Abstract
Despite the urgent need to conserve domesticated plant genetic resources, and developing ‘on farm’ strategies of conservation, the impact of traditional farming practices and of their interaction with ecological factors on the structure and evolutionary dynamics of the genetic variability of crop populations has been little documented. We assessed the genetic variability of 31 varieties of cassava (M. esculenta Crantz) traditionally grown by Makushi Amerindians from Guyana, using AFLP markers. We used a sample of 38 varieties from an ex situ core collection as a reference. Accessions of wild cassava were also included. While clonality of the varieties was expected due to the vegetative propagation of cassava, 21 varieties presented intravarietal polymorphism. Among the varieties from a single site in Guyana, genetic diversity was the same as that in the accessions from the core collection. We suggest that incorporation of volunteer seedlings, produced by sexual reproduction, into the stock of varieties grown by the Makushi plays a major role in explaining both intravarietal polymorphism and the high level of genetic diversity. No correspondence was found between the structure of molecular diversity and variation observed for agronomic traits that are targets for selection by cultivators. As found in previous studies, all wild forms of cassava clustered together and separately from the cultivated varieties in a Neighbour-Joining dendrogram. These results are consistent with the hypothesis of a limited domestication event in a restricted area, followed by rapid diffusion of cultivated phenotypes and convergent evolution. Our results show that local varieties are an important source of genetic diversity, and highlight the importance of the interaction between human and ecological factors in the dynamics of this diversity.
Similar content being viewed by others
Introduction
Defining the scientific bases from which efficient strategies for conserving biological diversity can be developed, still remains a challenge for the scientific community. A wide range of theoretical and empirical knowledge about wild plant and animal populations, has led to conservation strategies, but equivalent information is missing for populations of domesticated crops. Defining plans for ‘on farm’ or in situ conservation of plant genetic resources requires assessment of the pattern of both molecular and phenotypic diversity and of processes that drive evolution of crop populations (Miller et al., 1995). Traditional agrosystems are of particular interest because crop diversity within them is generally high. It is common to find numerous varieties in the same field. Moreover, related and interfertile wild forms sometimes occur sympatrically. Such situations offer opportunities for studying the effects of gene flow, disruptive selection and genetic drift in crop populations, which often are enmeshed in crop/wild/weed complexes (De Wet & Harlan, 1975). Conservation of traditional landraces must take into account the peculiarities of man-made environments, in which human action not only shapes landscapes but also is an important selective factor acting on populations of landraces.
Our research programme aims to evaluate the impact of traditional agricultural practices on varietal and genetic diversity, using cultivation of cassava (Manihot esculenta) by Amerindians as an experimental model. Cassava, a vegetatively propagated crop, is widely cultivated in traditional farming systems in Amazonia (McKey & Beckerman, 1993; Salick et al., 1997; Emperaire et al., 1998) where it exhibits a huge varietal and genetic diversity (Colombo, 1997; Second et al., 1997). This paper reports an assessment, using molecular markers, of the organization of genetic variability of cassava landraces in a single village of Makushi Amerindians in Guyana, South America.
Cassava is the Makushis’ staple crop. Makushi Amerindians practice slash-and-burn agriculture, with a cycle of two years of cultivation (2 crops) followed by one year of fallow. Every year, each farmer prepares a new field from secondary forest or fallow, and burns it. Cassava is propagated by means of stem cuttings, which are planted in the new field (first crop). In this study, AFLP markers were used to characterize the genotypes of a sample of the varieties present in the village studied, in order to: (i) assess the level of intravarietal diversity; (ii) study the relationship between the organization of genetic variability and the local taxonomy, and also the relationship between genetic variability and the diversity of four agronomic traits (degree of bitterness, colour of the root, dryness of the root and length of cultivation cycle), which are targets for selection by farmers; (iii) compare the diversity found in the village with that of a sample from a worldwide ex situ core collection [from the International Center for Tropical Agriculture (CIAT, Cali, Colombia)]; (iv) assess the level of genetic similarity of the local varieties and local and exogenous wild forms of cassava.
