Abstract
Twenty-three populations of Pinus pinaster (13 Aquitaine populations and 10 Corsican populations) were analysed at three microsatellite loci and 122 AFLP loci. The aims of the study were: (i) to compare levels of within-population and among-population diversity assessed with both kinds of markers; (ii) to compare Aquitaine and Corsican provenances of P. pinaster; and (iii) to know if both markers gave the same information for conservation purposes. Classical population genetics statistics were estimated and the ranking of populations obtained using different markers and/or parameters were compared by computing Spearman’s rank correlations. Even though microsatellites showed a higher within-population diversity, they showed the same level of differentiation as AFLP markers. Moreover, both markers also showed a higher genetic diversity in the Aquitaine provenance and a higher differentiation among Corsican populations. AFLPs and microsatellites gave different population diversity rankings. Consequently, the results do not support the potential population identification within each provenance for conservation purposes.
Similar content being viewed by others
Introduction
Different marker systems are currently available for monitoring genetic diversity in natural populations. Examples of markers with contrasting properties are AFLP markers (Vos et al., 1995) and microsatellites (Tautz, 1989). First, AFLPs are considered to be dominant and biallelic, whereas microsatellites are codominant and usually reveal a high number of alleles. Secondly, the AFLP technique provides many markers randomly spread in the genome whereas, in a diversity study, the number of analysed microsatellites is usually low. Although mutation rates may be different for different markers and may be considered ‘marker specific’, other evolutionary factors such as migration or drift affect the whole genome regardless of the genomic regions that are monitored by different marker systems. The comparative analysis between different markers is therefore intended to reveal the balance between different evolutionary forces contributing to genetic diversity. In this paper, we compare the levels of diversity within and among Pinus pinaster Ait. populations estimated by both types of markers (AFLP and microsatellites).
The natural range of P. pinaster is restricted to the western part of the Mediterranean basin, including continental France (mainly south-west and south-east), Corsica, Spain, Portugal, continental Italy (Liguria and Tuscany), Italian islands (Sardinia and Pantelleria), Morocco and Tunisia. Previous genetic diversity studies have been performed using terpene, protein and allozyme markers (Baradat & Marpeau-Bezard, 1988; Baradat et al., 1991; Bahrman et al., 1992; Petit et al., 1995; Bahrman et al., 1997). The results of these studies indicated a subdivision of the species into three groups: Atlantic (including south-west France, Portugal and Galicia in Spain), peri-Mediterranean (including Andalusia in Spain, Corsica, Maures in south-east France and Liguria in Italy) and North African (Moroccan Atlas, Algeria and Tunisia). They also suggested that the species differentiated into regional “races” after the last glaciation, beginning in the south-west of the Iberian Peninsula. Salvador et al. (2000) studied the genetic variability with 18 isozyme loci and concluded that the southern Iberian range could be the putative recolonization centre of the species after the last glaciation. Chloroplast microsatellites were recently used in P. pinaster for studying genetic diversity and colonization processes of the species (Vendramin et al., 1998; Ribeiro et al., 2000). As a whole, few studies focused on the fine scale description of genetic variability of P. pinaster within regional ‘races’.
Maritime pine is a widely afforested species in France. An intensive breeding programme is conducted with breeding populations originating from the Aquitaine and Corsican provenances. Natural resources of the species might be threatened in France. In south-west France, the artificial selection in the improvement programme may cause a reduction of the genetic variability. In the south-east, the threat may be due to the fires and to the bast scale Matsucoccus feytaudi Duc., which is a specific pest of P. pinaster (Burban et al., 1999). After proliferating in south-east France, the pest recently appeared in Corsica (Jactel et al., 1996) where it is spreading and producing extensive damage (Jactel et al., 1998).
In this paper, 13 Aquitaine populations and 10 Corsican populations of P. pinaster were screened with two different types of molecular markers: three nuclear microsatellite loci and 122 AFLP loci. The first objective of the study was to compare the levels of within- and among-population diversity assessed by both types of markers. The second objective was to analyse and compare the genetic diversity of P. pinaster populations from the Aquitaine provenance and from the Corsican provenance. Finally, we intended to draw recommendations for conservation purposes based on the comparative analysis of genetic diversity with different markers.
