Abstract
Mitochondrial DNA diversity of seven Silurus glanis populations (six from Greece and one from the Danube Delta) and three populations of the endemic Greek Silurus aristotelis was investigated. RFLP analysis of four regions of mitochondrial DNA (cytochrome b, D-loop, ND-5/6) amplified by PCR was used. Ten and nine haplotypes were found in S. glanis and S. aristotelis, respectively. No haplotype was shared between the two species. Significant geographical substructuring was observed in the distribution of haplotypes, with most populations possessing private haplotypes. These haplotypes can serve as genetic ‘tags’ and therefore warrant protection. Haplotype diversity was very low for all Greek S. glanis populations, possibly because the small size and large annual fluctuations of Greek inland waters do not support large fish populations. Nucleotide divergence was in the range of 0.00–0.52% among S. glanis populations, and 0.00–0.11% among S. aristotelis populations. Historical factors such as glaciations could account for these low values. The value of 6.75% sequence divergence of the two species refutes the classification of the two species in different genera, as proposed by some authors. This study constitutes the first attempt, based on mitochondrial molecular data, to address the complicated evolutionary history of the two species which belong to the widely distributed and economically important Siluridae family.
Similar content being viewed by others
Introduction
The Siluridae family consists of more than 100 freshwater fish species with Eurasian distribution (Kobayakawa, 1989; Bornbusch, 1995). In Europe it is represented by only two species: Silurus glanis, Europe’s largest freshwater fish, found from the river Rhine eastwards, including Asia Minor, and S. aristotelis which is endemic to the basin of the river Acheloos in Greece (Economidis & Banarescu, 1991).
Despite the increasing importance of S. glanis in aquaculture, little is known about the genetic characterization of natural populations. Studies on allozyme variation of the two species (Kotsinas et al., 1989; Chombard, 1993, Triantafyllidis et al., 1999) revealed low to moderate variability. Karyotype analysis of the two species has also shown few differences within the two species (Rab et al., 1994).
Another scientifically interesting problem is the taxonomic classification of the two species. Many authors (Bleeker, 1862; Banarescu, 1989) have reported that the species aristotelis belongs to the genus Parasilurus. This proposed genus consists of species with an eastern Asian and Japanese distribution. Others have questioned the taxonomic validity of the genus Parasilurus and have considered it synonymous with Silurus (Kobayakawa, 1989; Bornbusch, 1995).
Mitochondrial DNA has proved to be an excellent tool for examining biogeographical events, above or below the species level (Avise, 1994). It has emerged as a genetic marker able to discriminate stocks (Billington & Hebert, 1991) and to trace past isolation incidents, in a more effective way, in some cases, than the nuclear genome (Taylor et al., 1997).
This study constitutes the first attempt to analyse mtDNA variation of Silurus fish populations using the Restriction Fragment Length Polymorphism technique. The aim of this paper is to describe the geographical distribution of mtDNA variation in the studied populations, evaluate the usefulness of mtDNA analysis in stock identification, compare the results with the previous study on allozyme variation and examine the relationships of the two species in a phylogeographical context.
Materials and methods
Sample collections
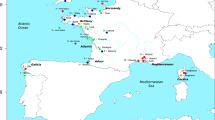
Samples from 10 different populations (Fig. 1) of catfishes were collected by traps and/or electrofishing. Three populations are of S. aristotelis and the rest belong to S. glanis. The two S. aristotelis populations from lakes Trichonida (TRI) and Amvrakia (AMV) are native and the third from lake Volvi (VOL) was recently introduced from Trichonida. Silurus glanis samples come from five different Greek presumably native unstocked river populations (Evros, EVR; Strymon, STN; Axios, AXI; Aliakmon, ALI; and Pineios, PIN), one hatchery population near the Strymon river (STH, probably established from a few individuals from the Strymon river) and one native population from Lake Puiu in the Danube Delta (DDN).