Materials and methods
Plant material from Guyana and sampling procedure
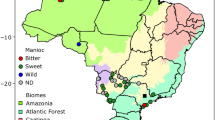
Varietal diversity and traditional farming practices were studied in Rewa, a 30 household Makushi village in Guyana (Fig. 1) which grows 76 cassava varieties. Hereafter the term ‘variety’ refers to the unit identified by farmers under a single name. Each variety is recognized by farmers on the basis of morphological features (Elias, 2000). Interviews and questionnaires were conducted, in which farmers were asked to characterize each variety they grew (Elias, unpubl. data). Based on the answers, we classified each variety by delimiting a priori classes for each of the following traits, which constitute four major agronomic groups: the colour of the root; the starch content of the root; the degree of bitterness (associated with cyanide content); and the length of the cultivation cycle (Table 1).
Of the 76 varieties grown in Rewa, 31 were chosen for AFLP analyses (Table 2). The varieties chosen represent each of the major agronomic groups defined above, and include both frequent and rare varieties. For each variety, three to five plants were collected, from one or several farmers depending on the variety. The sampling strategy therefore focused mainly on intervarietal diversity. Wild cassava populations (M. esculenta ssp. flabellifolia) (A.C. Allem, EMBRAPA Recursos Genéticos e Biotecnologia, Brazil, pers. comm.), are also present in the same region. Plants from three very small populations of wild cassava located near Rewa were also analysed (Table 3). Their map locations are given in Fig. 1.
Material from the CIAT core collection
CIAT provided us with 38 accessions from a core collection of cultivated cassava, chosen to maximize the extent of diversity (Roa et al., 1997). We also obtained from CIAT 14 accessions of wild cassava (M. esculenta ssp. flabellifolia and M. esculenta ssp. peruviana, which Roa et al. (1997) considered synonymous with M. esculenta ssp. flabellifolia), originating from different locations (Table 4).
Analyses of genetic variability with AFLP markers
DNA extraction of Manihot accessions from Guyana was performed on leaves dried for 48 h at 35°C, using the protocol described by Colombo (1997). We followed the AFLP protocol described by Vos et al. (1995) and the adaptation for silver staining detection of Le Thierry d’Ennequin (1999). We used A and G as selective nucleotides during pre-amplification for EcoRI and MseI primers, respectively. Two primer combinations selected for their polymorphism and readability (Y. Gutiérrez, CIAT, Cali, Columbia, personal communication) were used for final amplifications: EcoRI + AAC × MseI + GTA, and EcoRI + ACG × MseI + GGT.
Data analysis
AFLP data were recorded in terms of the presence or absence of each band level for each plant. Shannon indices of phenotypic diversity (Bowman et al., 1971) were calculated for the set of varieties from Guyana, and for the sample from the CIAT core collection. The similarity index (S) of Nei & Li (1979) and the distance matrix (D=1 − S) were calculated on the basis of interindividual pairwise comparisons. Neighbour-Joining dendrograms were constructed using the PHYLIP software (Felsenstein, 1993). Robustness of nodes was estimated by using 100 bootstrap resamplings. In order to evaluate how genetic variability was distributed among major agronomic groups and among varieties, analyses of variance on the molecular diversity were performed (variety level nested in the group level) using version 1.55 of the WINAMOVA software (Excoffier et al., 1992). Φ statistics (analogous to F statistics (Wright, 1965)) were also computed. Non-parametric tests for variance components and Φ statistics were conducted with 500 random permutations of the data set.
Results
Polymorphism revealed by AFLP markers
119 AFLP bands were detected (50 with AAC × GTA combination, and 69 with ACG × GGT combination), among which 112 were polymorphic within the whole sample (94% polymorphism). Considering only cultivated accessions, 99 AFLP bands were detected, among which 80 were polymorphic (81% polymorphism). Among wild accessions, 107 bands were detected, of which 98 were polymorphic (92% polymorphism). Twenty-three of these bands were found only in wild accessions, and 14 were specific to wild accessions from Guyana.
