Abstract
Two different genotypes of diploid Hordeum vulgare × H. bulbosum hybrids, which differ in their pattern of meiotic metaphase pairing behaviour, were investigated at MI and AI by genomic in situ hybridization (GISH). One hybrid, 102C2, showed a high frequency of bivalents at metaphase I whereas the other, 103K5, showed a high frequency of univalents. The GISH analysis of both hybrids established that pairing occurred only between chromosomes of different parental genomes and revealed that pairing frequency greatly exceeded recombination. Hybrid 102C2 had a significantly higher recombination frequency than 103K5, but in both hybrids recombination involved only distal chromosome regions. However, an interesting finding is that the ratio of recombination to pairing frequency in 103K5 (1:8.9) is twice as high compared with 102C2 (1:17). The hybrids also differed in chromosome stability; little chromosome elimination occurred in 102C2 but 103K5 showed extensive chromosome loss. It appears that the high frequency of bound arms at MI favours retention of H. bulbosum chromosomes and maintains stability of chromosome numbers in PMCs. Various ideas are put forward to explain the discrepancy between meiotic pairing frequency and recombination in these hybrids.
Similar content being viewed by others
Introduction
Diploid and tetraploid cytotypes of Hordeum bulbosum L. (2n=2x=14 and 2n=4x=28) are valuable sources of pest- and disease-resistance genes, which have been successfully transferred into cultivated barley (H. vulgare L.) (Xu & Kasha, 1992; Pickering et al., 1995). To achieve this, partially fertile triploid hybrids from the cross between cultivated barley (2n=2x=14) and H. bulbosum (2n=4x=28) were backcrossed to barley, and recombinants were produced that contained small segments of H. bulbosum DNA introgressed into the H. vulgare genome (Pickering et al., 1997). However, the number of recombinants and ‘chromosomally engineered’ plants obtained from crosses between the two species is very low (Lange & Jochemsen, 1975; Pickering, 1992). The reasons for this have been outlined by Pickering (1992) and briefly they are: (i) barley is a diploid species that cannot tolerate much genetic manipulation; (ii) the H. bulbosum genome is usually completely eliminated, resulting in haploid barley embryo and plantlet formation; (iii) the interspecific hybrids are often sterile and unstable; and (iv) intergenomic chromosome pairing is variable and genotype-dependent. Apart from the first problem, which is inherent for diploid species, the second and third of these barriers can easily be overcome by manipulating the parental genotypes and crossing environment. However, there still exists the more serious problem of recombination frequencies that are lower than expected in these hybrids (Pickering, 1991), a feature that is seen among interspecific crosses in other genera (Rick, 1969; Causse et al., 1994). Determining the reasons for low frequencies of recombinant progeny from H. vulgare × H. bulbosum hybrids has been difficult because conventional cytogenetic analyses are not sufficiently refined to draw meaningful conclusions. The use of a paracentric inversion in a diploid H. vulgare × H. bulbosum hybrid to assess recombination frequencies, by recording the numbers of bridges and fragments at meiotic anaphase (Pickering, 1991), was not efficient because the results only related to one particular chromosome and ‘hotspots’ of crossing-over are likely to occur throughout the genome (Pedersen et al., 1995).
Because genomic in situ hybridization (GISH) has been used successfully to identify parental chromosomes in H. vulgare × H. bulbosum hybrids (Schwarzacher et al., 1992b; Anamthawat-Jónsson et al., 1993) and also to visualize introgression of H. bulbosum chromatin into H. vulgare (Pickering et al., 1997), the objectives in this investigation were to use GISH to assess the extent of crossing-over between parental chromosomes by observations of anaphase I (AI) in pollen mother cells (PMCs) of two diploid H. vulgare × H. bulbosum hybrids, which differed in their chromosome pairing at metaphase I (MI) of meiosis. In addition, observations on chromosome segregation at AI in PMCs, the frequency of H. bulbosum chromosome elimination and any possible relationship between H. bulbosum chromosome elimination and MI pairing were made.
Materials and methods
Two different hybrids that have a common female parent (H. vulgare cv. Emir) were used, 102C2 (Emir × HB2032) and 103K5 (Emir × Cb3811/5). These hybrids were chosen because earlier observations had shown that the extent of their chromosome pairing at MI was significantly different, with 102C2 having a much higher MI pairing frequency than 103K5.
Growing conditions in the glasshouse were maintained at 21 ± 2°C/15 ± 2°C (day/night), with 16 h daylength. Anthers containing PMCs at MI or AI were fixed in Carnoy’s II solution (6 parts ethanol:3 parts chloroform:1 part glacial acetic acid) overnight at 4°C, then transferred to 70% ethanol and stored at 4°C until required.
