Abstract
A previous study of gene flow in Heterandria formosa, the least killifish, suggested that populations on the Florida peninsula may be close to equilibrium for migration and genetic drift, but the high average value of Nem calls that conclusion into question. In this study I examine patterns of gene flow at a fine scale within the St. Johns River drainage to test the predictions that (i) Nem within a drainage should be equal or greater than that among drainages, and (ii) gene flow within a drainage should approximate a one-dimensional stepping-stone system at equilibrium. I used four isozyme systems (six loci) that were polymorphic in the previous study to characterize population structure and gene flow. Nem within each river was equal to or greater than the average value reported in the previous study, which strengthens the equilibrium interpretation of the species-wide results. However, the results showed no isolation by distance within two forks of the St. Johns drainage (the upper St. Johns and the Ocklawaha rivers), but significant differentiation between the two forks. This result has two implications: (i) that populations within each river are not at equilibrium; and (ii) that there was a historical barrier to migration between the two forks. There were substantial differences among loci, but an (appropriate) application of Lewontin and Krakauer's test revealed no evidence that locus-specific forces led to a violation of the assumption of neutrality.
Similar content being viewed by others
Introduction
Disentangling the relative contributions of ongoing gene flow and historical affiliation (i.e. past gene flow) to the genetic structure of populations is one of the most challenging problems in evolutionary biology (Felsenstein, 1982; Slatkin, 1993). One methodological problem is that traditional population genetic techniques for estimating gene flow depend on the assumption of an equilibrium between migration and genetic drift (Wright, 1978), an assumption that has not often been systematically tested (but see Larson et al., 1984; Whitlock, 1992; Hellberg, 1994; Green et al., 1996). In most cases, nonequilibrium population structure has been invoked on an ad hoc basis, to explain counterintuitive results.
A prior study suggested that populations of the freshwater live-bearing fish Heterandria formosa on the Florida peninsula may be close to migration-drift equilibrium under a two-dimensional stepping-stone model (Baer, 1998). Two results from that study supported an equilibrium interpretation: (1) there was isolation by distance between pairs of populations separated by large geographical distances, but not between populations separated by smaller distances; and (2) the slope of the regression of the pairwise estimate of the effective number of migrants (M^) against geographical distance was almost exactly the predicted equilibrium value of −0.5 (Slatkin, 1991). However, the effective number of migrants was large (Nem=23.1, 95% CL=14.5, 37.5), a fact not readily reconciled with the equilibrium interpretation, which means that the best interpretation of the spatial pattern of genetic variation in H. formosa remains arguable.
The biology of the species leads to two predictions regarding equilibrium population structure. First, in a system at equilibrium everywhere, Nem within a river drainage should be equal to or greater than that among drainages. Secondly, the pattern of migration within a river drainage should approximate a one-dimensional stepping-stone model, with the resultant isolation by distance. Previous work did not include samples at a sufficiently small scale to test those predictions with confidence. In this study I test those predictions by examining the population genetic structure of H. formosa at a fine scale within the St. Johns River drainage on the Florida peninsula. The St. Johns drainage was chosen because it is the longest drainage in peninsular Florida; if isolation by distance occurs at the level of the river drainage, it should be detectable in the St. Johns drainage.
Natural history of Heterandria formosa
Heterandria formosa is a small (12–30 mm), prolific live-bearing fish, common throughout the coastal plain of the south-eastern United States, ranging from at least as far west as the western Mississippi river drainage in West Baton Rouge Parish, LA, throughout the Florida peninsula, and as far north as the Cape Fear River, NC (Martin, 1980). It occurs almost exclusively in slow-moving, heavily vegetated waters, and although typically locally abundant it may be patchily distributed throughout a river system. It is most abundant in fresh water, but is occasionally found in brackish waters (<20 p.p.t.) (Martin, 1980). On the Florida peninsula, reproduction typically begins in March and continues through October (personal observation), resulting in 2–4 generations per year (J. Leips, personal communication). Females superfetate (incubate embryos at different stages of development simultaneously), and give birth to small clutches (1–8 offspring) at intervals of a few days to two weeks (unpublished data). Heterandria formosa is the only North American representative of its genus; all congeners occur from the Yucatan peninsula southward into Central America.