Organization of genetic variability in the local varieties
The classification in major agronomic groups, the number of plants studied and the number of AFLP patterns detected for each variety are given in Table 1. Both monomorphic and polymorphic varieties can be identified. All the plants of a monomorphic variety have exactly the same AFLP patterns. However, the small sample size does not allow the conclusion that each of these varieties is clonal. The plants from the monomorphic varieties E22, E25, and E39 were collected from several farmers. In contrast, other varieties collected from different farmers displayed different genotypes among the farmers (e.g. polymorphic varieties E7, E12, E43). For the majority of polymorphic varieties, polymorphism was detected even within a single farm (e.g. individuals E40-3 and E40-4, which were collected from the same farm and display different AFLP patterns). This result is similar to those obtained by Second et al. (1997) and Mkumbira et al. (personal communication) in fields grown by Caboclos from Brazil and farmers from Malawi, respectively.
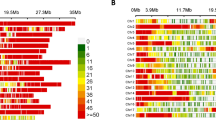
Figure 2 shows a Neighbour-Joining (NJ) dendrogram based on the Nei and Li similarity index, including all the plants from Rewa, and rooted with wild accessions from Guyana. The bootstrap values are shown for each node. The NJ dendrogram shows that two types of polymorphic varieties exist, homogeneous and heterogeneous. For the former (e.g. E5) all the plants studied cluster together in the dendrogram. For the latter (e.g. E6), the plants are dispersed in the dendrogram.
Neighbour-Joining dendrogram based on the Nei and Li similarity index, showing bootsrap values. Cassava varieties from Rewa, and wild accessions from Guyana are compared. For each cultivated accession, corresponding classes of the four major agronomic groups are indicated (codes are in Table 1).
There is obviously no agreement between the classification in major agronomic groups (shown in Fig. 2) and the organization of the molecular polymorphism as revealed by AFLP. In most cases, varieties that cluster together in the dendrogram belong to different major agronomic groups. Results given in Table 5 confirm that only a very small portion of the molecular variation is distributed between groups (from 0 to 9.8% depending on the trait under consideration). It is noticeable that the percentage of the total variance explained by within-variety diversity is larger than the proportion due to differentiation between groups. Finally, most of the variation is distributed between varieties (Φsc varies from 0.79 to 0.80). Only one variety of the sample was sweet (0 degree of bitterness). Although we can conclude that there is very little differentiation in AFLP pattern among bitter varieties according to their degree of bitterness, the material used in this study does not allow generalization of this conclusion to all the sweet and bitter varieties present in Guyana.
Structure of diversity in a single Makushi village, compared to the core collection
Figure 3 shows a rooted NJ dendrogram based on the Nei and Li similarity index, including all varieties from Rewa (the monomorphic or polymorphic and homogenous varieties are represented by one plant only), accessions from the core collection, as well as accessions of wild cassava from Guyana and those provided by CIAT. Excepting the wild accession PER417-6, considered to be a putative escapee from cultivation (C. Roa, CIAT, Cali, Columbia, pers. comm.), there is no interclustering between wild accessions and cultivated cassava whatever their geographical origins. The wild accessions from Guyana are distinct from the wild accessions provided by CIAT (although all have been identified as M. esculenta ssp. flabellifolia).
Accessions from the core collection and local varieties do not interpenetrate much in the dendrogram, and the nodes linking accessions from the core collection to local varieties are not supported by high bootstrap values. Also, the value of the Shannon index for the varieties grown in Rewa is 4.293, whereas that of the sample from the core collection is 4.289. This shows that the same amount of genetic variability is present in one small Amerindian village and in the core collection from CIAT.