Materials and methods
Plant material and DNA extraction
We sampled 13 populations in Aquitaine (south-west France). Populations were selected on the basis of historical records about their native origin [French National Forest Office (ONF), personal communication]. From palynological data, Paquereau (1964) indicates that P. pinaster appeared in Aquitaine about 8000 years ago and spread along the coast, where the species constituted some monospecific pine forests during the last 3000 years. The populations used in this study most likely originated from original stands after successive natural regenerations. Populations were sampled to represent the natural genetic variation of this species in south-west France.
The Corsican set of populations was selected among 30 previously collected populations (Durel & Bahrman, 1995). Ten of those populations, regularly distributed along the island, were chosen to cover the range of altitudes, from 10 m to 1105 m.
For the Aquitaine populations, needles were collected from about 30 young trees (3–5-years-old) randomly distributed over an area from 10 to 30 hectares for each population. In the Corsican populations, one seedling was sampled for each of about 30 open pollinated progenies per population. The mother trees of the open pollinated progenies were distributed on approximately 50 hectares. Genomic DNA was extracted from needles as described by Doyle & Doyle (1990).
Microsatellite and AFLP markers
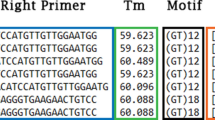
The 23 populations were screened with three nuclear microsatellites (FRPP91, FRPP94 and ITPH4516) and with two AFLP primer–enzyme combinations. Microsatellite amplifications were performed as described by Mariette et al. (2001) and AFLP markers were obtained following the protocol by Costa et al. (2000). Two primer–enzyme combinations, EcoRI + ACC/MseI + CCTG and EcoRI + ACG/MseI + CCCA, coded PEC 1 and PEC 2, respectively, were used. They were selected according to the number of amplified loci (respectively 110 and 81) and the number of polymorphic loci (respectively 29 and 28) detected in a mapped population (Costa et al., 2000). Microsatellite and AFLP loci were scored with the RFLP-SCAN version 3.0 (Scanalytics) software.
Statistical analyses
Data analysis of microsatellite markers.
For each locus, the allelic richness (A), the allelic frequencies, the observed heterozygosity (HO), the effective number of alleles (AE=1/(1 − HE)) and the fixation index (FIS) were calculated as described by Brown & Weir (1983). The analysis of diversity was also performed using Nei’s (1987) single-locus estimators: the within-population gene diversity (Hi within population i and HS mean value of Hi over several populations), the total gene diversity (HT) and the coefficient of gene differentiation among populations (GST). These parameters were computed by using the DIPLOIDE program (Antoine Kremer, Equipe de Génétique et Amélioration des Arbres Forestiers, Cestas, France).
Data analysis of AFLP markers.
To analyse AFLP markers, we assumed that each AFLP amplification product, regardless of its relative intensity, corresponded to a dominant allele at a unique locus. Polymorphic amplified loci were scored as ‘1’ for the presence and ‘0’ for the absence of a locus. Only loci with clearly amplified bands were used for data analysis.
The analysis of genetic diversity with AFLP fragments was done after estimating the frequencies of the alleles responsible for the presence and absence of fragments. If p and q are the alleles responsible for the presence and absence of an AFLP fragment, and if Q is the frequency of trees that do not exhibit the fragment, then Q=q2(1 – FIS) +qFIS, where FIS is the fixation index. Assuming that the true value of FIS is known, an asymptotically unbiased estimate of q is obtained by the use of a second order Taylor expression:

with Δ=FIS2 + 4(1 − FIS)Q and N the number of trees sampled per population.
The analyses were performed using the FIS value that was estimated based on the average value from the three microsatellites. We estimated the diversity over all loci and we also restricted the analysis to loci that showed an observed frequency smaller than 1 − (3/N), where N is the sample size, as recommended by Lynch & Milligan (1994). These authors showed that the bias introduced on the estimation of q due to small sample size was substantial when the null allele was rare. HS, HT and GST were computed by using the allelic frequencies as estimated by formula (1).
Analyses were performed by using the HAPDOM program (Antoine Kremer, Equipe de Génétique et Amélioration des Arbres Forestiers, Cestas, France).
For both types of marker, diversity parameters were estimated for each population and provenance (Corsica and Aquitaine). Mean values were estimated over all 23 populations, over all the Aquitaine populations and over all the Corsican populations.
