Abstract
Four to thirty-two copies of the rDNA 28S gene expansion segment D3 and flanking H14 stem were sequenced in six species of ticks (Ixodes: Ixodidae: Acari). Sequence match among species varied from 66% to 97%. Sequence length averaged 130 bases in I. persulcatus across eight Eurasian sites and averaged 186 bases in five other species across 19 Eurasian and North American sites. The difference in length represents one or more deletions totalling about 60 bases that correspond to stems S3 or S4 of the folded transcript. The typical transcript conformation was observed as one possible low energy structure in the five species of longer D3. The structure entails a basal loop with four stem/loop structures, S1–S4 (moving 5′ to 3′) atop stem H14. A secondary structure lacking S4 but possessing all other putative standard features of the D3 transcript is possible with the shorter I. persulcatus sequences. Interspecific sequence differences occur at higher frequency in loops and bulges vs. complementary pairing regions of stems. Insertion/deletion events (indels) and base substitutions accounted equally for sequence differences. Indels are flanked by similar sequences, suggesting that they occur by slippage during replication. The D3 of Ixodes species is composed of a degenerate set of subrepeats. Thus, unequal exchange among subrepeats may have caused the reduction in length of the I. persulcatus D3. Compensatory base substitution and compensatory insertion/deletion events are indicated by the failure of mutations to affect secondary structure. Transversions accounted for 64% of sequence differences and were biased toward the gain of G and U and the loss of A and C. This bias could re-establish intramolecular base pairing when disrupted by insertions or deletions that shift one side of a stem relative to the other. The distribution of sequence differences, biased substitution, and conservation of transcript conformation in D3 suggest selective constraint.
Similar content being viewed by others
Introduction
This report examines the nature and distribution of mutations in rDNA expansion segment D3 among six tick species (Ixodes: Ixodidae: Acari) with respect to the secondary structure of the folded transcript. Eukaryotic rDNA consists of arrays of a repeating unit that contains genes for 28S, 5.8S, and 18S rRNAs (Beckingham, 1982). The 28S rRNA gene is largely homologous to the prokaryote 23S rRNA gene but is larger owing to the presence of nonhomologous expansion segments that range from 10 to several hundred base pairs in length (Hancock et al., 1988). The number of expansion segments varies among eukaryotes (e.g. Kolosha & Fodor, 1990; Mercure et al., 1993) and is about a dozen in arthropods (e.g. Hancock & Dover, 1988; Gorab et al., 1995).
The function of some expansion segments, such as D3, is unknown. Others bind proteins during ribosome assembly (Chenuil et al., 1997) or may be required for transcript processing (Zarlenga & Dame, 1992). Yet, relaxed functional constraint may be inferred from observations that expansion segments evolve at up to 10× the rate of the gene core (Leffers & Andersen, 1993; Kuzoff et al., 1998) by accumulating point mutations (Waters et al., 1995) and changing length through the slippage-mediated multiplication or loss (Nunn et al., 1996; Vogler et al., 1997) of di- and tri-nucleotides (Holzman et al., 1996; Zarlenga et al., 1996).
The approximately 150-bp, AT-rich (usually) D3 expansion segment lies between highly conserved core sequences (Hancock & Dover, 1988). In spite of high rates of sequence evolution (Litvaitis et al., 1994), a characteristic secondary structure of the folded transcript is conserved (Gorab et al., 1995; Nunn et al., 1996). The typical RNA structure appears to consist of a central, basal loop with four intramolecularly bonded stems (and associated terminal loops) that sits atop a 14-bp stem, H14, which corresponds to sequences that flank the D3 sequence (Hancock et al., 1988). However, the validity of a folding pattern is uncertain when the sequence evolves rapidly (Gutell, 1993).
The present study examines sequence variation associated with intramolecular transcript folding in the D3 expansion segment of Ixodes affinis, I. pacificus, I. persulcatus, I. ricinus, I. scapularis and I. woodi. With the exception of I. woodi, all are sibling species of the I. ricinus complex. The distribution of insertions and deletions is mapped onto inferred transcript structures to determine if indels are limited to sequences that do not impact the intramolecular base pairing responsible for the formation of stems. Also, the direction of base substitution is inferred with reference to a phylogeny to examine if substitutions are biased in a way that conserves or restores intramolecular bonding.
Previously, sequencing of 4–32 copies of D3 in these six species revealed intraspecific variation among geographical isolates of I. persulcatus and substantial interspecific variation (McLain et al., 2001). Moreover, maximum likelihood, maximum parsimony, and neighbour-joining methods all supported a common phylogeny for the six tick species. I. woodi was inferred to be closer to the ancestral state. Geographical isolates of I. persulcatus formed one clade while I. affinis, I. pacificus, I. ricinus, and I. scapularis (two variants) formed another. Relative to other Ixodes species, I. persulcatus D3 sequences possess deletions totaling approximately 60 bases (McLain et al., 2001).