Discussion
Intra-varietal genetic variability
Each generation, a very low number of plants per variety (about 10% of the initial population of a field) contribute to the stock of cuttings planted the next generation. This should lead to a very strong bottleneck effect and to genetic uniformity within varieties in a few generations. Our results are not in agreement with this expectation. However, plants of the same variety were sometimes collected in fields of different farmers. Although sample size does not allow evaluation of the level of differentiation within varieties between farmers, the intravarietal diversity could be partly due to the genetic differentiation of plant material grown by different farmers. Exchange of cuttings between farmers is a factor that should lead to the genetic homogenization of varieties and loss of genotypic diversity, as expected from classical population genetics when migration is effective between a limited number of populations (Varvio et al., 1986). However, such exchanges also allow the existence of intravarietal and within-farm polymorphism in transition phases to equilibrium. In this case, the number of different genotypes present in a variety for a given farmer would depend on the frequency of introduction of cuttings from other farmers of the community or from other villages. Moreover, different farmers sometimes use the same names for different plants that differ for morphological and agronomic features. Recent inquiries seem to reveal such inconsistency of taxonomy among farmers, at least for some varieties, particularly those named after their origin, or the quality of the root (Elias, 2000).
Our results show, however, that intravarietal genetic variability is also found among plants collected from the stock of a single farmer. In this case, one could invoke confusion between plants belonging to different varieties but sharing similar phenotypes. For instance, individuals 7-4 and 7-3 are representatives of the same variety (‘fat boy stick’) coming from the same farmer, but they have different AFLP patterns. Genetically plant 7-3 is very close to 47-2 (‘cement stick’). Morphological descriptions and ethnological observations suggest that these two varieties have very similar features and are frequently confused by the farmers. Situations like this one could be common and would partly explain the intravarietal polymorphism observed.
Another explanation for within-variety polymorphism could be the incorporation, in the farmer’s planting stock, of volunteer plants originating from seeds produced by unmanaged sexual reproduction. Our inquiries have shown that the Makushi often multiply volunteer cassavas. This has also been reported among other Amerindian tribes (Boster, 1984; Salick et al., 1997; Emperaire et al., 1998) and in Africa (Chiwona-Karltun et al., 1998), but it has always been considered as marginal and of rare occurrence. Sexual reproduction of cassava in Makushi fields would allow recombination between varieties, because intercropping varieties is the general rule and because cassava has a predominantly outcrossing mating system (Rogers, 1965). Volunteer cassavas could therefore correspond to new genotypes, different from the parental varieties. While most of them, because of novel combinations of morphological characters, are considered by the Makushi as new varieties, we have observed that some are deliberately assigned to a known variety with which they share most of their morphological features. Although the frequency of this process is uncertain, we suspect it could be a major factor accounting for introduction of intravarietal genetic diversity to a greater degree than previously suspected (Elias, 2000).
Finally, somatic mutations transmitted through vegetative propagation must also be considered as a factor explaining intravarietal polymorphism. It is noticeable, for example, that for variety 23 only one band of 55 amplified differentiates individual 23-4 from the other plants of this variety.
Comparison of genetic variability between local varieties and the core collection and wild forms
Genetic diversity of the varieties grown in Rewa, assessed by the Shannon index of phenotypic diversity, is as high as genetic diversity of the sample from the core collection. Similar findings have been reported previously for cassava in other traditional farming systems in the Amazon (Colombo, 1997; Second et al., 1997). A small number of varieties would provide for the Makushi culinary requirement. However, they cultivate more than 76 varieties of cassava. Although Makushi have a strong preference for highly productive and yellow rooted varieties, there is no absolute selection. A farmer very rarely discards even a low-yielding variety, but maintains it at a low frequency. This corresponds to a strategy of risk management in uncertain farming conditions, but the practice is also motivated by social or cultural reasons. Because diversity is prized for its own sake, Makushi cultivators are eager to acquire new varieties through multiplication of volunteer cassava or through exchanges (within the community, or with other villages). These practices ensure that genetic variability is introduced and maintained between and within varieties, as witnessed by the morphological and molecular diversity we observed.