Levels of differentiation (GST) were calculated among all 23 populations, among the Aquitaine populations, among the Corsican populations and between the Aquitaine provenance and the Corsican one.
Comparison of diversity and differentiation between the Corsican and Aquitaine provenance.
Resampling methods were used to estimate the differences between the two provenances [Aquitaine (A) and Corsica (C)] for the parameters AE, HO, HS, HT, FIS and GST. For example, in the case of HS, bootstrapping was used, where all the individuals were resampled with replacement within each population of each provenance A and C, and HS was computed in each provenance as well as the difference of HS computed for each provenance (HSAq − HSCor). Resampling was repeated 1000 times. The distribution of (HSAq − HSCor) was then compared with the null hypothesis (HSAq − HSCor = 0). Sampling variances were obtained by bootstrapping over individuals in populations for AE, HO, HS, HT and FIS and by bootstrapping over populations for GST. The sample sizes in the Aquitaine and in the Corsican provenances were uneven, therefore the rarefaction method as described by Petit et al. (1998) was used for estimating the unbiased allelic richness at each locus, using 50, 100 and 200 as sample sizes for each provenance.
Comparison of allele frequencies among populations within provenances.
The geographical pattern of the studied populations was analysed by using the principal component analysis (PCA) and Mantel tests. PCA was performed using the allele frequencies for each microsatellite and the frequencies of loci obtained in the genotypic AFLP analysis. In the case of microsatellites only allele frequencies of the five most frequent alleles were used. The ACP program (Philippe Baradat, Equipe de Génétique et Amélioration des Arbres Forestiers, Cestas, France) was used to compute the PCA analysis.
Mantel tests were performed for comparing the pairwise population geographical distance (in kilometres) with the pairwise population genetic distance matrix (Nei, 1978). The strength of the relationship between both matrices was measured using the Mantel matrix-correspondence test (Mantel, 1967; Sokal, 1979). The null hypothesis refers to the absence of association between the elements of the pairs of matrices. A normalized Z-test was performed in which the observed value after 999 permutations should be significantly larger than that expected by chance in order for an association to be accepted as valid. The program used to compute Mantel tests was TFPGA version 1.3 (Mark P. Miller, Northern Arizona University, U.S.A.).
Intra- and inter-marker comparisons.
Diversity assessed at different markers results from either ‘marker specific’ factors (such as mutation) or ‘marker non-specific’ factors (as for example drift or migration). Hence one would expect that values of diversity of a given population as assessed with different markers should be different. However, if two populations went through different evolutionary histories involving drift and migration, then they would differ for their level of diversity in a same manner for two different kinds of markers. In other words, one expects congruent rankings of the populations between markers. As a result, the comparative analysis of diversity between markers is not based on the comparison of levels of diversity within a population, but on the comparison of the ranking of different populations by correlation analysis.
For each type of marker (AFLP or microsatellites), A, AE, HO and HS were estimated for each population (and provenance). Estimates of these diversity statistics were then compared between populations by computing Spearman’s rank coefficient correlation (Sokal & Rohlf, 1995). A significant correlation would indicate congruence between different factors and the impact of ‘marker non-specific’ factors on levels of diversity. Conversely, lack of correlation is likely to reveal that ‘marker specific’ factors are sufficiently different between populations to generate different levels of diversity.
Results
Within-population diversity
Microsatellites.
Considering all the 23 populations, the three microsatellites showed a high level of polymorphism with an average allelic richness A of 27.33, ranging from 23 to 31 alleles. HO was 0.645, ranging from 0.584 to 0.690, and HT was 0.832, ranging from 0.742 to 0.903 (Table 1).
AFLP markers.
For the PEC 1 and the PEC 2, 56 and 66 amplified loci were analysed, respectively. The number of polymorphic loci in at least one population was 23 (41%) for PEC 1 and 36 (55%) for PEC 2. Thus, on a whole, 122 loci were analysed and 48% of them were polymorphic.
Using the fixation index FIS obtained from microsatellites to estimate allelic frequencies (eqn 1), the total diversity was 0.161 for PEC 1, 0.194 for PEC 2 and 0.179 for all the studied loci. Using only the 36 polymorphic markers, as suggested by reducing the number of loci from Lynch & Milligan (1994), the levels of diversity were increased [0.321, 0.297 and 0.305 for PEC 1, PEC 2 and both PECs, respectively (Table 2)].