Materials and methods
Sequence alignment and folding
Details concerning (i) sample sizes, (ii) geographical origins, and (iii) DNA isolation, PCR amplification, subcloning, and dideoxy sequencing procedures are provided in McLain et al. (2001). Summary information is provided in Table 1 . D3 and H14 sequences were aligned with GENETIC DATA ENVIRONMENT (Smith et al., 1994) which uses the CLUSTAL-W program (Higgins et al., 1992) to maximize nucleotide identity of a primary structural alignment. Alignment parameters were set at 10 for both fixed and open gap penalties and for transition weighting. The phylogeny of sequences was inferred with maximum likelihood and maximum parsimony using PAUP v.4.0b3 (Swofford, 1999; Fig. 1 ). For these nonprotein-encoding sequences, PAUP treats gaps inserted into alignments as missing data, not as fifth characters.
Transcript folding patterns based on minimum energy structures were generated using RNAFOLD program of PC/GENE (IntelliGenetics, Inc. Mountain View, CA, USA). The folding program uses the method of Zuker & Steigler (1981) with energies as defined in Freier et al. (1986). RNAFOLD does not provide suboptimal, higher energy structures. Higher (but still relatively low) energy transcript structures were created by using RNAFOLD to fold one subsequence at a time, which prohibited intramolecular bonding between subsequences. This was done to determine whether the entire sequence could produce a structure with four different stem loops atop H14 as has been inferred for D3 of other species (e.g. Hancock et al., 1988). These structures – hereafter referred to as constrained structures – were modelled on the folding of a consensus sequence either for I. persulcatus populations or for all other Ixodes species. Consensus sequence structures were almost as stable as corresponding minimum energy structures.
Statistics
The observed and expected numbers of substitutions (transitions and transversions) resulting in the gain or loss of a given base were compared using chi-square (χ2) tests to determine whether substitutions were biased. The inferred direction of base change (e.g. A→G) was provided for 165 substitutions by PAUP (Swofford, 1999). Assuming transitions were no more likely to be conserved than transversions (see below), the expected number of losses of base j in transcripts was estimated as the product, s · pj, where pj is the proportional representation of that base and s is the number of substitutions of inferred direction. The expected number of gains of a base via substitutions at other bases was estimated as the quotient, s · (1 − pj)/(Σ[1 − pi])= s · (1 − pj)/3.
Proportional representation, pj, was [number of conserved base j in the ancestral transcript + number of independent base substitutions to j arising from mutations at other (nonconserved) bases] ÷ [number of G, A, U and C of the ancestral transcript that were conserved throughout the phylogeny + number of independent substitutions arising from all nonconserved bases]. The ancestral Ixodes D3 was taken to be the I. woodi sequence, based on its position in the phylogeny (Fig. 1), with the exception of nine bases. These nine bases represented cases where both the direction of sequence evolution, including small insertion/deletion events, was not inferred and where the non-I. woodi alternative produced a slightly lower energy folded transcript (see below). The ancestral I. persulcatus sequence was assumed to be the same as for other Ixodes species except as adjusted by insertions and deletions present in all geographical isolates of I. persulcatus.
The total sample size in χ2 tests equalled the denominator in the calculation of proportional representation. Thus, conserved descendant subsequences and shared mutations are not treated as independent subsamples (see Vawter & Brown, 1993).
An analysis was conducted to determine whether minimum energy conformations were significantly less consistent than constrained structures (see above) that were modelled on a constrained consensus sequence structure. The consensus sequence structure was of slightly higher energy than the minimum energy structure and resembled the D3 transcript structure inferred in other studies (e.g. Hancock et al., 1988). The analysis utilized all 66 positions in the alignment at which there were no deletions among the eight I. persulcatus sequences. The 5′ to 3′ distance between pairing partners in the constrained consensus sequence was subtracted from the corresponding distance in individual constrained structures to yield residuals. The same process was applied to the individual minimum energy structures relative to the minimum energy structure of the consensus sequence. Residuals for each nonpairing base were based on the mean position of other bases sharing its loop. The magnitude of the residuals was compared with a two-way ANOVA with sequence and structure (constrained or minimum energy) as categorical sources of variation. Residuals were smaller for structures more similar to the structure of the consensus sequence.