Sampling sites. Silurus aristotelis populations: TRI, Lake Trichonida; AMV, Lake Amvrakia; VOL, Lake Volvi. S. glanis populations: EVR, Evros river; STN, Strymon river; STH, Strymon river hatchery; AXI, Axios river; ALI, Aliakmon river; PIN, Pineios river. Sample 10 came from the Danube Delta (DDN) and is not indicated on the map.
mtDNA RFLP analysis
DNA was extracted from the liver or muscle of 247 individuals using the method of Taggart et al. (1992). Two adjacent mtDNA segments were amplified. One (ND-5/6) encompassed the complete NADH-dehydrogenase 5/6 region and the other (HN20) comprised the cytochrome b gene and the control region. PCR conditions were as described in Bernatchez et al. (1995). Three to 9 μL of the PCR product were subsequently digested with 18 restriction endonucleases, electrophoretically separated in 1.5% agarose gel, stained with ethidium bromide and visualized under UV light. The following restriction enzymes were tested: AluI, AseI, AvaII, BfaI, Bst NI, DdeI, EcoRI, HaeIII, HhaI, HinfI, HpaII, MboI, RsaI, TaqaI, HincII, HpaI, NruI and PstI. The last four enzymes had no restriction sites in the ND-5/6 region of S. glanis and they were subsequently not used.
Data analysis
Distinct single endonuclease patterns were identified by a specific letter. Each fish was assigned a multi-letter code that described its composite mtDNA genotype-haplotype. The raw data were fragment profiles, but we inferred site differences among haplotypes from changes in fragment profiles as these could be accounted for by the gain or loss of particular restriction sites. The data were analysed using various programs contained in the REAP (McElroy et al., 1991) and the PHYLIP 3.57c (Felsenstein, 1995) computer packages. The significance level in the MONTE program was obtained by 10 000 randomizations. NST (Lynch & Crease, 1990) was used to estimate the degree of population subdivision at the nucleotide level. A Mantel test (NTSYS computer package, Rohlf, 1990) was used to compare distance matrices calculated from allozyme (Triantafyllidis et al., 1999) and mtDNA data.
Results
Restriction analysis
The sizes of PCR-amplified mtDNA segments for both species were ≈ 2.4 kilobases (kb) for ND-5/6 and 2.2 kb for HN20. Routine screening of all populations revealed a total of 130 sites in S. glanis and 127 sites in S. aristotelis corresponding to 538 nucleotides and 536 nucleotides, respectively, ≈ 3% of the mitochondrial genome. Fragment patterns generated by each enzyme are available upon request. Four (AluI, RsaI, HinfI, HhaI) and four (AluI, RsaI, HaeIII, Bst NI) restriction enzymes out of 14 were polymorphic in the ND-5/6 and HN20 regions of S. glanis, whereas the respective values for S. aristotelis were two (HaeIII, HhaI) and four (AluI, HinfI, HhaI, BfaI). No haplotype was shared between individuals from the two species. Thus, for better presentation of the results, composite genotypes and corresponding frequencies for the populations of the two species are presented separately in Tables 1 and Table 2.
RFLP haplotype distribution and diversity
Ten different haplotypes (composite genotypes) were found in S. glanis populations (Table 1). The number of observed haplotypes within populations varied from one to six. Eight out of 10 haplotypes were private, i.e. population-specific. Greek populations were dominated by one or two haplotypes, whereas six haplotypes were found in the Danube Delta population. This was reflected in the values of the haplotype diversity index, which varied from 0.00 in most Greek populations to 0.79 in the Danube Delta population (Table 1). Pairwise sequence divergence estimates among these haplotypes varied from 0.10 to 0.73%.
Within the S. aristotelis populations nine different haplotypes were found, with seven of them being private (Table 2). Pairwise sequence divergence estimates among these haplotypes varied from 0.09 to 0.39%.
Relationship among haplotypes and populations
The test for geographical heterogeneity in haplotype frequencies among all populations, through the MONTE program of the REAP package, resulted in a significant outcome. All 16 pairwise comparisons between natural populations within the species were also significant (P < 0.01). On the other hand, differences between an introduced (Volvi) or a hatchery (Strymon Hatchery) population with its parental (Trichonida and Strymon Natural, respectively) population were not significant.