Comparison of diversity in Aquitaine and Corsica
At the single population level, Aquitaine populations showed higher within-population diversity levels than Corsican populations. This result was significant at the 5% level both for microsatellites (Table 1) and AFLP markers (Table 2). However, locus FRPP94 showed a significantly higher effective number of alleles and within-population diversity for Corsican populations (Table 1). The fixation index within Corsican populations was significantly higher for locus FRPP91 and over all loci (Table 1).
At the provenance level, the Aquitaine provenance showed significantly higher levels of diversity than the Corsican one, both for microsatellite and AFLP markers (Table 1 and Table 2). However, loci FRPP94 and PEC 1 showed significant higher diversity parameters for the Corsican provenance.
With 50, 100 and 200 as sample sizes for each provenance, the rarefaction method gave a higher number of alleles in Corsica for FRPP91 (27.4 in Corsica and 21.5 in Aquitaine for a sample size of 200) and for FRPP94 (18.3 in Corsica and 11.7 in Aquitaine). In contrast, ITPH4516 showed more alleles in the Aquitaine provenance (22.4 in Aquitaine and 18.8 in Corsica). The Corsican provenance showed a higher mean allelic richness (A=23.33) than the Aquitaine provenance [A=21.33 (Table 1)].
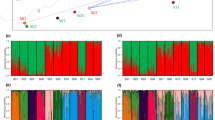
Considering the distribution of allelic frequencies of microsatellites, the provenances did not share the same common alleles for FRPP91 and for ITPH4516 (Fig. 1). For FRPP91, one allele showed divergent frequencies in Corsican and Aquitaine provenances (0.680 in Corsica and 0.004 in Aquitaine). For all loci, each provenance exhibited private alleles: for FRPP91, one rare allele was specific to Aquitaine and six were specific to Corsica; for FRPP94, three alleles were found in Aquitaine against nine in Corsica and for ITPH4516, eight alleles were found in Aquitaine vs. three in Corsica (Fig. 1). Corsica showed a higher number of rare alleles (18 against 12 in Aquitaine).
AFLP markers also showed private alleles. Considering both PECs, seven polymorphic loci exhibited bands in Aquitaine which were absent from Corsica. No specific allele was found in Corsica.
Subdivision of genetic diversity among populations and between provenances
The average differentiation among all populations was 11.1% for the microsatellite markers (Table 3), varying from 6% for FRPP94 to 18.2% for FRPP91. For AFLP markers the level of differentiation was 10.2% for the genotypic analysis, when all 122 loci were considered, and reduced to 6.1%, following Lynch & Milligan (1994). The differentiation levels estimated with the two primer–enzyme combinations were found to be significantly different. The PEC 2 showed a significantly higher differentiation than the PEC 1 (data not shown).
The genetic differentiation assessed with microsatellite markers was significantly higher than the differentiation assessed with AFLP markers only when we used the Lynch & Milligan (1994) method of analysis.
The differentiation among Corsican populations was higher than the differentiation among Aquitaine populations with both microsatellite and AFLP markers (Table 3). The difference was significant for microsatellites and for the analysis of AFLP when the 122 loci were considered.
The differentiation between provenances (9.2% for microsatellites and 4.9% for AFLP markers) represented more than half of the total differentiation (Table 3).
Comparison of levels of diversity assessed with different measures (A and Hi)
Allelic richness and expected heterozygosity estimated with microsatellite loci within each population exhibited similar rankings of populations within each provenance. The rankings of populations given by A calculated over all microsatellites were positively and significantly (5% level) correlated with rankings given by Hi, both for Aquitaine populations (rS=0.765, P=0.018) and for Corsican populations (rS=0.710, P=0.035). In the Aquitaine populations, the correlation was also significantly positive for each microsatellite.
For Corsican populations, pairwise comparison of diversity statistics between loci did not always indicate congruent results. For FRPP91, the ranking given by HO was positively and significantly correlated with the ranking given by HO on FRPP94. The ranking given by A on FRPP91 was also positively and significantly correlated with the ranking given by HO on FRPP94 and by HO on ITPH4516. The same analysis was performed for the Aquitaine provenance and no correlation was ever found.