The number of Watson–Crick pairs (U·A, G·C), wobble pairs (G·U) and unpaired bases in the constrained structure of the consensus sequence was used to estimate the expected number of losses of paired bases and gains in unpaired bases as a function of the number of mutations inferred from the phylogeny. Simplifying assumptions were made with respect to calculations for indels: (i) insertions within a helix would cause a 1-base bulge, adding to the number of unpaired bases, without disturbing pairing of adjacent bases; (ii) complementary insertions opposite unpaired bases would form a pair, reducing the number of unpaired bases, regardless of adjacent substructure; (iii) the deletion of a paired base resulted in the loss of pairing by the undeleted partner, increasing the number of unpaired bases; (iv) indels were 1-base long; and (v) indels affected paired and unpaired bases in proportion to their frequency. These assumptions lead to a conservative statistical test because they slightly underestimate the effect of mutations on the loss of paired bases. Expected changes in the number of paired and unpaired bases as functions of the numbers of base substitutions and indels occurring in helical portions of stems were compared, using a χ2 test, to the observed number of changes summed across the set of sequences from which the consensus sequence was derived.
The expected numbers of changes in pairing status were calculated from the following consensus sequence values, p, the proportion of paired bases in stems, and q, the proportion of Watson–Crick pairs among paired bases, and from ND, NI, and NS, the numbers, respectively, of deletions, insertions, and base substitutions in helical portions of stem-loop structures as inferred from the phylogeny. The number of paired bases lost was calculated by (ND · p · 2) (= a) + (NS · p · q · 2 · [10/12]) (= b) + (NS · p · {1 − q} · 2 · [4/6]) (= c), while the number gained was calculated by (NI · (1 − p) · 2 [6/16]) (= d) + (NS (1 −p) 2 · [26/60]) (= e). Numbers in square brackets are rates at which base substitutions would cause a change in pairing status (see Hancock & Vogler, 1998) given that all bases were equally frequent as was the case for bases in helical portions of stems (χ32=1.567, P=0.710). Gains and losses in the number of unpaired bases were calculated as, respectively (a/2) + (NI − {d/2}) + b + c, and ND· (1 − p) + (d/2) + e.
Owing to the large number of statistical tests conducted, the sequential Bonferroni adjustment (Rice, 1989) was applied across all analyses to control type I error and to maintain an overall α-level of 0.05 for assessing statistical significance. All tests for which P < 0.05, indicating significance in isolation, had P-values that were sufficiently small to retain significance after application of the sequential Bonferroni adjustment.
Results
Sequence variation
No intraspecific sequence variation was found in Ixodes affinis, I. pacificus, I. ricinus, and I. woodi. Another species, I. scapularis, had two variant forms of D3 and H14 that occurred together at several North American collection sites (GA, NC, MD) and one or the other of these at remaining sites (see McLain et al., 2001). Sequences of I. persulcatus varied between but not within collection sites. Superimposing sequence variation over the maximum parsimony phylogeny of Fig. 1 yielded 179 base substitutions among five species and eight geographical isolates of a sixth species. Transversions dominated over transitions 115–64. The proportion of transversions in the I. persulcatus clade (43/64=0.672) was not different from the proportion in the remainder of the phylogeny (72/115=0.626) (χ12=0.528, P=0.425).
Aligned D3 plus H14 sequences matched to a variable degree among Ixodes species (where percentage match=100 · number of matches/length of shorter sequence). The phylogenetically basal Ixodes woodi sequence (Fig. 1) matched sequences of the eight geographical isolates of I. persulcatus at an average level of 66.0% (88/133.4; SD=4.2%) but matched sequences of the other four Ixodes species at 90.6% of their bases (168.2/185.8; SD=4.8%). Within the I. persulcatus clade the average level of matching was only 82.0% (106.4/129.7; SD=7.5%) compared to 88.2% (163.6/185.7; SD=5.9%) within the multispecies clade (Fig. 1). Between the two clades, sequences matched at an average level of 67.1% (88.8/132.3; SD=3.6%; N=40).
Given differences in sequence length between the clades [129.7 (for I. persulcatus) – 185.7 (for other Ixodes species) = 56], gaps were necessarily inserted into I. persulcatus sequences to maximize the degree of base matching with other Ixodes species. Most of the difference in sequence length could be accounted for by a single large deletion. Three almost equally precise alignments, in terms of base matching and number of indels, are possible between the 3′ ends of the D3 consensus sequences for I. persulcatus geographical isolates and the other Ixodes species (Table 2 ). These alternative alignments exposed a large deletion in the I. persulcatus D3 of either 54, 45, or 56 base pairs. However, the alignment maximizing percentage match among all sequences has six deletions of 1–29 bases instead of a single large deletion at the 3′ end (Fig. 2 ).