Nucleotide divergence was in the range of 0.00–0.52% among S. glanis populations, and 0.00–0.11% among S. aristotelis populations, based on restriction site data. Strong geographical patterning was observed in the distribution of the S. glanis mtDNA haplotypes, in both the FITCH (distance) phenogram (not shown) and the majority rule consensus tree of Wagner parsimony trees (Fig. 2). As shown, haplotypes 2g, 3g, 4g and 5g, which are characterized by the patterns AluI B in the ND-5/6 region and AluI B, C and RsaI B in the HN20 region form a cluster. These haplotypes were found in the Greek populations of the Evros, Axios, Aliakmon and Pineios rivers. Bootstrap analysis supports this clustering with 96% confidence. Haplotype 1g was found in the Strymon (Greece) and Danube Delta populations.
Majority rule consensus tree clustering the 10 Silurus glanis mtDNA haplotypes described in Table 1. Numbers at the forks indicate the percentage of times the group consisting of the haplotypes located to the right of the fork occurred among the trees, out of 1000 trees. The tree was rooted with haplotype 6, which is found in the Danube Delta population.
Because of the big differences in fragment profiles, site differences could not be inferred in all cases when comparing the two species. Accordingly, fragment rather than site data were used for the analysis of the relationship of the two species. The average divergence between the two species based on fragment data was 6.75%. The UPGMA dendrogram showing the relationships between the different populations, based on nucleotide divergence values, is presented in Fig. 3. The differentiation of the two species is evident in this dendrogram. For S. glanis, the Evros, Axios, Aliakmon and Pineios populations form one group whereas the rest comprise one second cluster, which is concordant with the previous trees on haplotypes and the UPGMA tree constructed based on sequence divergence values of S. glanis populations only (not shown).
Discussion
The present study has revealed low levels of mtDNA divergence and variation within the two European silurid catfishes. Levels of intraspecific mtDNA haplotype sequence divergence for freshwater fish reported in the literature (Billington & Hebert, 1991) were as high as 10%. However, in our study values of intraspecific mtDNA haplotype sequence divergence did not exceed 0.39% and 0.73% in S. aristotelis and S. glanis, respectively. Many authors (Avise, 1989; Cronin et al., 1993) suggest that this pattern may occur in areas that have recently been colonized following glaciation, because of bottlenecks at the time of population origin.
Ecological characteristics can also play significant roles in determining the distribution and genetic structure of modern-day populations. Wilson & Hebert (1996) also discovered comparably low levels of mtDNA divergence in lake trout (Salvelinus namaycush) populations in eastern North America. Apart from the importance of glaciation events, they emphasized that lake trout as a top predator (as is S. glanis) may be expected to have smaller population sizes than species at lower trophic levels, which could account for low levels of mtDNA variation. However, additional sampling throughout the distribution of S. glanis is required to test if these low diversity values could be considered a species-specific trait.
Despite the low levels of divergence among haplotypes, considerable genetic differentiation was observed among populations. Eight out of 10 haplotypes for S. glanis and seven out of nine haplotypes for S. aristotelis were private. As a result, most populations can easily be distinguished because some samples do not share any haplotypes. This is reflected in the significant outcome of all pairwise comparisons between native populations for geographical heterogeneity of mtDNA haplotype distribution.
The differentiation of the wild S. glanis population is also evident by the NST estimate of 0.84, according to which only about 15% of the overall genetic diversity observed was within populations. This value is higher than those reported for various groups of invertebrates and vertebrates (range 0.03–0.075; compiled in Lynch & Crease, 1990). However, such high NST values have been reported after RFLP analysis of the mtDNA diversity of two more Greek freshwater fish: Salmo trutta (Apostolidis et al., 1996) and Leuciscus cephalus (Imsiridou et al., 1998). Long-time isolation of the populations coupled with bottleneck and subsequent genetic drift phenomena, which resulted in stochastic loss of haplotypes, were proposed as a possible explanation.
This strong geographical patterning in the distribution of the mtDNA haplotypes is evident in all trees constructed from relationships either of haplotypes or of populations (Figs 2 and 3). The populations (or haplotypes) of Evros, Axios, Aliakmon and Pineios rivers seem to form one monophyletic group, which diverged from the rest and has remained isolated. The Strymon population does not cluster with the rest of the Greek populations. Loss of haplotypes, characteristic of the other populations, because of stochastic forces and/or introgression of the Danubian origin haplotype 1g, after movement of fish from the Danube through river captures or human influence, could explain this pattern.