Comparison of levels of diversity assessed with different markers
For each type of analysis, the rankings of populations given by PEC 1 and PEC 2 were not significantly correlated for both Aquitaine and Corsican populations (Table 4). Rankings given by PEC 1 and PEC 2 were checked separately but were never significantly correlated (Table 5); they were positively and significantly correlated, as expected, with rankings given by both PECs, in general.
When comparing Hi values given by microsatellites and AFLP markers, no positive and significant correlation was found (Table 4).
Geographical pattern
The two first axes of the principal component analysis separated Aquitaine populations from Corsican ones (Fig. 2).
Mantel tests performed on Aquitaine populations did not detect any geographical pattern with AFLP markers or with microsatellites. A significant correlation (r=0.360, P=0.018) between the genetic distances matrix and the geographical distances matrix was found for Corsican populations when microsatellites were used.
Discussion
Maritime pine is a highly polymorphic species as shown by the values of diversity measures and comparisons with other species. For example, in this study, the average value of HO for microsatellites was 0.645 and the average value of A was 27.33. These values are similar to the values found in other pine species (Smith & Devey, 1994; Echt et al., 1996). The ultimate aim of this study was to derive conservation decisions based on a comparative survey of genetic diversity conducted with different markers. The comparative analysis was intended to reveal the differential impact of ‘marker specific’ factors vs. ‘marker non-specific’ factors on diversity. In addition, as conservation should be based on genetic diversity at the whole genome level, the comparative analysis will provide insights on the genomic heterogeneity for diversity and differentiation. Therefore, we will first discuss congruencies and discrepancies between markers, and then consider their consequences on conservation strategies.
Congruent results of differentiation obtained with microsatellites and AFLPs
As for diversity, some evolutionary factors such as migration may have an impact on differentiation regardless of the marker, whereas others (such as mutation) may be considered as ‘marker specific’. For most markers, the contribution of mutation to differentiation has generally been ignored because mutation rates of traditional isozyme markers are negligible when compared to migration rates. This may not be the case when comparing microsatellites and AFLP. Because microsatellite markers show higher mutation rates than other molecular markers (Weber & Wong, 1993), they are expected to exhibit lower levels of differentiation. Furthermore, as reported by Estoup et al. (1995), homoplasy due to the high mutation rate may increase the underestimation of differentiation. On the other hand, the dominant property of AFLP markers is also responsible for a bias in the estimation of differentiation. Jenczewski et al. (1999) showed, both with simulation and experimental results, that FST is biased upward when using RAPD phenotypes to estimate allelic frequencies and Krutovskii et al. (1999) showed that RAPDs can reasonably estimate population differentiation if sample sizes are larger than about 30 individuals. In our case, sample sizes were around 30 individuals for each population and some loci were fixed for the presence of amplified loci so an overestimation of differentiation for AFLPs cannot be excluded. Despite these expected discrepancies, our results lead to similar conclusions for genetic differentiation assessed with microsatellite and AFLP markers. Although ‘absolute values’ of differentiation may slightly differ between markers, they lead to similar conclusions (Table 3): (i) when all populations were considered, a high differentiation among populations was found both with microsatellites and AFLP markers; (ii) furthermore, both markers indicate that a predominant part of the total differentiation is due to the subdivision between the Aquitaine and Corsican provenances; and (iii) lastly the two markers reveal that there is more differentiation among the Corsican populations than among the Aquitaine populations. The congruent results obtained with different markers most likely indicate that marker differences (mutation rates, homoplasy, estimation biases) had only a minor impact on differentiation. Conversely, migration and drift may still be the prevailing factors involved. This is illustrated by the contrasting difference between the Aquitaine and Corsican forest conditions. The Aquitaine populations were sampled within a broad continuous forest occupying a flat area of about 1 million hectares. In contrast, the Corsican populations came from an heterogeneous discontinuous forest distributed in mountainous regions. Consequently there is likely to be more gene flow among stands in Aquitaine than in Corsica.
Interestingly our results (Table 3) could be compared with previous reports based on different markers, but also on different sampling of populations. GST values estimated on six populations chosen throughout the range of the species (Petit et al., 1995) were 0.170, 0.161 and 0.139, respectively, for protein, isozyme and terpene loci. Since the sampling area was larger, differentiation was also higher. However this study also indicated congruent results of differentiation across three different marker systems.