Ancestral Ixodes spp. D3 and H14 transcript. (A) Location of mutations (excepting I. persulcatus). Arrows point to positions of base substitutions. Wedges and numbers indicate positions and sizes of insertions. Encircled bases indicate where deletions occurred. Stems S1–S4 and H14 are labelled. (B) Transcript sequence that would remain after deletions (indicated by arrows with number of deleted bases) and insertions (indicated by pie sectors) that characterize I. persulcatus.
The observation that the deletion in the I. persulcatus sequence could be placed at different points along the 3′ end of the multispecies consensus suggested that the D3 of Ixodes species may be composed of repeated subsequences. Internal alignment of consensus sequences of geographical isolates of I. persulcatus and other Ixodes species indicated the possible presence of up to five mutationally degenerated repeats that together would constitute the entire D3 (Table 3 ).
The minimum number of insertion/deletion events (indels) suggested by the phylogeny was 89 and encompassed 127 bases. Most indels (71=79.8%) were only 1-bp long. The number of these that were flanked on one or both sides by the same base (29=40.8%) was not different than the number expected by chance (= 31.62, using binomial expansion and adjusting for base composition; χ21=0.55, P=0.407). However, five of eight indels of 2 bp were flanked by the same sequence while and nine of 10 indels of 3–5 bp were flanked by the same sequence or a similar sequence (= same for all but one base). The frequency with which indels of greater than 1 bp matched flanking sequences (14/18=77.8%) was significantly greater than that expected by chance (= 27.3%) (χ21=23.05, P < 0.001 using binomial expansion and adjusting for base composition).
Intramolecular base pairing
For all Ixodes species except I. persulcatus, the minimum energy conformation (≈−74 kcal mol−1) of the D3 transcript contained 2 simple stem loops and one bifurcated stem atop the non-D3 stem, H14 (Fig. 3 ). This folded structure did not conform to the standard structure, which is four simple stem loops associated with a common basal loop atop the H14 stem (Hancock et al., 1988). However, the standard structure was obtained in a slightly higher energy conformation (≈−60 kcal mol−1) simply by unzipping complementary pairs at the base of the bifurcated stem (Fig. 3). This structure was very similar in all Ixodes species less I. persulcatus (Fig. 4 ). Now, the folded transcript had 13–14 complementary base pairs in the H-14 stem that supported four stem loops (S1, S2, S3 and S4 moving 5′ to 3′). Interior bulges occurred in S1 (1 bulge) and S3 (2 bulges). S2, always the smallest, occurred relatively close to S3. S1 and S4 were always relatively far from, respectively, S2 and S3. These features were very similar to those described for D. melanogaster (Hancock et al., 1988) except that in the fruit fly it is S4, not S3, that contains two interior bulges.
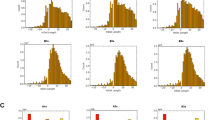
Minimum energy conformations of I. persulcatus transcripts (≈−45 kcal mol−1) did not resemble those of other Ixodes species and did not show much correspondence between geographical isolates within the species (Fig. 5 ). In some cases (e.g. sequences representing western Russia and the Altai Mountains), the folding pattern did not result in a series of stem loops associated with a central, basal loop. Constrained structures (≈−38 kcal mol−1) of I. persulcatus sequences were based on the assumption that a single large deletion resulted in the loss of bases corresponding to the 3′ end of S3 and virtually all of S4 (of other Ixodes species; Fig. 4). With sequences aligned, homologous bases form stems 1–3 for both constrained structures of I. persulcatus and constrained structures of other Ixodes species (Table 2). Even the bases of terminal loops are largely conserved between transcript structures of I. persulcatus and the other Ixodes species.
Constrained structures of I. persulcatus were significantly more similar to each other than were corresponding minimum energy structures (F1,910=352.621, P < 0.001; Fig. 5). Sequences from western Russia and the Altai Mountains were significantly more similar to consensus structures than were other sequences (Tukey HSD post-hoc pairwise comparisons, P < 0.001). These other sequences did not differ significantly among themselves with respect to similarity to consensus structures.
Sequence variation vs. transcript conformation
The number of intramolecular hydrogen-bonded bases in low-energy conformations of transcripts of Ixodes species (other than I. persulcatus) averaged 120 (64% of total) while the number of bases in loops and interior bulges averaged 66.5 (36% of total). However, based on the folded transcript of the sequence inferred to be ancestral to I. ricinus complex members, base substitutions have been more likely to occur within 65 bases corresponding to loops and bulges of the transcript instead of within 124 bases corresponding to complementary, intramolecular base-paired portions of transcript stems (χ21=10.566, P=0.001; Fig. 2). Indels also were observed at a higher frequency in bases corresponding to loops, bulges, and junctures between loops and stems rather than in bases corresponding to portions of stems with intramolecular base paring (χ21=12.641, P < 0.001).