Levels of congruence between mtDNA and allozyme variation data
MtDNA haplotype diversity is considered equivalent to heterozygosity of protein polymorphism (Nei, 1987) and as such it is interesting to note that there was agreement with an allozyme variation analysis on the same populations (Triantafyllidis et al., 1999). In both studies, the central European population of the Danube Delta showed the highest value (Table 1). Additionally, the three Greek S. glanis populations (of the Pineios, Strymon and Evros rivers) with zero haplotype diversity also exhibited the lowest values of observed (allozyme) heterozygosity. Congruence was also noticed for S. aristotelis populations (Table 2).
However, greater differentiation between populations was observed at the mtDNA level than at nuclear loci. Allozyme analysis did not reveal alleles which were characteristic of populations or, if it did, these alleles were of low frequency (<0.05). On the other hand, many mtDNA haplotypes were private for some populations, even fixed in certain cases. This pattern has been reported before (Taylor et al., 1997). It has been associated with the heightened sensitivity of mtDNA as a marker of population structure, because of its fourfold lower effective population size. Consequently, in populations of recent origin the effects of genetic drift are first evident at the mtDNA level.
Another discrepancy between the two methods regards relationships between populations as indicated by genetic distance. The Mantel test, employing 10 000 permutations, showed that there is no correlation between the genetic distance matrices calculated with the two methods (r = – 0.021, P = 0.48). Differences were also observed in the phylogenetic relationships between populations, because the Strymon population does not cluster with the rest of the Greek populations in the present analysis. Discordance between the two methodologies has been reported before (e.g. Degnan, 1993; Bernatchez & Osinov, 1995). Therefore, this study emphasizes once more the need for multidisciplinary genetic analyses (Triantaphyllidis et al., 1997).
Taxonomic and phylogenetic relationship between the two species
Average sequence divergence between the two species based on fragment data is 6.75%. When fragment data were used to estimate the genetic divergence within species, the values obtained were ≈20% lower than those obtained with restriction site data. Some authors (Bernatchez et al., 1991) also suggest that at above 5% divergence, undetectable homoplasy in either restriction site or fragment data analyses leads to an underestimation of the real value. Thus, the estimated value of 6.75% divergence should be considered a minimum. Even so, the value should be within the range of 1.4–16% reported for congeneric freshwater fish (Billington & Hebert, 1991). Therefore, our data support previous morphological studies on catfish (Kobayakawa, 1989; Bornbusch, 1995) according to which the genus Parasilurus should be synonymized with Silurus.
Estimating the time of divergence between phylogenetic assemblages requires certain assumptions on which there is not always agreement. For instance, Bernatchez et al. (1991) assume a rate of 2% nucleotide divergence per million years for Coregonus fish, whereas Martin & Palumbi (1993) propose a rate of 0.5–0.9% for salmonid fish. Even if the most conservative value is used, the time of divergence calculated is sometime during the Middle to Late Miocene. Our results are concordant with other authors’ views (Bianco, 1990; Economidis & Banarescu, 1991) that the ancestral Silurus population was established in central Europe at around that time.
Silurus aristotelis is found only in western Greece. The fish fauna of western Greece comprises species that are believed to have reached the area and remained isolated there since Miocene or at least early Pliocene times (Economidis & Banarescu, 1991). It is proposed therefore that the ancestors of S. aristotelis dispersed from the original Silurus stock in central Europe, somewhere around that time, and moved south reaching Greece. Based on the low divergence values between S. glanis haplotypes, S. glanis seems to have spread in Europe later, during the Pleistocene.
The results of this study suggest that protection measures and a conservation programme of the two silurid species be undertaken, for both financial and ecological reasons. First, a number of private haplotypes, which were sometimes fixed in Greek populations, were revealed. These haplotypes can serve as ‘genetic tags’ for stock identification in future aquaculture programmes. Secondly, low levels of mtDNA diversity were found, a result in agreement with previous allozyme analysis. This impoverishment of genome variability could be a threat for the future survival of the species.