For GISH, meiotic chromosome preparations from anthers of hybrids were made as follows: anthers containing the appropriate stage were digested in 4% cellulase (Onozuka R-10, Merck) and 1.3% pectolyase (P-3026, Sigma) for 90 min at 37°C. After the enzyme mix was removed with small pieces of filter paper, the anthers were washed twice with distilled water and the contents squeezed out in one or two drops of fresh fixative (3 parts ethanol: 1 part glacial acetic acid) and spread evenly on clean glass slides assisted by a gentle stream of air, then left to air dry. Slides were stored at −20°C until needed.
Genomic DNA, isolated from young leaves of H. bulbosum, was labelled with digoxigenin-11-dUTP (Boehringer Mannheim) by nick-translation according to the manufacturer’s instructions. Total genomic DNA from H. vulgare was broken into 200–500 bp fragments by autoclaving and used as blocking DNA. The method of in situ hybridization followed Schwarzacher et al. (1992a) with minor modifications. Forty-five μL of denatured probe mixture, containing 100 ng of DIG-labelled probe and a 50-fold excess of H. vulgare blocking DNA, was applied to each slide. The slides were then heated to 80°C for 10 min and the temperature lowered to 37°C for incubation overnight on a thermal cycler (Hybaid, Omnislide). The hybridization mix consisted of 55% formamide, 2 × SSC, 10% (w/v) dextran sulphate, 0.15% SDS and 150-fold excess of salmon sperm DNA (200–500 bp). Hybridization was followed by a stringent wash in 50% formamide in 2 × SSC at 42°C. Sites of probe hybridization were detected using the HNPP Fluorescent Detection Set (Boehringer Mannheim). Chromosomes were counterstained with DAPI (1 μg/mL, 4′,6′-diamidino-2-phenylindole) in 1 × PBS buffer. Slides were mounted in antifade mountant (1.8% DABCO in 90% glycerol v/v, 20 mM Tris pH 7.5) and examined under a Zeiss epi-fluorescence photomicroscope. Photographs were taken on Fujichrome colour reversal film (Provia 400), digitized using a Nikon LS-1000 scanner, and the images processed using Adobe Photoshop 3.0.5 software.
Results
GISH analysis of pairing at MI and recombinant chromosomes at AI in hybrids
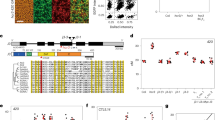
Using GISH it was possible to distinguish clearly H. vulgare and H. bulbosum chromosomes and to show that pairing always occurred between chromosomes of H. vulgare and H. bulbosum. However, the pattern of chromosome pairing in the two hybrids was quite different. There was significantly greater chromosome pairing in hybrid 102C2 than in 103K5 (Table 1; Fig. 1a, b, c, d, e). The former had a mean bivalent frequency of 6.68 per PMC with a range of 5–7 bivalents per PMC, compared with 2.64 bivalents per PMC with a range of 0–5 for the latter. There were also differences in the types of bivalent formed. In 102C2, 35% of the bivalents were ring-shaped (with two bound arms) whereas in 103K5 only 11.7% were ring-shaped. From this it can be seen that the frequency of bound arms was significantly greater in 102C2 than in 103K5 (χ21=412.1, d.f.=1, P < 0.001).
GISH using Hordeum bulbosum DNA as probe on meiotic metaphase I (a – e) and anaphase I (f, g) chromosome preparations in PMCs of the barley × H. bulbosum hybrids 102C2 and 103K5. The labelled H. bulbosum chromosomes appear red and barley chromosomes are blue with the DAPI counterstain. Arrowheads indicate recombinant barley chromosomes. Arrows indicate recombinant H. bulbosum chromosomes. (a) Hybrid 102C2 with four ring and two rod bivalents; (b) hybrid 102C2 with two ring and five rod bivalents; (c) a 12 chromosome cell of hybrid 103K5 with two rod bivalents; (d) a 13 chromosome cell of hybrid 103K5 with two rod bivalents; (e) complete chromosome complement of hybrid 103K5 at MI with one ring and two rod bivalents; (f) hybrid 102C2 with four recombinant barley chromosomes (arrowheads) and three recombinant H. bulbosum chromosomes (arrows); (g) hybrid 103K5 with one recombinant barley chromosome (arrowhead). Bars represent 10 μm.