In Ixodes species (less I. persulcatus) base substitutions were not more likely to be observed at positions corresponding to sequences deleted in I. persulcatus (χ21=0.004, P=0.956). However, indels were much more likely to occur at bases corresponding to those deleted from I. persulcatus sequences (χ21=26.468, P < 0.001). Most of these deletions corresponded, in the folded transcript, to a portion of S3, the loop region between S3 and S4, and a portion of S4 (Fig. 2).
In the approximately 90 bases of helical portions of S1, S2, and S3 of eight I. persulcatus sequences, 18 base substitutions and 31 indels were observed. Using constrained structures modelled after the constrained consensus sequence (Fig. 3), these mutations were expected to result in the overall loss of 48.17 paired bases (= 3.01 pairs/sequence) and a gain of 31.47 unpaired bases. However, the observed net loss of only 12 paired bases (= 0.75 pairs/sequence) and a net ‘gain’ of –3 unpaired bases were significantly less than expected (χ21=64.915, P < 0.001). Consequently, the difference between the consensus and individual constrained structures in the number of paired bases in S1, S2, and S3 was not significantly different from 0 (t23=0.598, P=0.405).
Among the five other Ixodes species, 10 base substitutions and 20 indels occurred in the approximately 90 bases of helical portions of stems 1–4. These mutations were expected to result in the loss of 19.48 pairing bases and a gain of 27.65 unpaired bases. However, there was a gain of 1.18 pairing bases and 4.84 unpaired bases (χ21=40.729, P < 0.001).
Base composition
The GC content of the combined D3 and H14 stem of I. persulcatus (59.3%) did not differ from that of other Ixodes species (63.5%; χ23=1.062, P=0.567). The base composition of helical portions of transcript stems was significantly different from that of loops and bulges (χ23=15.529, P=0.003). Loops and bulges were rich in A (36.5%) but had very little U (6.3%). In contrast, stems had relatively little A (14.3%) and relatively more U (19%). The percentage of G was slightly higher in stems than in loops and bulges (35.7% vs. 25.4%) while the percentage of C was similar (31.0% vs. 31.7%).
Within the I. persulcatus clade, substitutions did not result in significant differences in the loss of some bases relative to others (χ23=6.137, P=0.132; Tables 4, 5 . Also, substitutions did not result in significant differences in the gain of some bases relative to others (χ23=1.284, P=0.484). However, elsewhere in the phylogeny, substitutions resulted in significant differential loss of some bases relative to others (χ23=15.285, P=0.003). Bases C and A were lost from the transcript at higher rates than predicted by chance while bases G and U were lost at lower rates than predicted by chance (Table 5). Also, substitutions resulted in significant differential gain of some bases relative to others (χ23=14.150, P=0.004; Tables 4, 5. Bases G and, to a lesser extent, U were gained at higher than expected rates while bases C and, to a lesser extent, A were gained at lower than expected rates (Table 5).
The mutational bias toward transversions did not differ among bases corresponding to either helical portions of stems or to loops of the folded transcript (χ21=0.420, P=0.532). Also, the mutational bias toward accumulation of G and U did not vary among bases contributing either to helical portions of stems or to loops (χ21=1.098, P=0.190).
Discussion
Alignment and phylogeny
Phylogenetic inference is influenced by sequence length, the rate of base substitution, and the method used to achieve homologous nucleotide alignment (Mugridge et al., 2000). Alignment based on conserved features of secondary structure is advocated (Mugridge et al., 2000) except where sequence evolution is significantly affected by replication slippage which obscures homology (Hancock & Vogler, 2000). Replication slippage appears to account for about half of the sequence variation observed among Ixodes species. Thus, alignment by primary structure is justified. However, the D3 plus H14 sequence is quite small and any phylogeny based solely on this is suspect.
Two major features of the inferred phylogeny are relevant to conclusions drawn from some of the forgoing analyses, the placement of I. woodi and the grouping of I. persulcatus sequences into a single clade. I. woodi is not considered a member of the same subgenus as the other species based on morphology (Robbins & Keirans, 1992). Thus, its position in the current phylogeny is supported independently. Moreover, I. persulcatus is considered a single, widespread species on morphological grounds (Filippova, 1991). In the phylogeny, most sequence variation occurred along branches from I. woodi to I. persulcatus and from the I. persulcatus clade to the multispecies clade. However, given the uncertain homology of much of the I. persulcatus sequence, doubt pertains to the placement of the I. persulcatus clade. Sequences were very similar between I. woodi and Ixodes species other than I. persulcatus, and especially between Ixodes species of the multispecies clade (Fig. 1). Thus, any placement of the I. persulcatus clade within the multispecies clade would have only minor quantitative effects on analyses.