References
Apostolidis, A. P., Karakousis, Y. and Triantaphyllidis, C. (1996). Genetic differentiation and phylogenetic relationships among Greek Salmo trutta L. (brown trout) populations as revealed by RFLP analysis of PCR amplified mitochondrial DNA segments. Heredity, 77: 608–618.
Avise, J. C. (1989). Gene trees and organismal histories: a phylogenetic approach to population biology. Evolution, 43: 1192–1208.
Avise, J. C. (1994). Molecular Markers, Natural History and Evolution. Chapman and Hall, New York.
Banarescu, P. M. (1989). Zoogeography and history of the freshwater fish fauna of Europe In: Holcik, J. (ed.). The Freshwater Fishes of Europe , I/II: 88–107. Aula Verlag, Wiesbaden.
Bernatchez, L. and Osinov, A. (1995). Genetic diversity of trout (genus Salmo) from its most eastern native range based on mitochondrial DNA and nuclear gene variation. Mol Ecol, 4: 285–297.
Bernatchez, L., Colombani, F. and Dodson, J. J. (1991). Phylogenetic relationships among the subfamily Coregoninae as revealed by mitochondrial DNA restriction analysis. J Fish Biol(Suppl A), 39: 283–290.
Bernatchez, L., Glemet, H., Wilson, C. C. and Danzmann, R. G. (1995). Introgression and fixation of Arctic charr (Salvelinus alpinus) mitochondrial genome in an allopatric population of brook charr (Salvelinus fontinalis). Can J Fish Aquat Sci, 52: 179–185.
Bianco, P. G. (1990). Potential role of the paleohistory of the Mediterranean and Paratethys basins on the early dispersal of Euromediterranean freshwater fishes. Ichthyol Explor Freshw, 1: 167–184.
Billington, N. and Hebert, P. D. N. (1991). Mitochondrial DNA diversity in fishes and its implications for introductions. Can J Fish Aquat Sci, 48: 80–94.
Bleeker, P. (1862). Notice sur les generes Parasilurus Eutropiichthys Eudeutropius et Pseudopangasius. Versl Akad Amsterdam, 14: 390–399.
Bornbusch, A. H. (1995). Phylogenetic relationships within the Eurasian catfish family Siluridae (Pisces: Siluriformes), with comments on generic validities and biogeography. Zool J Linn Soc, 115: 1–46.
Chombard, C. (1993). Etude des Variations Electrophoretiques de Populations de Silure Glane, Silurus glanis (L. 1758). DEA de Biodiversité, Génétique et Évolution, Paris.
Cronin, M., Spearman, W., Wilmot, R., Patton, J. and Bickham, J. (1993). Mitochondrial DNA variation in chinook (Oncorhynchus tshawytscha) and chum (O. keta) detected by restriction enzyme analysis of polymerase chain reaction (PCR) products. Can J Fish Aquat Sci, 50: 708–715.
Degnan, S. M. (1993). The perils of single gene trees –mitochondrial versus single-copy nuclear DNA variation in white eyes (Aves: Zosteropidae). Mol Ecol, 2: 219–225.
Economidis, P. S. and Banarescu, P. M. (1991). The distribution and origins of freshwater fishes in the Balkan Peninsula, especially in Greece. Int Rev ges Hydrobiol, 76: 257–283.
Felsenstein, J. (1995). PHYLIP(Phylogeny inference package), version 3.57c. Department of Genetics, SK-50, University of Washington, Seattle, WA.
Imsiridou, A., Apostolidis, A. P., Durand, J. D., Briolay, J., Bouvet, Y. and Triantaphyllidis, C. (1998). Genetic differentiation and phylogenetic relationships among Greek chub Leuciscus cephalus L. (Pisces, Cyprinidae) populations as revealed by RFLP analysis of mitochondrial DNA. Biochem Syst Ecol, 26: 415–429.
Kobayakawa, M. (1989). Systematic revision of the catfish genus Silurus with description of a new species from Thailand and Burma. Jap J Ichthyol, 36: 155–186.