Congruent results of diversity obtained with microsatellites and AFLPs
Among the different evolutionary factors that have an impact on genetic diversity, mutation can be considered as ‘marker specific’. However there are other factors such as genetic drift and migration that can be considered as ‘non-marker specific’. For example, a reduction in population size in a given population will decrease the level of diversity for any marker (AFLP and microsatellites) regardless of its mutation rates. As a result, one would expect that populations that had different evolutionary histories would also differ in their levels of diversity, following a similar trend for both markers. Most of the diversity measures estimated with different markers showed a significantly lower level in Corsican populations than in Aquitaine populations (Table 1 and Table 2). For microsatellites, the Corsican populations showed a lower allelic richness than the Aquitaine ones but the alleles tended to be different in each population. For AFLPs, there are more pronounced differences between the two provenances for the analysis restricted to the polymorphic markers (according to the rules of Lynch & Milligan, 1994) than for the analysis conducted on the whole set of data.
The congruent results of diversity levels between the two markers suggest that population differences in diversity are more related to ‘marker non-specific’ than ‘marker specific’ causes. Among the former, population history may be advocated to interpret these results. The frequent occurrence of fire over large areas, the more severe climatic conditions in Corsica and the mountainous topography may have induced repetitive sequences of restriction and expansion of maritime pine forest, leading to progressive genetic erosion. Furthermore genetic erosion can not be prevented by gene flow from the continent, as Corsica is isolated by a few hundred kilometres from France or Italy. An alternative hypothesis, and not exclusive with the previous one, may be that the number of founder populations that became established on the Corsican island was limited.
Discrepancy between AFLPs and microsatellites for measuring within-population diversity
If the estimates of levels of diversity with different markers were congruent for the comparison between the Corsican and Aquitaine provenances, they do not lead to the same conclusions for the comparison of populations within each of the two provenances. There is no significant correlation among the diversity estimates calculated for the AFLPs and microsatellites (Table 4). Considering AFLP markers alone, a striking result was that the two primer–enzyme combinations provided different rankings of populations, particularly in Corsica. Furthermore, rankings given by two different microsatellite loci were rarely significantly and positively correlated. When marker systems were compared, neither A nor Hi, estimated by microsatellites, gave similar rankings to Hi estimated by AFLP markers. The discrepancy found between markers may be due to either biological and/or technical causes. ‘Marker non-specific’ factors (drift, migration, population history) did not differ sufficiently between different populations to generate different levels of diversity. This would result in the lack of differentiation for population diversity. Hence populations within the Corsica or Aquitaine provenance would be homogeneous for their level of diversity due, for example, to intensive and equal gene flow, and the absence of drift within each provenance. This hypothesis is supported by the statistical multipopulation comparison of diversity (Table 5), showing that, except with microsatellites in Corsican populations, no diversity difference could be detected among populations. The technical interpretation is related to the sampling used. The difference in levels of diversity may be so small among populations that our sampling strategy did not allow the detection of any significant differences. All differences among diversity estimates are therefore so small that the ranking observed is only generated by the noise of the estimation.
Conservation implications
The comparative analysis of genetic diversity with different markers has provided interesting insights not only into the organization and subdivision of diversity but also into the genetic mechanisms that contributed to the extant structure of diversity. All markers indicated that the Corsican and Aquitaine provenance are strongly differentiated. This is suggested by the high GST values between the two provenances and by the principal component analysis (Fig. 2). It is also illustrated by the important differences in allele frequencies for the microsatellite locus FRPP91 (Fig. 1). Conservation actions are therefore recommended to save diversity for each of the two provenances. Our results indicate however, that sampling of candidate populations or trees for conservation should be achieved differently in the two provenances. The Aquitaine provenance is characterized by a low differentiation for allele frequencies (extremely low GST values) and a low differentiation for within-population diversity, and these results are again consistent across markers. As a result ex situ or in situ conservation could be based on a random sample of trees throughout the Aquitaine area. On the other hand the Corsica provenance is characterized by a higher population differentiation for allele frequencies (GST values) and a slightly higher differentiation for within-population diversity. Consequently, it is expected that different populations may contain different alleles, as suggested by the distribution of allelic richness among populations. Sampling guidelines for conservation would therefore recommend concentrating on a few populations that would sample the total diversity. At this stage, our results indicate that a more intense monitoring of diversity should be conducted in the Corsican provenance in order to identify the candidate populations for conservation. To conclude on the conservation issue, we would also recommend complementing the present study with a parallel investigation on adaptive traits (drought, pest and insect resistance) and other economic traits (growth and form) before a definite strategy is adopted.