Sequence variation
The percentage match of D3 and H14 sequences among tick species varies from a low of 66.0% (I. woodi: I. persulcatus) to a high of 97.2% (I. ricinus: I. scapularis). Surprisingly, the percentage match among geographical isolates of I. persulcatus is less than that for any pair-wise combination among the other five species. However, the relatively low degree of sequence conservation in I. persulcatus may not be associated with highly variable secondary structure of the folded transcript. It is possible to produce low, but not minimum, energy structures that lack stem 4 but that otherwise match the structure of the D3 transcript inferred for other species (e.g. Hancock et al., 1988). Principal features of secondary structure appear to be conserved among the other Ixodes species and their geographical isolates where the base sequence is also conserved.
Based on the phylogeny, base substitutions have occurred at about twice the rate of indels (see Hancock, 1995). However, as over 1/4 of the indels were greater than one base in length, substitutions and indels affected a similar number of bases. Indels of variable rDNA sequences may represent mutations caused by slippage (Hancock, 1995; Hancock & Vogler, 2000) that occurs during replication (Levinson & Gutman, 1987). Replication slippage is suggested by the observation that indels are often flanked by one or more similar sequences. Indels were more common toward the 3′ end of the expansion segment, especially where the ancestral I. persulcatus sequence suffered one or more deletions encompassing about 50 base pairs. Most indels of the tick D3 are mono- or dinucleotides. This is observed in other hypervariable regions of rDNA (Hancock & Vogler, 2000).
A deletion in I. persulcatus is a more parsimonious explanation for variation in D3 length than is a duplication in the other species. The position of I. persulcatus in the phylogeny would require the duplication to have arisen twice. Also, the potential to form the more conventional secondary structure with the D3 transcript of tick species other than I. persulcatus is consistent with the retention of an ancient conformation.
The loss of over 1/3 of the D3 sequence in I. persulcatus may have occurred via unequal crossing over (Coen & Dover, 1983). The putative recognition of now-degenerate subrepeats of about 45 base pairs suggests the potential for a relatively small-scale unequal crossover event. The deletion may also have been facilitated by ability of the subrepeats to foldback during DNA replication.
RNA folding patterns
With the exception of I. persulcatus, D3 and H14 transcripts of Ixodes species can assume very similar minimum and low energy conformations. The low, but not minimum, energy conformation (= constrained structure) contains all standard secondary structural features hypothesized to characterize eukaryotes (Michot & Bachellerie, 1987; Michot et al., 1990). Conserved secondary structures need not be minimum energy structures, but they are usually close in stability to minimum energy structures (Hancock & Vogler, 1998). In Ixodes species, the constrained structure may need to be stabilized by protein–RNA interactions (Brimacombe et al., 1983).
Alternative alignments of the I. persuslcatus consensus sequence with that of other Ixodes species (Table 2) indicates the loss in I. persuslcatus of sequences corresponding to one or the other of these structural features: the 3′ half of S3 through the 5′ half of S4 or the central loop region between S3 and S4 plus all of S4. Constrained structures have stems 1–3 that are homologous to the same stems of transcripts of other Ixodes species. However, no homologous stem 4 is present in constrained structures of I. persulcatus. Loss of S4 would be consistent with observations of some other organisms (Michot et al., 1990; Nunn et al., 1996). S4 may also differ enormously in length among closely related arthropods (Nunn et al., 1996). Thus, S4 may not be not critical to either ribosome function or the ability to bury D3 in the whole of a folded 28S transcript.
The general lack of resemblance between minimum energy secondary structures of I. persuslcatus geographical isolates argues against the structural significance of these folded transcripts. Perhaps, their small size relaxes selection on particular transcript features that are otherwise necessary so that the D3 will not interfere with ribosome function (Clark, 1987; Hancock & Dover, 1988). Alternatively, the functional I. persulcatus D3 transcript may assume a higher energy structure. The constrained transcript structures are similar in spite of significant variation in the primary sequence. Moreover, constrained structures retain features inferred to be present in a variety of organisms (Michot & Bachellerie, 1987; Hancock et al., 1988; Michot et al., 1990; Nunn et al., 1996). Thus, for a given set of homologous sequences, the actual secondary structures may be those of the lowest energies that permit structural consistency while encompassing all primary sequence variation.
Compensatory base substitutions and compensatory insertions/deletions (Hancock & Dover, 1990) can maintain features of secondary structure of expansion segment transcripts and may indicate selection for conservation of those features (Gorab et al., 1995). However, the slippage-mediated gain and loss of short sequences can be neutral with respect to overall rates of base pairing, suggesting that folded secondary structures arise as an intrinsic property of rRNA transcripts (Hancock & Vogler, 2000).