Observations on AI cells showed that recombination frequency was significantly higher in 102C2 than in 103K5 (χ21=7.13, P < 0.01) (Table 2; Fig. 1f, g). In 103K5, only one or two recombinant chromosome arms were seen in any single PMC and 74% of PMCs showed no recombinant arms. In 102C2, the number of PMCs without recombinant arms was similar (71%) but the PMCs that did show recombination had significantly more recombinant arms. One PMC with as many as seven recombinant arms was seen and the mean frequency of recombinant arms in PMCs showing recombination was 1.84 compared to 1.24 in 103K5.
Pairing frequency was much higher than recombination frequency in both hybrids (χ21=2107.6, P < 0.001 and χ21=249.8, P < 0.001 for 102C2 and 103K5, respectively) and a comparison of the mean ratios per PMC of the recombinant arms at AI to the bound arms at MI showed that this was twice as high in 103K5 (1:8.9) compared with 102C2 (1:17).
In both hybrids, synapsed chromosome arms were bound predominantly at distal regions and the recombination events involved only small terminal segments (Fig. 1f, g). No evidence of recombination from double crossovers was observed. Hybrid 102C2 showed an ordered arrangement of bivalents at MI and of segregation at AI whereas 103K5 showed irregular chromosome congression and little evidence of polar migration of chromosomes.
Chromosome elimination in PMCs of hybrids
Differences in chromosome number were seen among PMCs of both hybrids and these only involved the loss of H. bulbosum chromosomes. However, there were clear differences in the frequency of chromosome loss in the two hybrid combinations. Elimination of one or two H. bulbosum chromosomes was seen in ≈50% of 173 PMCs of hybrid 103K5 (Fig. 1c, d) but elimination was only rarely observed in PMCs of 102C2. PMCs of 103K5 could be classified into three types: type A (48.6%) with the complete chromosome complement (7 H. vulgare chromosomes + 7 H. bulbosum chromosomes or 7V + 7B); type B (38.7%) with 13 chromosomes (7V + 6B) and type C (12.7%) with 12 chromosomes (7V + 5B). Significant differences were found for the mean number of bound arms per PMC between types A and B (χ21=18.0, P < 0.001) and between types A and C (χ21=13.0, P < 0.001), whereas the difference between types B and type C was not significant.
Discussion
Apart from one report of homoeologous chromosome pairing in PMCs of diploid and triploid H. vulgare × H. bulbosum hybrids using C-banding techniques (Xu & Snape, 1988), conventional meiotic analysis has not been adopted as an efficient method to establish homoeology between H. vulgare and H. bulbosum chromosomes and to study recombination frequency in hybrids between the two species. However, genomes from H. vulgare and H. bulbosum, which are genetically very closely related, can be differentiated by GISH on root-tip chromosome preparations (Schwarzacher et al., 1992b; Anamthawat-Jónsson et al., 1993). GISH has also been shown to be an efficient technique for the study of homoeologous chromosome pairing and recombination in wheat–rye hybrids (Le & Armstrong, 1991; King et al., 1994; Fernández-Calvín et al., 1995; Benavente et al., 1996, 1998), and in other interspecific and intergeneric hybrids (Parokonny et al., 1997; King et al., 1999). However, GISH has not been used previously for studying meiotic behaviour in diploid H. vulgare × H. bulbosum hybrids. GISH is therefore a valuable additional technique to conventional genome analysis, and has enabled us to readily clarify some aspects of the reduced recombination in hybrids between H. vulgare and H. bulbosum.
Our observations have confirmed the lower level of meiotic pairing in hybrid 103K5 and shown that there is a higher frequency of recombination as measured by recombinant H. vulgare chromosomes at AI in hybrid 102C2. In both hybrids there was more pairing at MI compared with the recombination frequency at AI and there are at least three possible explanations for this discrepancy (see also Benavente et al., 1996, 1998). First, there may be a limitation in the ability of GISH detection in the degree of resolution of fluorescence microscopy on meiotic chromosome preparations at AI, as chromosomes of these hybrids at AI are more highly condensed than at MI. Secondly, the size of introgressed segments is related to their ease of detection. Jiang & Gill (1994) reported that in the successful cases of mapping low- or single-copy sequences, the targeted DNA sequences on chromosomes with reliable frequency of signal detection were all more than 10 kb long. Thus small introgressed segments may go undetected. The third possible explanation to account for this discrepancy is that a majority of bound arms between chromosomes of different parental genomes at MI in hybrids may be attributable to remnants of prophase pairing which is not related to crossing-over. We believe that the third explanation is the most likely and is consistent with the observations in a study of wheat–rye hybrids by Orellana (1985), and if proven, would establish that the paucity of recombinant progeny from H. vulgare × H. bulbosum hybrids results largely from low levels of crossing-over between chromosomes of different parental genomes.