Kotsinas, A., Karakousis, Y. and Triantaphyllidis, C. (1989). Biometric and isozymic comparison of the species Silurus glanis and Silurus aristotelis (Pisces, Siluridae).. Proceedings of the 11th Congress of the Biol. Soc. of Greece, Komotini. 138–139. (in Greek with English summary).
Lynch, M. and Crease, T. J. (1990). The analysis of population survey data on DNA sequence variation. Mol Biol Evol, 7: 377–394.
Martin, A. P. and Palumbi, S. R. (1993). Body size, metabolic rate, generation time and the molecular clock. Proc Natl Acad Sci USA, 90: 4087–4091.
McElroy, D., Moran, P., Bermingham, E. and Kornfield, J. (1991). The restriction enzyme analysis package. version 4.0. University of Maine, Orono, ME.
Nei, M. (1987). Molecular Evolutionary Genetics.. Columbia University Press, New York.
Rab, P., Karakousis, Y. and Peios, C. (1994). Karyotype of Silurus aristotelis with reference to the cytotaxonomy of the genus Silurus (Pisces, Siluridae). Fol Zool, 43: 75–81.
Rohlf, F. J. (1990). NTSYS -pc. Numerical taxonomy and multivariate analysis system. Exeter Software, New York.
Taggart, J. B., Hynes, R. A., Prodohl, P. A. and Ferguson, A. (1992). A simplified protocol for routine total DNA isolation from salmonid fishes. J Fish Biol, 40: 963–965.
Taylor, E. B., Harvey, S., Pollard, S. and Volpe, J. (1997). Postglacial genetic differentiation of reproductive ecotypes of kokanee Oncorhynchus nerka in Okanagan Lake, British Columbia. Mol Ecol, 6: 503–517.
Triantafyllidis, A., Ozouf-Costaz, C., Rab, P., Suciu, R. and Karakousis, Y. (1999). Allozyme variation in European silurid catfishes Silurus glanis and Silurus aristotelis. Biochem Syst Ecol. in press.27
Triantaphyllidis, G. V., Criel, G. R. J., Abatzopoulos, T. J., Thomas, K. M., Peleman, J., Beardmore, J. A. and Sorgeloos, P. (1997). International study on ArtemiaLVII. Morphological and molecular characters suggest conspecificity of all bisexual European and North African Artemia populations. Mar Biol, 129: 477–487.
Wilson, C. C. and Hebert, P. D. N. (1996). Phylogeographic origins of lake trout (Salvelinus namaycush) in eastern North America. Can J Fish Aquat Sci, 53: 2764–2775.
Acknowledgements
The authors would like to thank Prof. C. Triantaphyllidis and Dr A. Apostolidis for their help and guidance throughout this work, Dr R. Guyomard and Dr F. Krieg for their collaboration, Dr Z. Scouras and Dr A. Kouvatsi for their fruitful discussions, Dr R. Suciu and all the fishermen for providing samples, and finally two anonymous reviewers for their valuable comments. Financial support provided by the European Commission within the framework ERVCIPACT930274 project is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Triantafyllidis, A., Abatzopoulos, T. & Economidis, P. Genetic differentiation and phylogenetic relationships among Greek Silurus glanis and Silurus aristotelis (Pisces, Siluridae) populations, assessed by PCR–RFLP analysis of mitochondrial DNA segments. Heredity 82, 503–509 (1999). https://doi.org/10.1038/sj.hdy.6885140
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/sj.hdy.6885140
Keywords
This article is cited by
-
Whole genome sequencing reveals high differentiation, low levels of genetic diversity and short runs of homozygosity among Swedish wels catfish
Heredity (2021)
-
Genetic diversity and population genetic structure of Wels (Silurus glanis Linnaeus, 1758) in the northwest of Iran
Environmental Biology of Fishes (2015)
-
Genetic differentiation among Greek lake populations of Carassius gibelio and Cyprinus carpio carpio
Genetica (2009)
-
Phylogeography and genetic structure of the Mediterranean killifish Aphanius fasciatus (Cyprinodontidae)
Marine Biology (2007)
-
Genetic Structure and Low-genetic Diversity Suggesting the Necessity for Conservation of the Chinese Longsnout Catfish, Leiocassis longirostris (Pisces: Bagriidae)
Environmental Biology of Fishes (2006)