Finally, our results show that the scarcity in the core collection of local varieties leads, to some extent, to an under-representation of the genetic variability of cassava, because high diversity and novel genotypes are characteristic of traditional farming systems in comparison to the few cultivars found in modern farming systems. These results highlight the need for redefining the biological unit for strategies of conservation of local cassava diversity based not only on the major agronomic groups or even on the variety, but also on ecological and human factors that contribute to the differentiation of the genetic stocks. Traditional practices, especially intercropping and incorporation of volunteer seedlings, would promote the contribution of recombinant genotypes to the cultivated stocks, therefore allowing selection and adaptation to continue in this mainly vegetatively propagated crop. Even though the extents of traditional practices and their consequences for the genetic diversity of local varieties must be evaluated more precisely, it is already obvious that the loss of these practices could lead to a long-term reduction of genetic variability in local cassava varieties. Conservation strategies should therefore aim to maintain such processes.
Phylogenetic studies (Schaal et al., 1997; Olsen & Schaal, 1999) and analyses of the structure of the genetic variability in the cassava species complex (Fregene et al., 1994; Colombo, 1997; Roa et al., 1997) tend to confirm Allem’s (1994) hypothesis that M. esculenta ssp. flabellifolia is the wild progenitor of cultivated cassava. In our study, wild forms clustered independently from all cultivated varieties. Similar patterns were observed in previous studies by Colombo (1997) and Roa et al. (1997). Such a pattern is consistent with an hypothesis of a limited (or even unique) domestication event in a restricted area, followed by diffusion of the cultivated phenotypes, as has been suggested, for example, for wheat (Heun et al., 1997). Olsen & Schaal (1999) reached the same conclusion for cassava from phylogeographic analyses using molecular sequences. They proposed that cultivated cassava originated in southern Amazonia. This scenario seems all the more plausible as fixation of interesting phenotypes, and their diffusion throughout the Amazonian basin, would have been facilitated by the vegetative reproductive mode. The pattern of genetic differentiation between varieties that shows very short internodes and long-terminal branches in the dendrogram, could therefore be interpreted as a consequence of a radiation-type divergence of these varieties for morphological features under human selective pressures, from a common ancestral domesticated stock, combined with their fixation through vegetative propagation. In addition, the very low level of genetic differentiation between the major varietal groups defined above, even with the expectation of strong linkage disequilibrium as a consequence of the vegetative reproduction mode, would suggest that convergent evolution for the agronomic features defining them has occurred. However, confirming these results requires a more precise evaluation of the agronomic features used for this classification than one based on questionnaires and interviews only.
Our results also show that domesticated cassava in Guyana is more similar (in terms of AFLP markers) to wild cassava from Brazil, which is closer to the putative centre of domestication, than it is to wild cassava in Guyana. This leads us to hypothesize that gene flow from wild cassava is not a significant evolutionary factor in cultivated populations in Guyana and that the local wild gene pool has contributed little to the diversity of the cultivated pool.
The interpretations offered here depend on the reliability of AFLP markers in reflecting evolutionary relationships between taxa at the infraspecific level. Several studies have already shown the reliability of AFLP markers for taxonomic purposes in cultivated species including lettuce (Hill et al., 1996) and tea (Paul et al., 1997). In rice, analyses of cultivated varieties of the indica and japonica groups, carried out using RFLP, AFLP and ISSR markers, have shown that similar branching patterns are obtained whatever the type of markers used (Blair et al., 1999). However, one cannot discard the possibility that some of the AFLP markers correspond to very rapidly evolving sequences (Reamon-Büttner et al., 1999), and that saturation and homoplasy could therefore occur, at least for such markers, even at the infraspecific level. However, this hypothesis is difficult to equate with the cultivated/wild dichotomy of AFLP markers found in cassava.
References
Allem, A. C. (1994). The origin of Manihot esculenta Crantz (Euphorbiaceae). Genet Res Crop Evol, 41: 133–150.
Blair, M. W., Panaud, O. and McCouch, S. R. (1999). Application of ISSR amplification for the fingerprinting of rice cultivars and for the analysis of microsatellites motif frequencies in rice genome. Theor Appl Genet, 98: 780–792.
Boster, J. S. (1984). Classification, cultivation, and selection of Aguaruna cultivars of Manihot esculenta (Euphorbiaceae). In: Prance, G. T. and Kallunki, J. A. (eds) Ethnobotany in the Neotropics, Advances in Economic Botany, Vol. 1, pp. 127–173. New York Botanical Garden, Bronx, NY.