Zoogeography of the St. Johns River drainage
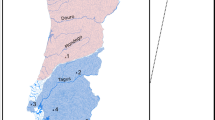
The St. Johns drainage (Fig. 1) is unique among peninsular drainages in that it contains a number of species with disjunct distributions in mainland US drainages (Swift et al., 1986), as well as numerous endemic lotic species or subspecies with sister taxa either on the US mainland or in the Suwannee/Santa Fe drainage to the west (Burgess & Franz, 1978). The distributional patterns support a western colonization of the St. Johns drainage via a Santa Fe-Ocklawaha connection following the Wicomico high stand in the early Pleistocene. Subsequent Pleistocene sea level rises repeatedly inundated large portions of the St. Johns drainage, but several streams draining west into the lower St. Johns and south into the lower Ocklawaha were never flooded and probably served as refugia for freshwater species (Burgess & Franz, 1978). The major geographical features of the St. Johns drainage therefore appear to date from the mid-Pleistocene, a sufficient time for the system to have equilibrated given the values of Nem determined in the prior study (Crow & Aoki, 1984).
Map of the St. Johns river drainage. Numbers represent collecting sites; sample sizes are in parentheses. 1, Blue Cypress Lake, Indian River Co. (58); 2, US Hwy 192 bridge (71); 3, State Hwy (SH) 520 bridge (68); 4, SH 50 bridge (69); 5, SH 46 bridge (61); 6, SH 415 bridge (72); 7, SH 44 bridge (65); 8, SH 40 bridge (62); 9, Salt Springs, Marion Co. (64); 10, SH 19 bridge (47); 11, Orange Springs boat ramp, Marion Co. (48); 12, Marion County Rd. 315 bridge (47); 13, Gore's Landing, Marion Co. (48); 14, Moss Bluff park, Marion Co. (48); 15, Marion County Rd. 42 bridge (48); 16, Haines Creek, Lake County Rd. 44 bridge (48).
Materials and methods
Collection and electrophoresis
Fish were collected from nine sites on the upper St. Johns river in June 1995 and from seven sites on the Ocklawaha River in September 1995 (Fig. 1). These sites were separated by an average of 26 km of linear river distance. I attempted to collect fish from the lower St. Johns (below the confluence of the St. Johns and the Ocklawaha) in June 1995, but the species was extremely sparsely distributed and I was unable to obtain reasonable sample sizes (i.e. <≈3–4 per site). Samples were stored and prepared following the protocol of Trexler (1988). I stained for four enzyme systems with six putative loci that the previous study showed were polymorphic throughout the species range. Phosphoglucomutase (Pgm-1 and Pgm-2) and isocitric dehydrogenase (Idh-1 and Idh-2) were run on Tris/Citrate pH 7.0 buffer. Glucose phosphate isomerase (Gpi) and mannose phosphate isomerase (Mpi) were run on Tris/EDTA/Borate pH 8.0 buffer (Morizot & Schmidt, 1990). Starch gels (12%) were run under constant voltage until xylene cyanole tracking dye had migrated ≈12 cm from the origin; generally this took from 12 to 16 h, depending on the buffer system. Enzymes were stained according to modifications of protocols listed in Morizot & Schmidt (1990). Comparisons between sites were carried out in a stepwise fashion, with 12 individuals from two (usually) geographically adjacent sites run on the same gel. Upon completion of the initial survey, samples of each variant electromorph (=‘allele’) were re-run in ‘line-up’ gels to assign relative mobility and establish the veracity of the initial interpretation. Only putative variants that were repeatably demonstrable were scored as variants; otherwise they were scored as the most common allele at that site. Most variants were repeatably demonstrable. Allele frequencies at all sites at all loci included in the analyses are presented in the Appendix.
Model of dispersal
I assumed a priori that dispersal in the St. Johns/Ocklawaha drainage fits a one-dimensional stepping-stone model (Kimura & Weiss, 1964). Each collecting site represents a ‘stone’, and distance between sites was determined by riverine distance scaled from 1:126 720 maps using a planimeter. The matrix of pairwise distances between sites is available upon request.
