. It is unclear whether low-level gene amplification has any effect on the level of gene expression. Similarly, the significance of high protein expression for the oncogenic function of c-MYC still needs further investigation.
The use of a sensitive signal amplification technique for FISH (16) enabled us to identify subtle c-MYC gene copy number alterations in the analyzed tumor samples. In this context, it must be considered that a low-level increase (1–3 extra gene copies) may also represent a structural alteration of the long arm of chromosome 8 rather than a c-MYC gene specific phenomenon. Gain of chromosome 8q, including the locus of the c-MYC gene was indeed reported by several cytogenetic (6) and CGH studies (4) on pancreatic cancer. Duplication of chromosome 8q also raises the possibility that genes located on chromosome 8q other than c-MYC may be amplified and exert an oncogenic effect either dependent or independent of c-MYC overexpression. Hence, the selection for 8q amplification in pancreatic cancer may also be for a neighboring gene rather than for c-MYC itself. Interestingly, one such candidate gene located next to c-MYC is PVT1 (8q24), a transcriptional activator of c-MYC.
It has been suggested by several authors that amplification and/or overexpression of c-MYC is frequently associated with more advanced tumor stages (9, 11, 26). In consequence, the increase of c-MYC gene and/or protein expression would be expected to occur preferentially in cases with advanced local tumor extension and/or metastatic spread. Statistical analysis of our current data did not reveal any significant correlation between tumor size, stage, and c-MYC up-regulation, suggesting that activation of c-MYC is not involved in locoregional pancreatic tumor spread. However, it has to be noted that the number of pT1 and pT4 tumors in our collective is relatively small compared with that of pT2 and pT3 cancers, which may obscure statistical correlation. In addition, gene amplification and protein overexpression were detected in only 5/17 (29.4%) and 6/19 (31.6%) lymph node metastases, respectively, indicating that c-MYC activiation is not primarily involved in metastatic dissemination of pancreatic cancer. The failure by Wong et al. (19) to identify c-MYC amplification in lung metastases in their study seems to strengthen our observation.
Noninvasive precursor lesions of pancreatic adenocarcinoma have now been generally recognized, and recently a new nomenclature and classification system of these intraductal lesions has been proposed (17). In our series, low-level c-MYC amplification and protein expression was detected in PanIN-3 lesions of two cases in which adjacent invasive cancer showed similar findings of c-MYC amplification and overexpression. In contrast, two pancreatic cancers with normal c-MYC copy numbers and absence of c-MYC expression contained foci of PanIN-2 without gene amplification or protein overexpression. Although numbers are too low to allow conclusions, these findings suggest that at least in some pancreatic cancers, c-MYC alterations occur in preinvasive stages of tumor development. Reports on c-MYC deregulation in premalignant lesions of the bladder and colon (23, 24, 25) are in support of our findings. In this study, activation of c-MYC (either by gene amplification or protein overexpression) was found less frequently in lymph node metastases as compared with primary pancreatic tumors, and it did not correlate with either tumor stage or tumor size. Therefore we assume that c-MYC plays a role in the development of pancreatic neoplasia rather than in the local or metastatic spread of invasive carcinoma.
Statistical analysis of our results revealed a significant correlation (α = 0.033) between c-MYC protein overexpression and histological tumor grade. This suggests that enhanced c-MYC expression may contribute to dedifferentiation of tumor cells and thus promote tumor development. This hypothesis is supported by reports on cellular dedifferentiation in various cell lines containing multiple copies of c-MYC gene (10). Recently, the histological grade of pancreatic adenocarcinoma was proposed to be an independent prognostic factor (27). Therefore, up-regulation of c-MYC seems to be of prognostic significance, albeit not as an independent prognostic factor.