Bowman, K. D., Hutcheson, K., Odum, E. P. and Shenton, L. R. (1971). Comments on the distribution of indices of diversity. Stat Ecol, 3: 315–359.
Chiwona-Karltun, L., Mkumbira, J., Saka, J., Bovin, M., Mahungu, N. M. and Rosling, H. (1998). The importance of being bitter – a qualitative study on cassava cultivar preference in Malawi. Ecol Food Nutr, 10: 1–27.
Colombo, C. (1997). Etude de la diversité génétique des maniocs américains (Manihot esculenta Crantz) par les marqueurs moléculaires (RAPD et AFLP). PhD Dissertation, ENSAM, Montpellier.
Dewet, J. M. J. and Harlan, J. R. (1975). Weeds and domesticates: evolution in the man-made habitat. Econ Bot, 29(2)99–107.
Elias, M. (2000). Le manioc chez les Makushi: une richera à valoriser. In: Grenand, P. (ed.) Avenir des Peuples des Forêts Tropicals, Volume Régional Caraïbes. EC DG VIII; Bruxelles, in press.
Emperaire, L., Pinton, F. and Second, G. (1998). Une gestion dynamique de la diversité variétale du manioc en Amazonie du Nord-Ouest. Nature, Sci Société, 6: 27–42.
Excoffier, L., Smouse, P. E. and Quattro, J. M. (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: applications to human mitochondrial DNA restriction data. Genetics, 131: 479–491.
Felsenstein, J. (1993). Phylip. Phylogeny Inference Package, Version 3.5c. Department of Genetics, University of Washington, Seattle, WA.
Fregene, M. A., Vargas, J., Angel, F., Tohme, J., Asiedu, R. A., Akorada, M. O. et al (1994). Chloroplast DNA and nuclear ribosomal DNA variability in cassava (Manihot esculenta Crantz) and its wild relatives. Theor Appl Genet, 89: 719–727.
Heun, M., Schäfer-Pregl, R., Klawan, D., Castagna, R., Accerbi, M., Borghi, B. et al (1997). Site of einkorn wheat domestication identified by DNA fingerprinting. Science, 278: 1312–1314.
Hill, M., Witsenboer, H., Zabeau, M., Vos, P., Kesseli, R. and Michelmore, R. (1996). PCR-based fingerprinting using AFLPs as a tool for studying genetic relationships in Lactuca spp. Theor Appl Genet, 93: 1202–1210.
Lethierry d’Ennequin, M. (1999). Domestication et évolution du génome des céréales: l’exemple du millet ( Setaria italica L. Beauv.). PhD Dissertation, University of Paris VI.
McKey, D. B. and Beckerman, S. (1993). Chemical ecology, plant evolution and traditional cassava cultivation systems. In: Hladik, C. M., Hladik, A., Pagezy, H., Linares, O. F., Koppert, G. J. A. and Froment, A. (eds) Tropical Forests, People and Food, pp. 83–112. UNESCO, Paris.
Miller, K., Allegretti, M. H., Johnson, N. and Johnson, B. (1995). Measures for conservation of biodiversity and sustainable use of its components. In: Heywood, V. H. (ed.) Global Biodiversity Assessment. United Nations Environment Programme. Cambridge University Press, Cambridge.
Nei, M. and Li, W. (1979). Mathematical model for studying genetic variations in terms of restriction endonucleases. Proc Natl Acad Sci USA, 76: 5269–5273.
Olsen, K. M. and Schaal, B. A. (1999). Evidence on the origin of cassava: Phylogeography of Manihot esculenta. Proc Natl Acad Sci USA, 96: 5586–5591.
Paul, S., Wachira, F. N., Powell, W. and Waugh, R. (1997). Diversity and genetic differentiation among populations of Indian and Kenyan tea (Camellia sinensis (L.) O. Kuntze) revealed by AFLP markers. Theor Appl Genet, 94: 255–263.
Reamon-Büttner, S. M., Schmidt, T. and Jung, C. (1999). AFLPs represent highly repetitive sequences in Asparagus officinalis L. Chrom Res, 7: 297–304.