Relationships between M^ and distance
Recent theoretical work (Slatkin & Maddison, 1990; Slatkin, 1991, 1993) has established that under certain conditions there are simple relationships between the pairwise estimate of FST, the among-population component of genetic variance (Wright, 1978), and geographical distance in a system at migration-drift equilibrium. Using the statistic M^=(1/FST−1)/4, under a one-dimensional stepping-stone model at equilibrium, M^=4Nem/k, where k is geographical distance. Under a (symmetric) two-dimensional model at equilibrium, M^≈Nem/√k. These equations lead to the prediction that the slope of the regression of log10(M^) against log10(k) should equal −1.0 at equilibrium under a one-dimensional stepping-stone model, and approximately -0.5 under a two-dimensional model (Slatkin & Maddison, 1990; Slatkin, 1991).
Data analysis
Observed genotype frequencies at each locus at each site were compared to the values expected under the assumption of Hardy-Weinberg equilibrium using the exact probabilities option in BIOSYS-1 (Swofford & Selander, 1989). Level of significance was held at the experiment-wide level of 0.05 using the Bonferroni correction 0.05/n, where n is the number of comparisons (Trexler, 1988).
Hierarchical F-statistics were calculated using the Wright78 option of BIOSYS-1, which estimates FST using the equation of Wright (1978). Hierarchical levels are individual within total (FIT), individual within site (FIS), site within river (FSR) and river within total (drainage; FRT).
Pairwise values of M^ were calculated using the program of Slatkin (1993) provided in the GENEPOP population genetics data analysis package (Raymond & Rousset, 1995). M^ was calculated using Nei's GST (Nei, 1973) as an estimator of FST, to avoid negative values. The matrix of pairwise values of M^ is available upon request.
Ordinary least squares (OLS) regressions of log10(M^) against log10(distance) were calculated using the Regression protocol in Excel 5.0. Regressions were calculated for all pairs of sites, and for the reduced matrices of St. Johns sites only and Ocklawaha sites only. The 95% confidence intervals around calculated regression slopes were determined using OLS formulae, but with the number of populations rather than the number of pairwise comparisons as degrees of freedom; simulations using Mantel's randomization test showed that the two methods agree to two decimal places (Hellberg, 1994).
If there was not a significant regression of M^ on distance, I tested for population subdivision using the chi-squared method of Workman & Niswander (1970).
Global (i.e. not pairwise) estimates of Nem were calculated from the intercept of the regression, which provides an estimate of Nem at one unit of distance (assumed equivalent to one ‘stepping-stone’; Slatkin, 1991); 95% confidence limits were calculated using the number of populations to establish degrees of freedom. If there is no relationship between FST and distance, the regression intercept equals the mean M^.
Results
Out of 96 possible combinations of sites and loci, 78 were polymorphic. Of the 78 tests for deviation from Hardy–Weinberg equilibrium, two had individual probabilities below 0.05, and none was significant at the critical level of 0.05/78=0.0007. Therefore, all sites and loci were included in the subsequent analysis.
The value of FIS for Pgm-2 in the upper St. Johns is large and positive (0.179), whereas in the Ocklawaha it is small (-0.030). For all other combinations of locations and loci, FIS is small. This pattern is similar to that found in the species-wide study, wherein FIS for Pgm-2 was highly variable among populations, and sometimes large and positive (Baer, 1998). The simplest explanation for this observation is that there is a null allele(s) present at the Pgm-2 locus in some populations, and that nulls are more common in the St. Johns than in the Ocklawaha. Heterozygotes with a null allele will appear as homozygotes on a gel.
The hierarchical F-statistics reveal several interesting patterns of population differentiation (Table 1). F-statistics averaged over loci show that there is relatively little population subdivision within rivers (FSR), but considerable differentiation between the two rivers (FRT). However, the pattern varies greatly among loci. Although three loci are highly variable (E[H]>0.2), almost all the between-river component of variance is attributable to differentiation at Mpi. Because only one out of three highly polymorphic loci contributed to the variance, resampling across loci to determine confidence intervals is unnecessary.