In 102C2 and 102K5, rod bivalents predominate (Tables 1 and Table 3), but even in 102C2, which had almost complete bivalent formation, there is less chromosome pairing at MI than in cultivated barley. In barley seven ring bivalents are usually observed in most cells (Nilsson & Pelger, 1991). Recombinant segments at AI confirmed that crossing-over occurs distally, as is the case in barley (Pedersen et al., 1995), and suggested that there was a reduced recombination in the proximal regions of the chromosome arms. GISH results on root-tip chromosome preparations of recombinants from the progeny of H. vulgare × H. bulbosum hybrids also revealed that introgressions of H. bulbosum chromatin are located distally (Pickering et al., 1997; Pickering, unpubl. obs.).
Although the high-pairing hybrid 102C2 showed higher recombination frequency than the low-pairing hybrid 103K5, when a comparison is made of the recombination frequency in comparison to the number of bound arms at MI, recombination frequency is then higher in 103K5 than 102C2. In other words, when the chromosomes of the hybrids are paired at MI, they are then more likely to undergo recombination in 103K5 than 102C2. This seems at variance to what might have been expected and at present we have no ready explanation for the phenomenon. It is possible that the localization of chiasmata is less precisely controlled in 103K5, which results in the formation of more interstitial chiasmata and the exchange of larger segments, which are then more readily detected by GISH (see above).
We have also found that of the three types of PMCs in 103K5, type A had a significantly higher proportion of bound arms than types B and C, as would be expected, but there was no significant difference in the number of bound arms between types B and C. It is possible therefore that a high formation of bound arms promotes the retention of H. bulbosum chromosomes and maintains the stability of chromosome numbers and pairing. Support for this suggestion came from the analysis of 90 different genotypes of diploid H. vulgare × H. bulbosum hybrids (Pickering, unpubl. obs.), where a negative correlation (r=−0.528, P < 0.01) between the number of univalents and mean chromosome number was found.
In summary, the occurrence of chromosome pairing provides a basis for recombination in H. vulgare × H. bulbosum hybrids, albeit at a lower frequency than in H. vulgare itself. Because high meiotic pairing seems to be associated with high recombination in the hybrids, selection of high-pairing hybrids has resulted in the development of recombinant barley plants with introgressed H. bulbosum chromatin. Several of these recombinants possess resistance to fungal and/or viral pathogens and will provide breeders with a novel source of resistance genes.
References
Anamthawat-Jónsson, K., Schwarzacher, T. and Heslop-Harrison, J. S. (1993). Behaviour of parental genomes in the hybrid Hordeum vulgare× H. bulbosum. J Hered, 84: 78–82.
Benavente, E., Fernández-Calvín, B. and Orellana, J. (1996). Relationship between the levels of wheat–rye metaphase I chromosomal pairing and recombination revealed by GISH. Chromosoma, 105: 92–96.
Benavente, E., Orellana, J. and Fernández-Calvín, B. (1998). Comparative analysis of the meiotic effects of wheat ph1b and ph2b mutations in wheat × rye hybrids. Theor Appl Genet, 96: 1200–1204.
Causse, M. A., Fulton, T. M., Cho, Y. G., Ahn, S., Chunwongse, J., Wu, K. et al (1994). Saturated molecular map of the rice genome based on an interspecific backcross population. Genetics, 138: 1251–1274.
Fernández-Calvín, B., Benavente, E. and Orellana, J. (1995). Meiotic pairing in wheat-rye derivatives detected by genomic in situ hybridization and C-banding – A comparative analysis. Chromosoma, 103: 554–558.
Jiang, J. and Gill, B. S. (1994). Nonisotopic in situ hybridization and plant genome mapping: the first 10 years. Genome, 37: 717–725.
King, I. P., Reader, S. M., Purdie, K. A., Orford, S. E. and Miller, T. E. (1994). A study of the effect of a homoeologous pairing promoter on chromosome pairing in wheat/rye hybrids using genomic in situ hybridization. Heredity, 72: 318–321.
King, I. P., Morgan, W. G., Harper, J. A. and Thomas, H. M. (1999). Introgression mapping in the grasses. II. Meiotic analysis of the Lolium perenne/Festuca pratensis triploid hybrid. Heredity, 82: 107–112.