References
Bahrman, N., Baradat, P. and Petit, R. J. (1992). Structuration de la variabilité génétique du Pin maritime dans l’ensemble de son aire naturelle. Colloque international en hommage à Jean Pernès. Publications du Bureau des Ressources Génétiques, pp. 352–368.
Bahrman, N., Plomion, C., Petit, R. J. and Kremer, A. (1997). Contribution of two-dimensional electrophoresis of proteins to maritime pine genetics. Ann Sci For, 54: 225–236.
Baradat, P. and Marpeau-Bezard, A. (1988). Le pin maritime, Pinus pinaster Ait. Biologie et génétique des terpènes pour la connaissance et l’amélioration de l’espèce. PhD Thesis, University of Bordeaux I.
Baradat, P., Marpeau, A. and Walter, J. (1991). Terpene markers. In: Müller-Starck, G. and Ziehe, M. (eds) Genetic Variation in European Populations of Forest Trees, pp. 40–66. Sauerländer’s Verlag, Frankfurt-am-Main.
Brown, A. H. D. and Weir, B. S. (1983). Measuring genetic variability in plant populations. In: Tanksley, S. D. and Orton, T. J. (eds) Isozymes in Plant Genetics and Breeding Part A, pp. 219–239. Elsevier Science Publishers, Amsterdam.
Burban, C., Petit, R., Carcreff, E. and Jactel, H. (1999). Rangewide variation of the maritime pine bast scale Matsucoccus feytaudi Duc. (Homoptera: Matsucoccidae) in relation to the genetic structure of its host. Mol Ecol, 10: 1593–1602, 10.1046/j.1365-294x.2001.01297.x.
Costa, P., Pot, D., Dubos, C., Frigerio, J. M. et al (2000). A genetic map of Maritime pine based on AFLP, RAPD and protein markers. Theor Appl Genet, 100: 39–48.
Doyle, J. J. and Doyle, J. L. (1990). Isolation of plant DNA from fresh tissue. Focus, 12: 13–15.
Durel, C. E. and Bahrman, N. (1995). Analyse de la diversité génétique des peuplements de Pin maritime de Corse. Gestion et exploitation de la ressource génétique. Rev For Fr, 47: 509–522.
Echt, C. S., May-Marquardt, P., Hseih, M. and Zahorchak, R. (1996). Characterization of microsatellite markers in eastern white pine. Genome, 29: 1102–1108.
Estoup, A., Tailliez, C., Cornuet, J. -M. and Solignac, M. (1995). Size homoplasy and mutational processes of interrupted microsatellites in two bee species, Apis mellifera and Bombus terrestris (Apidae). Mol Biol Evol, 12: 1074–1084.
Jactel, H., Ménassieu, P. and Burban, C. (1996). Découverte en Corse de Matsucoccus feytaudi Duc. (Homoptera: Margarodidae), cochenille du Pin maritime. Ann Sci For, 53: 145–152.
Jactel, H., Ménassieu, P., Ceria, A., Burban, C. et al (1998). Une pullulation de la cochenille Matsucoccus feytaudi provoque un début de dépérissement du Pin maritime en Corse. Rev For Fr, 50: 33–45.
Jenczewski, E., Prosperi, J. -M. and Ronfort, J. (1999). Differentiation between natural and cultivated populations of Medicago sativa (Leguminosae) from Spain: analysis with random amplified polymorphic DNA (RAPD) markers and comparison to allozymes. Mol Ecol, 8: 1317–1330, 10.1046/j.1365-294x.1999.00697.x.
Krutovskii, K. V., Erofeeva, S. Y., Aagaard, J. E. and Strauss, S. H. (1999). Simulation of effects of dominance on estimates of population genetic diversity and differentiation. J Hered, 60: 499–502.