In summary, this study demonstrates that deregulation of c-MYC is common in pancreatic cancer. Gene amplification consists predominantly of a low–copy number gain and is therefore more likely to represent a structural aberration of chromosome 8 than a distinct amplification of the c-MYC gene. Low concordance between gene amplification and protein overexpression indicates that multiple regulation mechanisms can be operational at different genetic or epigenetic levels. c-MYC overexpression is significantly associated with poor tumor differentiation, which has been postulated as an independent prognostic factor in pancreatic adenocarcinoma. Although no significant correlation was found with tumor stage or metastasis, c-MYC activation was detected in some high-grade PanIN lesions. This raises the possibility that c-MYC activation is involved in earlier stages of development and progression of pancreatic neoplasia, rather than in late stages of local spread and lymph node metastasis of invasive adenocarcinoma.
References
Sakorafas GH, Tsiotou AG, Tsiotos GG . Molecular biology of pancreatic cancer; oncogenes, tumour suppressor genes, growth factors, and their receptors from a clinical perspective. Cancer Treat Rev 2000; 26: 29–52.
Sirivatanauksorn V, Sirivatanauksorn Y, Lemoine NR . Molecular pattern of ductal pancreatic cancer. Langenbecks Arch Surg 1998; 383: 105–115.
Schleger C, Arens N, Zentraf H, Bleyl U, Verbeke C . Identification of frequent chromosomal aberrations in ductal adenocarcinoma of the pancreas by comparative genomic hybridisation (CGH). J Pathol 2000; 191: 27–32.
Mahlamäki EH, Höglund M, Gorunova L, Karhu R, Dawiskiba S, Andrén-Sandberg Å, et al. Comparative genomic hybridization reveals frequent gains of 20q, 8q, 11q, 12p, and 17q, and losses of 18q, 9p, and 15q in pancreatic cancer. Genes Chromosom Cancer 1997; 20: 383–391.
Armengol G, Knuutila S, Lluís F, Capellá G, Miró R, Caballín MR . DNA copy number changes and evaluation of MYC, IGF1R, and FES amplification in xenografts of pancreatic adenocarcinoma. Cancer Genet Cytogenet 2000; 116: 133–141.
Gorunova L, Höglund M, Andrén-Sandberg Å, Dawiskiba S, Jin Y, Mitelman F, et al. Cytogenetic analysis of pancreatic carcinomas: intratumour heterogeneity and nonrandom pattern of chromosome aberrations. Genes Chromosom Cancer 1998; 23: 81–99.
Cole MD . The myc oncogene: its role in transformation and differentiation. Annu Rev Genet 1986; 20: 361–384.
Evan GI, Littlewood TD . The role of c-myc in cell growth. Curr Opin Genet Dev 1993; 3: 44–49.
Schwab M, Amler LC . Amplification of cellular oncogenes: a predictor of the clinical outcome in human cancer. Genes Chromosom Cancer 1990; 1: 181–193.
Koskinen PJ, Alitalo K . Role of myc amplification and overexpression in cell growth, differentiation and death. Semin Cancer Biol 1993; 4: 3–12.
Nesbit CE, Tersak JM, Prochownik EV . MYC oncogenes and human neoplastic disease. Oncogene 1999; 18: 3004–3016.
Yamada H, Sakamoto H, Taira M, Nishimura S, Shimosata Y, Terada M, et al. Amplifications of both c-Ki-ras with a point mutation and c-myc in primary pancreatic cancer and its metastatic tumours in lymph nodes. Jpn J Cancer Res 1986; 77: 370–375.
Sakorafas GH, Lazaris A, Tsiotou AG, Koullias G, Glinatsis MT, Golematis BC . Oncogenes in cancer of the pancreas. Eur J Surg Oncol 1995; 21: 251–253.
Zojer N, Fiegl M, Müllauer L, Chott A, Roka S, Ackermann J, et al. Chromosomal imbalances in primary and metastatic pancreatic carcinoma as detected by interphase cytogenetics: basic findings and clinical aspects. Br J Cancer 1998; 77: 1337–1342.
Klöppel G . Pathology of nonendocrine pancreas tumours. In: Go VLW, DiMagno EP, Gardner JD, Lebenthal E, Reber HA, Scheele GA, editors. The pancreas: biology, pathobiology and disease. New York: Raven Press; 1993. p. 871–898.