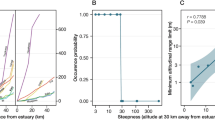
The regressions of log10(M^) against log10(distance) reveal how hierarchical population structure can influence estimators of gene flow. When all pairwise comparisons from both rivers are considered, the slope of the regression is−0.89 (95% CL=−0.53, −1.18), with r2=0.26 (Fig. 2a). This result is consistent with that predicted from a one-dimensional stepping-stone system at, or close to, migration-drift equilibrium, and indicates significant isolation by distance. However, when the upper St. Johns and the Ocklawaha are considered separately, a qualitatively different pattern emerges. Although there is significant differentiation among sites in both rivers (St. Johns: χ296=149.2, P<0.001; Ocklawaha: χ266=163.4, P<0.0001), in neither case does regression of log10(M^) against log10(distance) yield a significant regression (Fig. 2b,c). Thus, there is no isolation by distance among sites within either the upper St. Johns or the Ocklawaha.
The influence of hierarchical structure is also evident in the calculation of the effective number of migrants, Nem, from the regression intercept. When all populations are considered together, Nem= 406.4 (95% CL=112.5, 1467.9), much higher than within either the upper St. Johns by itself (Nem=100.0 [14.7, 680.0]) or the Ocklawaha by itself (Nem=20.8 [1.17, 368.6]). This result is an artifact of the apparent relationship between FST and distance that occurs when subpopulations from two differentiated populations are inappropriately pooled without considering hierarchical structure. If Nem between the two rivers is calculated using the between-river component of genetic variance (FRT), the value of Nem is 3.13, substantially less than the within-river values, as expected. This discrepancy underscores the importance of determining the appropriate spatial scale at which to estimate gene flow (Slatkin, 1993).
Discussion
The results of this study reveal differences in evolutionary dynamics at different geographical levels, and possibly among enzyme loci as well. Taking the multilocus data at face value for the moment, several conclusions follow. For the purpose of comparison to the species-wide study, the most important, albeit unsurprising, result is that the magnitude of gene flow (Nem) within each river of the drainage is equal to or greater than that over the peninsula as a whole. This reinforces the conclusion that populations on the peninsula as a whole may not be far from migration-drift equilibrium, even given the large value of Nem.
The lack of isolation by distance within rivers is more problematic. One possibility is that there is sufficient long-distance gene flow for the system to approximate an island model, although it seems unlikely that the true probability of migration between any pair of sites is independent of distance. Alternatively, it may be that population structure within rivers is not at equilibrium, probably because of repeated local extinctions and recolonizations. In either case, this result at least partly explains the lack of isolation by distance at short distances observed on the entire peninsula. McClenaghan et al. (1985) found a similar lack of isolation by distance in the poeciliid Gambusia holbrooki in the Savannah River drainage, suggesting that this pattern may be a somewhat general feature of poeciliid natural history.
The differentiation between the Ocklawaha and the upper St. Johns most likely results from a historical barrier to gene flow. The Rodman dam is a current barrier to gene flow between the Highway 19 site and sites further upstream on the Ocklawaha, but it was constructed in the late 1960s, and it seems unlikely that migration between the Highway 19 site and the upper St. Johns is high enough to have caused the Highway 19 population to diverge from the upper St. Johns to the degree it has in only 30 years (≈100 generations). One possible scenario is that one (or more) of the Pleistocene sea level rises brought salt water upstream on the present-day lower St. Johns as far as the confluence of the Ocklawaha and the St. Johns, thus effectively isolating the two rivers (Gilbert, 1987; R. Franz, personal communication).
Of more general interest is the fact that the calculated value of Nem depends on whether all populations within the drainage are considered together or separately. It should be noted that the method of calculating Nem from the intercept of the regression of log(M^) on log(distance) differs from the standard island model method of Wright (1978). Because the apparent relationship between FST and distance over the entire drainage is an artifact, the estimate of Nem from the regression intercept in that case is too high. However, in the case in which there is a real relationship between FST and distance (probably a more common situation than the converse), Nem calculated by Wright's formulation will be an underestimate. In any event, attempts to infer gene flow from population genetic data must take the hierarchical structure into account (Trexler, 1988; Slatkin, 1993). Phylogeographical methods (Avise, 1989) may often be useful in determining the appropriate scope for gene flow analyses (Slatkin, 1993; Baer, 1998).