Lange, W. and Jochemsen, G. (1975). The offspring of diploid, triploid and tetraploid hybrids between Hordeum vulgare and H. bulbosum. In: Proc. Int. Barley Genet. Symposium III 252–259.
Le, H. T. and Armstrong, K. C. (1991). In situ hybridization as a rapid means to assess meiotic pairing and detection of alien DNA transfers in interphase cells of wide crosses involving wheat and rye. Mol Gen Genet, 225: 33–37.
Nilsson, N. -O. and Pelger, S. (1991). The relationship between natural variation in chiasma frequencies and recombination frequencies in barley. Hereditas, 115: 121–126.
Orellana, J. (1985). Most of the homoeologous pairing at metaphase I in wheat-rye hybrids is not chiasmatic. Genetics, 111: 917–931.
Parokonny, A. S., Marshall, J. A., Bennett, M. D., Cocking, E. C., Davey, M. R. and Power, J. B. (1997). Homoeologous pairing and recombination in backcross derivatives of tomato somatic hybrids [Lycopersicon esculentum(+) L. peruvianum]. Theor Appl Genet, 94: 713–723.
Pedersen, C., Giese, H. and Linde-Laursen, I. (1995). Towards an integration of the physical and the genetic chromosome maps of barley by in situ hybridization. Hereditas, 123: 77–88.
Pickering, R. A. (1991). Comparison of crossover frequencies in barley (Hordeum vulgare) and H. vulgare× H. bulbosum hybrids using a paracentric inversion. Genome, 34: 666–673.
Pickering, R. A. (1992). Monosomic and double monosomic substitutions of Hordeum bulbosum L. chromosomes into H. vulgare L. Theor Appl Genet, 84: 466–472.
Pickering, R. A., Hill, A. M., Michel, M. and Timmerman-Vaughan, G. M. (1995). The transfer of a powdery mildew resistance gene from Hordeum bulbosum L. to barley (H. vulgare L.) chromosome 2 (2I). Theor Appl Genet, 91: 1288–1292.
Pickering, R. A., Hill, A. M. and Kynast, R. G. (1997). Characterization by RFLP analysis and genomic in situ hybridization of a recombinant and a monosomic substitution plant derived from Hordeum vulgare L. × H. bulbosum L. crosses. Genome, 40: 195–200.
Rick, C. M. (1969). Controlled introgression of chromosomes of Solanum pennellii into Lycopersicon esculentum: segregation and recombination. Genetics, 62: 753–768.
Schwarzacher, T., Anamthawat-Jónsson, K., Harrison, G. E., Islam, A. K. M. R., Jia, J. Z., King, I. P. et al (1992a). Genomic in situ hybridization to identify alien chromosomes and chromosome segments in wheat. Theor Appl Genet, 84: 778–786.
Schwarzacher, T., Heslop-Harrison, J. S., Anamthawat-Jónsson, K., Finch, R. A. and Bennett, M. D. (1992b). Parental genome separation in reconstructions of somatic and premeiotic metaphase of Hordeum vulgare× H. bulbosum. J Cell Sci, 101: 13–24.
Xu, J. and Kasha, K. J. (1992). Transfer of a dominant gene for powdery mildew resistance and DNA from Hordeum bulbosum into cultivated barley (H. vulgare). Theor Appl Genet, 84: 771–777.
Xu, J. and Snape, J. W. (1988). The cytology of hybrids between Hordeum vulgare and H. bulbosum revisited. Genome, 30: 486–494.
Acknowledgements
A grant from the University of Auckland Graduate Research Fund is gratefully acknowledged. We also thank Karen Sheath and Marc Jacobs for technical advice and Joshua Salter for helpful comments on an earlier version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, L., Pickering, R. & Murray, B. Direct measurement of recombination frequency in interspecific hybrids between Hordeum vulgare and H. bulbosum using genomic in situ hybridization. Heredity 83, 304–309 (1999). https://doi.org/10.1038/sj.hdy.6885710
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/sj.hdy.6885710
Keywords
This article is cited by
-
Crossover patterning in plants
Plant Reproduction (2023)
-
High-density linkage mapping in a pine tree reveals a genomic region associated with inbreeding depression and provides clues to the extent and distribution of meiotic recombination
BMC Biology (2013)
-
A comparison of recombination frequencies in intraspecific versus interspecific mapping populations of Nasonia
Heredity (2010)
-
GISH and BAC-FISH study of apomictic Beta M14
Science in China Series C: Life Sciences (2007)
-
Mapping of Rym16 Hb , the second soil-borne virus-resistance gene introgressed from Hordeum bulbosum
Theoretical and Applied Genetics (2006)