Raap AK, van de Corput MPC, Vervenne RAW, van Gijlswijk RPM, Tanke HJ, Wiegant J . Ultra-sensitive FISH using peroxidase-mediated deposition of biotin- or fluorochrome tyramides. Hum Mol Genet 1995; 4: 529–534.
Hruban RH, Adsay NV, Albores-Saavedra J, Compton C, Garrett ES, Goodman SN, et al. Pancreatic intraepithelial neoplasia. A new nomenclature and classification system for pancreatic duct lesions. Am J Surg Pathol 2001; 25: 579–586.
Liehr T, Grehl H, Rautenstrauss B . FISH analysis of interphase nuclei extracted from paraffin-embedded tissue. Trends Genet 1995; 11: 377–378.
Wong AJ, Ruppert JM, Eggleston J, Hamilton SR, Baylin SB, Vogelstein B . Gene amplification of c-myc and N-myc in small cell carcinoma of the lung. Science 1986; 233: 461–464.
Sauter G, Carroll P, Moch H, Kallioniemi A, Kerschmann R, Narayan P, et al. C-myc copy number gains in bladder cancer detected by fluorescence in situ hybridization. Am J Pathol 1995; 146: 1131–1139.
Krystal G, Birrer M, Way J, Nau M, Sausville E, Thompson C, et al. Multiple mechanisms for transcriptional regulation of the myc gene family in small-cell lung cancer. Mol Cell Biol 1988; 8: 3373–3381.
Marcu KB, Bossone SA, Patel AJ . MYC function and regulation. Annu Rev Biochem 1992; 61: 809–860.
Erisman MD, Rothberg PG, Diehl RE, Morse CC, Spandorfer JM, Astrin SM . Deregulation of c-myc gene expression in human colon carcinoma is not accompanied by amplification or rearrangement of the gene. Mol Cell Biol 1985; 5: 1969–1976.
Finley GG, Schulz NT, Hill SA, Geiser JR, Pipas JM, Meisler AI . Expression of the myc gene family in different stages of human colorectal cancer. Oncogene 1989; 4: 963–971.
Christoph F, Schmidt B, Schmitz-Dräger BJ, Schulz WA . Over-expression and amplification of the c-myc gene in human urothelial carcinoma. Int J Cancer 1999; 84: 169–173.
Heerdt BG, Molinas S, Deitch D, Augenlicht LH . Aggressive subtypes of human colorectal tumours frequently exhibit amplification of the c-myc gene. Oncogene 1991; 6: 125–129.
Lüttges J, Schemm S, Vogel I, Hedderich J, Kremer B, Klöppel G . The grade of pancreatic ductal carcinoma is an independent prognostic factor and is superior to the immunohistochemical assessment of proliferation. J Pathol 2000; 191: 154–161.
Acknowledgements
This work was supported by grants from the Tumorzentrum Heidelberg/Mannheim (Grant IV/10) and the Fakultät für Klinische Medizin, Klinikum Mannheim (Grant 932311).
The authors are grateful to Dr. C. Weiβ, biostatistician, for statistical advice.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Schleger, C., Verbeke, C., Hildenbrand, R. et al. c-MYC Activation in Primary and Metastatic Ductal Adenocarcinoma of the Pancreas: Incidence, Mechanisms, and Clinical Significance. Mod Pathol 15, 462–469 (2002). https://doi.org/10.1038/modpathol.3880547
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/modpathol.3880547
Keywords
This article is cited by
-
Momordicae Semen inhibits migration and induces apoptotic cell death by regulating c-Myc and CNOT2 in human pancreatic cancer cells
Scientific Reports (2023)
-
C9orf16 represents the aberrant genetic programs and drives the progression of PDAC
BMC Cancer (2022)
-
Musashi2 promotes EGF-induced EMT in pancreatic cancer via ZEB1-ERK/MAPK signaling
Journal of Experimental & Clinical Cancer Research (2020)
-
Multiclass cancer classification in fresh frozen and formalin-fixed paraffin-embedded tissue by DigiWest multiplex protein analysis
Laboratory Investigation (2020)
-
A unifying paradigm for transcriptional heterogeneity and squamous features in pancreatic ductal adenocarcinoma
Nature Cancer (2020)