Roa, A. C., Maya, M. M., Duque, M. C., Tohme, J., Allem, A. C. and Bonierbale, M. W. (1997). AFLP analysis of relationships among cassava and other Manihot species. Theor Appl Genet, 95: 741–750.
Rogers, D. J. (1965). Some botanical and ethnological considerations of Manihot esculenta. Econ Bot, 19: 369–377.
Salick, J., Cellinese, N. and Knapp, S. (1997). Indigenous diversity of cassava: generation, maintenance, use and loss among the Amuesha, Peruvian upper Amazon. Econ Bot, 51: 6–19.
Schaal, B., Carvalho, L. J. C. B., Prinzie, T., Olsen, K., Hernandez, M., Cabral, G. et al (1997). Phylogenetic relationships and genetic diversity in Manihot species. Afr J Root Tuber Crops, 2: 147–149.
Second, G., Allem, A. C., Mendes, R. A., Carvalho, L. J. C. B., Emperaire, L., Ingram, C. et al (1997). Molecular marker (AFLP)-based Manihot and cassava numerical taxonomy and genetic structure analysis in progress: implications for their dynamic conservation and genetic mapping. Afr J Root Tuber Crops, 2: 140–147.
Varvio, S. L., Chakraborty, R. and Nei, M. (1986). Genetic variation in subdivided populations and conservation genetics. Heredity, 57: 189–198.
Vos, P., Hogers, R., Bleeker, M., Reljans, M., van de Lee, T., Hornes, M. et al (1995). AFLP: a new technique for DNA fingerprinting. Nucl Acids Res, 21: 4407–4414.
Wright, S. (1965). The interpretation of population structure by F-statistics with special regard to mating systems. Evolution, 19: 395–420.
Acknowledgements
This study was supported in part by the European Commission (DGVIII, Future of Tropical Forest Peoples (APFT) Program), that provided funds for fieldwork. The Bureau des Ressources Génétiques (France) supported all genetic analyses. We are grateful to the government of Guyana, specially the Ministry of Amerindian Affairs and the Environmental Protection Agency, for allowing us to carry out our fieldwork in Guyana. We are also grateful to the National Agronomic Research Institute (Georgetown, Guyana) and to the Amerindian Research Unit of the University of Guyana for their useful advice. We wish to thank personnel of the CIAT, particularly Dr Joseph Tohme and Carolina Roa, for their great help, and for providing us with DNA samples. Many thanks also to Dr Antonio Costa Allem, for identifying accessions of M. esculenta ssp. flabellifolia. Our greatest debt of gratitude is to the Makushi Amerindians of Rewa for their hospitality and interest in participating in our study. Many thanks to Dr Bruno Toupance for adapting the PHYLIP program, and to Professor Doyle McKey, Dr Bernard Lejeune and two anonymous reviewers for their useful comments on this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Elias, M., Panaud, O. & Robert, T. Assessment of genetic variability in a traditional cassava (Manihot esculenta Crantz) farming system, using AFLP markers. Heredity 85, 219–230 (2000). https://doi.org/10.1046/j.1365-2540.2000.00749.x
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1046/j.1365-2540.2000.00749.x
Keywords
This article is cited by
-
Lost Grains and Forgotten Vegetables from Japan: the Seikei Zusetsu Agricultural Catalog (1793–1804)
Economic Botany (2019)
-
Genetic basis for folk classification of oca (Oxalis tuberosa Molina; Oxalidaceae): implications for research and conservation of clonally propagated crops
Genetic Resources and Crop Evolution (2017)
-
Mining and characterization of EST-SSR markers for Zingiber officinale Roscoe with transferability to other species of Zingiberaceae
Physiology and Molecular Biology of Plants (2017)
-
Development of TRAP (Target Region Amplification Polymorphism) as New Tool for Molecular Genetic Analysis in Cassava
Plant Molecular Biology Reporter (2015)
-
Genetic analysis of Pongamia pinnata (L.) Pierre populations using AFLP markers
Tree Genetics & Genomes (2014)