If constrained structure models are correct, large numbers of mutations do not appear to have significantly affected the number of paired and unpaired bases in either the I. persulcatus or Ixodes multispecies clades. As base substitutions and indels are approximately equally likely in helical portions of stems, both compensatory substitutions (Hancock & Vogler, 1998) and compensatory indels (Hancock & Dover, 1990) are indicated. Covariation between sites in a sequence that maintains secondary structure suggests the action of natural selection. However, selection need not imply a functional role (Gerbi, 1986). D3 secondary structure may reflect function or the necessity of folding in a manner that does not impair ribosome function.
Selection is further indicated by the observation that mutations are concentrated in loops and bulges where secondary structure should be relatively less impacted. Thus, the location and size of loops is largely conserved among species. For example, on average 66 of 186 bases occur in loops and bulges of the folded transcript. Yet, the standard deviation about this mean is only 3.2 bases, or 1.7% of total sequence length. Thus, both the distribution of mutations and their failure to reduce base pairing argues for selection to preserve the characteristic conformation of the transcript.
Base composition
Base composition of D3 can be as high as 85% GC (Chan et al., 1983). However, with 51–62% GC, Ixodes species are unusual among invertebrates, which generally have a GC content of 30–35% (Nunn et al., 1996). In some organisms the GC content of genomes is greater in high temperature environments (Bernardi et al., 1988), possibly because hydrogen bonding between GC pairs is more stable than for AT pairs at higher temperatures (Wada & Suyama, 1986; Nunn et al., 1996). However, this would not appear to explain high GC content in these ticks which occur predominately in temperate regions. The high GC content of Ixodes extends to sequences flanking D3 and H14. For instance, the GC composition of the approximately 180 base pairs between D3 and D4 is 48.0–52.6% GC in the six Ixodes species (unpubl. data). Thus, it may not be the GC content of D3 in particular that is high but, rather, some larger part of the 28S rRNA gene.
Loops and bulges of some rRNA transcripts are markedly AU-biased (McLain et al., 1995; Vawter & Brown, 1993). In D3 transcripts this may represent a compositional constraint (Nunn et al., 1996) associated with ribosomal protein recognition (Peattie et al., 1981). However, this pattern is not present in Ixodes species where loops and bulges are A-rich but U-poor. This contrasts with Ixodes species stems which are A-poor. Also, compared to loops and bulges stems have more G and U.
These patterns may reflect selection to maintain a high degree of intramolecular base pairing in transcript stems while reducing this potential in loops and bulges. Adenine pairs only with U and even then only forms two hydrogen bonds. Therefore, subsequences rich in A are unlikely to form stable stems if subjected to frequent insertions or deletions. Stem-forming regions with higher percentages of G and U may be able to continue to base pair even if noncompensating insertions or deletions occur. This is because G could hydrogen bond with either C or U while U could hydrogen bond with either A or G if an indel caused one side of a stem to slip relative to the other. Also, alternate stable configurations might be possible for stem regions enriched with G and U. For instance, minimum and low energy conformations of the Ixodes D3 are very different but retain high levels of intramolecular base pairing.
Biased base substitution
If base substitutions were biased toward accumulation of G and U, the effect would be to better enable intramolecular base pairing where it was absent (Fig. 6 ). In fact, there are 14 ways in which substitutions to G or U would establish base pairing but only 6 ways in which substitutions to A or C would do so (Fig. 6). Moreover, of 20 substitutions establishing base pairing, 18 are transversions and only 2 are transitions.
Base substitutions capable of preserving (upper set of boxes) or establishing (lower set) complementary base pairing. The two middle rows represent bases (pairing or not) that are opposite each other in a stem. An ellipse in a box indicates that a mutation from the base in the row to the base listed above the column would result in a hydrogen-bonded pair.
In the tick D3, transversions were almost twice as likely to account for sequence evolution as were transitions. In addition, substitutions favoured the accumulation of G and U (except in the I. persulcatus clade). The bias toward transversions or toward greater G and U content of tick transcripts appears to be a consequence of mutational processes because both helical portions of stems and loops exhibit similar biases. Thus, it appears that random sequence evolution could promote intramolecular base pairing and compliment any selective pressures favouring low energy secondary transcripts. However, only transitions can preserve complementary pairing where it already exists (Fig. 6). Also, with complementary base pairing, no substitution bias would be advantageous if all four bases were equally frequent in the transcript.