Lynch, M. and Milligan, B. G. (1994). Analysis of population genetic structure with RAPD markers. Mol Ecol, 3: 91–99.
Mantel, N. (1967). The detection of disease clustering and a generalized regression approach. Cancer Res, 27: 209–220.
Mariette, S., Chagné, D., Decroocq, S., Vendramin, G. G. et al (2001). Microsatellite markers for Pinus pinaster Ait. Ann Sci For, 58: 203–206.
Nei, M. (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89: 583–590.
Nei, M. (1987). Molecular Evolutionary Genetics. Columbia University Press, New York.
Paquereau, M. M. (1964). Flores et climats post-glaciaires en Gironde. Actes Société Linnéenne Bordeaux, 101: 1–156.
Petit, R. J., Bahrman, N. and Baradat, P. H. (1995). Comparison of genetic differentiation in maritime pine (Pinus pinaster Ait.) estimated using isozyme, total protein and terpenic loci. Heredity, 75: 382–389.
Petit, R. J., El Mousadik, A. and Pons, O. (1998). Identifying population for conservation on the basis of genetic markers. Cons Biol, 12: 844–855.
Ribeiro, M. M., Plomion, C., Petit, R. J., Vendramin, G. G. et al (2001). Variation of chloroplast single-sequence repeats in Portuguese maritime pine (Pinus pinaster Ait.). Theor Appl Genet, 102: 97–103, 10.1007/s001220051623.
Salvador, L., Alía, R., Agúndez, D. and Gil, L. (2000). Genetic variation and migration pathways of maritime pine (Pinus pinaster Ait.) in the Iberian peninsula. Theor Appl Genet, 100: 89–95.
Smith, D. N. and Devey, M. E. (1994). Occurrence and inheritance of microsatellites in Pinus radiata. Genome, 37: 977–983.
Sokal, R. (1979). Testing statistical significance of geographic variation patterns. Syst Zool, 28: 227–232.
Sokal, R. and Rohlf, F. J. (1995) Biometry. Freeman, New York.
Tautz, D. (1989). Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucl Acids Res, 17: 6463–6471.
Vendramin, G. G., Anzidei, M., Madaghiele, A. and Bucci, G. (1998). Distribution of genetic diversity in Pinus pinaster Ait. as revealed by chloroplast microsatellites. Theor Appl Genet, 97: 456–463, 10.1007/s001220050917.
Vos, P., Hogers, R., Bleeker, M., Reijans, M. et al (1995). AFLP: a new technique for DNA fingerprinting. Nucl Acids Res, 23: 4407–4414.
Weber, J. L. and Wong, C. (1993). Mutation of human short tandem repeat. Hum Mol Genet, 2: 1123–1128.
Acknowledgements
We are very grateful for the technical help provided by Cécile Cabrero and Audrey Lartigue. We thank Dominique Charon and the French National Forest Office (ONF) for the help in collecting material in Aquitaine. This work was supported by grants from France (Ministère de l’Agriculture et de la Pêche-DERF no. 61.21.04/98 and Bureau des Ressources Génétiques) and the European Union (INCO, ERBIC-08CT-970200). The authors thank two anonymous reviewers and Don Stratton for their comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mariette, S., Chagné, D., Lézier, C. et al. Genetic diversity within and among Pinus pinaster populations: comparison between AFLP and microsatellite markers. Heredity 86, 469–479 (2001). https://doi.org/10.1046/j.1365-2540.2001.00852.x
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1046/j.1365-2540.2001.00852.x
Keywords
This article is cited by
-
Genotyping ex situ trees of Abies nebrodensis translocated from the original Sicilian population to enrich the gene pool
Journal of Forestry Research (2023)
-
Genetic variability of Araucaria angustifolia in the Argentinean Parana Forest and implications for management and conservation
Trees (2018)
-
Population structure and historical demography of Dipteronia dyeriana (Sapindaceae), an extremely narrow palaeoendemic plant from China: implications for conservation in a biodiversity hot spot
Heredity (2017)
-
Differential landscape effects on the fine-scale genetic structure of populations of a montane conifer from central Mexico
Tree Genetics & Genomes (2017)
-
Influence of ecological and geological features on rangewide patterns of genetic structure in a widespread passerine
Heredity (2015)