Patterns of allele frequencies at individual loci suggest that different loci may be subject to different evolutionary dynamics. Because migration, drift, and inbreeding will, on average, affect all loci equally, it is tempting to attribute substantial differences in FST among loci to locus-specific forces, especially selection. Lewontin & Krakauer (1973) proposed that under a model of neutral evolution, σ2=kF2/(n−1), where σ2 is the expected variance in FST among loci, k is a constant dependent on the underlying distribution of allele frequencies among populations with expectation ≈2 (but see Ewens, 1977), n is the number of subpopulations, and F is the mean value of FST across loci. The expected variance, σ2F, is distributed approximately as χ2/df, where df=number of loci. The ratio of the observed variance in FST among loci, s2F, can then be compared to the expected variance by a χ2-test. Although the LK test is not a valid test of selection for a variety of reasons (Nei & Maruyama, 1975; Robertson, 1975; Ewens, 1977), it is a useful heuristic test for locus-specific effects (i.e. selection and/or mutation) IF an island model describes the migration pattern (Lewontin & Krakauer, 1975). Because there is no relationship between FST and distance within rivers, I performed an LK test on the between-river component of genetic variance, FRT (Table 1); this is equivalent to an island model with two populations. When all six loci are considered, the test is significant (χ26=16.50, P<0.02). However, when only the three highly polymorphic loci are considered, the result is not significant (χ23=4.03, P≲0.25). Because this is a study of gene flow and not of selection, it is the highly polymorphic loci that are of interest, inasmuch as they contribute most of the variance among populations. Therefore, the observed differences in FST among loci are consistent with those expected as a result of neutral evolution.
References
Avise, J. C. (1989). Gene trees and organismal histories: a phylogenetic approach to population biology. Evolution, 43: 1992–1208.
Baer, C. F. (1998). Species-wide population structure in a southeastern US freshwater fish, Heterandria formosa: gene flow and biogeography. Evolution, 52: 183–193.
Burgess, G. H. and Franz, R. (1978). Zoogeography of the aquatic fauna of the St. Johns river system with comments on adjacent peninsula faunas. Am Midl Nat, 100: 160–170.
Crow, J. F. and Aoki, K. (1984). Group selection for a polygenic behavioral trait: estimating the degree of population subdivision. Proc Natl Acad Sci USA, 81: 6073–6077.
Ewens, W. J. (1977). Population genetics theory in relation to the neutralist-selectionist controversy. In: Harris, H. & Hirschhorn, K. (eds) Advances in Human Genetics, vol. 8, pp. 67–134. Plenum Press, New York.
Felsenstein, J. (1982). How can we infer geography and history from gene frequencies? J Theor Biol, 96: 9–20.
Gilbert, C. R. (1987). Zoogeography of the freshwater fish fauna of southern Georgia and peninsular Florida. Brimleyana, 13: 25–54.
Green, D. M., Sharbel, T. F. and Kearsley, J. (1996). Postglacial range fluctuation, genetic subdivision and speciation in the western North American spotted frog complex, Rana pretiosa. Evolution, 50: 374–390.
Hellberg, M. E. (1994). Relationships between inferred levels of gene flow and geographic distance in a philopatric coral Balanophyllia elegans. Evolution, 48: 1829–1854.
Kimura, M. and Weiss, G. H. (1964). The stepping stone model of population structure and the decrease of genetic correlation with distance. Genetics, 49: 561–576.
Larson, A., Wake, D. B. and Yanev, K. P. (1984). Measuring gene flow among populations having high levels of genetic fragmentation. Genetics, 106: 293–308.
Lewontin, R. C. and Krakauer, J. (1973). Distribution of gene frequency as a test of the theory of the selective neutrality of polymorphisms. Genetics, 74: 175–195.
Lewontin, R. C. and Krakauer, J. (1975). Testing the heterogeneity of F values. Genetics, 80: 397–398.
Martin, F. D. (1980). Heterandria formosa Agassiz, Least Killifish. In: Lee, D. S., Gilbert, C. R., Hocutt, C. H., Jenkins, R. E., McAllister, D. E. & Stauffer, J. R., Jr (eds) Atlas of North American Fishes, pp. 547. North Carolina State Museum of Natural History, Raleigh, NC.
McClenaghan, L. R., JR, Smith, M. H. and Smith, M. W. (1985). Biochemical genetics of mosquitofish. IV. Changes of allele frequencies through time and space. Evolution, 39: 451–460.
Morizot, D. C. and Schmidt, M. E. (1990). Starch gel electrophoresis and histochemical visualization of proteins. In: Whitmore, D. H. (ed.) Electorphoretic and Isoelectric Focusing Techniques in Fisheries Management, pp. 24–80. CRC Press, Boca Raton, FL.
Nei, M. (1973). Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA, 70: 3321–3323.
Nei, M. and Maruyama, T. (1975). Lewontin-Krakauer test for neutral genes. Genetics, 80: 395
Raymond, M. and Rousset, F. (1995). GENEPOP (version 1.2). Population genetics software for exact tests and ecumenicism. J Hered, 86: 248–249.
Robertson, A. (1975). Remarks on the Lewontin-Krakauer test. Genetics, 80: 396
Slatkin, M. (1991). Inbreeding coefficients and coalescence times. Genet Res, 58: 167–175.
Slatkin, M. (1993). Isolation by distance in equilibrium and non-equilibrium populations. Evolution, 47: 264–279.
Slatkin, M. and Maddison, W. P. (1990). Detecting isolation by distance using phylogenies of genes. Genetics, 126: 249–260.
Swift, C. C., Gilbert, C. R., Bortone, S. A., Burgess, G. H. and Andyerger, R. W. (1986). Zoogeography of the freshwater fishes of the southeastern United States: Savannah river to Lake Ponchartrain. In: Hocutt, C. H. & Wiley, E. O. (eds). The Zoogeography of North American Freshwater Fishes, pp. 213–266. John Wiley and Sons, New York.
Swofford, D. L. and Selander, R. B. (1989). BIOSYS-1. Release 1.7. University of Illinois Press, Urbana, IL.
Trexler, J. C. (1988). Hierarchical organization of genetic variation in the sailfin molly, Poecilia latipinna (Pisces: Poeciliidae). Evolution, 42: 1006–1017.
Whitlock, M. C. (1992). Nonequilibrium population structure in forked fungus beetles: extinction, colonization, and the genetic variance among populations. Am Nat, 139: 952–970.
Workman, P. L. and Niswander, J. D. (1970). Populations studies on southwestern Indian tribes. II. Local genetic differentiation in the Papago. Am J Hum Genet, 22: 24–49.
Wright, S. (1978). Evolution and the Genetics of Populations, vol.4. Variability Within and Between Populations. University of Chicago Press, Chicago.
Acknowledgements
I thank J. Travis for his ongoing support, both financial and intellectual, of these studies. B. Maruca and R. Rogers helped with electrophoresis, and G. Coffey and H. Rodd helped collect fish. I thank L. Horth, G. Lebuhn, M. Ptacek, and M. Ruckelshaus for comments on the manuscript, and M. Hellberg, S. Karl, J. Trexler, and T. Turner for enlightening discussion. This project was supported by NSF grant DEB 92–20849 to J. Travis.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Baer, C. Population structure in a south-eastern US freshwater fish, Heterandria formosa. II. Gene flow and biogeography within the St. Johns River drainage. Heredity 81, 404–411 (1998). https://doi.org/10.1046/j.1365-2540.1998.00410.x
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1046/j.1365-2540.1998.00410.x
Keywords
This article is cited by
-
Paleoclimatic modeling and phylogeography of least killifish, Heterandria formosa: insights into Pleistocene expansion-contraction dynamics and evolutionary history of North American Coastal Plain freshwater biota
BMC Evolutionary Biology (2013)
-
Variation in predation pressure as a mechanism underlying differences in numerical abundance between populations of the poeciliid fish Heterandria formosa
Oecologia (2006)