Studies of selectively neutral pseudogenes and other noncoding sequences indicate that the mutational process may be biased (Kvarnheden et al., 1998) and may have a significant effect on base composition (Petrov & Hartl, 1999). In fact, the local base composition itself may impact the process (Morton, 1997). DNA repair may also be biased toward certain substitutions (Bouzekri et al., 1998). Substitution bias often favours the accumulation of T (Lobuglio et al., 1993; Dowton & Austin, 1997; Kvarnheden et al., 1998; Petrov & Hartl, 1999) as was observed in Ixodes ticks.
Conclusions
The results indicate that low-energy folding of the D3 expansion segment of most Ixodes tick species can produce the hypothesized standard eukaryote folded transcript structure. In I. persulcatus, a similar secondary structure can be produced with the transcript but it lacks stem 4, apparently owing to a deletion of approximately 50 base pairs. Much smaller insertions and deletions occur at a high rate. These could cause opposite sides of transcript stems to slide relative to each other. Compensatory indels and mutational biases toward transversions and substitution of G and U for A and C may promote the re-establishment of intramolecular pairing in helical portions of transcript stems.
References
Beckingham, K. (1982). Insect rDNA. Cell Nucl, 10: 205–263.
Bernardi, G., Mouchiroud, D. and Gautier, C. (1988). Compositional patterns in vertebrate genomes. conservation and change in evolution. J Mol Biol, 28: 7–18.
Bouzekri, N., Taylor, P. G., Hammer, M. F. and Jobling, M. A. (1998). Novel mutation processes in the evolution of a haploid minisatellite, MSY1: array homogenization without homogenization. Hum Mol Genet, 7: 655–659.
Brimacombe, R., Maly, P. and Zwieb, C. (1983). The structure of ribosomal RNA and its organization relative to ribosomal protein. Nucl Acids Res Mol Biol, 28: 1–48.
Chan, Y. -L., Olvera, J. and Wool, I. G. (1983). The structure of rat 28S ribosomal ribonucleic acid inferred from the sequence of nucleotides in a gene. Nucl Acids Res, 11: 7819–7831.
Chenuil, A., Solignac, M. and Bernard, M. (1997). Evolution of the large-subunit ribosomal RNA binding site for protein L23/25. Mol Biol Evol, 14: 578–588.
Clark, C. G. (1987). On the evolution of ribosomal RNA. J Mol Biol, 25: 343–350.
Coen, E. S. and Dover, G. A. (1983). Unequal crossing over and the coevolution of X and Y arrays in D. melanogaster. Cell, 33: 849–855.
Dowton, M. and Austin, A. D. (1997). Evidence for AT-transversion bias in wasp (Hymenoptera. Symphyta) mitochondrial genes and its implications for the origin of parasitism. J Mol Evol, 44: 398–405.
Filippova, N. A. (1991). The Ixodes ricinus complex. Modern Acarol, 1: 109–118.
Freier, S. M., Kierzek, R., Jaeger, J. A., Subimoto, N., Caruthers, M. H. and Neilson, T. et al.1986). Improved free-energy parameters for predictions of RNA duplex stability. Proc Natl Acad Sci USA, 83: 9373–9377.
Gerbi, S. (1986). The evolution of eukaryotic ribosomal DNA. Biosystems, 19: 247–258.
Gorab, E., Lacoba, M. and Botella, L. M. (1995). Structural constraints in expansion segments from a midge 26S rDNA. J Mol Evol, 41: 1016–1021.
Gutell, R. R. (1993). Collection of small subunit (16S- and 16S-like) ribosomal RNA proteins. Nucl Acids Res, 21: 3051–3054.
Hancock, J. M. (1995). The contribution of DNA slippage to eukaryotic nuclear 18S rRNA evolution. J Mol Evol, 40: 629–639.
Hancock, J. M. and Dover, G. A. (1988). Molecular coevolution among cryptically simple expansion segments in eukaryotic 26S/28S rRNA. Mol Biol Evol, 5: 377–392.
Hancock, J. M. and Dover, G. A. (1990). ‘Compensatory slippage’ in the evolution of ribosomal RNA genes. Nucl Acids Res, 18: 5949–5954.
Hancock, J. M. and Vogler, A. P. (1998). Modelling the secondary structures of slippage-prone hypervariable RNA regions: the example of the tiger beetle 18S rRNA variable region V4. Nucl Acids Res, 26: 1689–1699.
Hancock, J. M. and Vogler, A. P. (2000). How slippage-derived sequences are incorporated into rRNA variable-region secondary structure: implications for phylogeny reconstruction. Mol Phyl Evol, 14: 366–374.
Hancock, J. M., Tautz, D. and Dover, G. A. (1988). Evolution of the secondary structures and compensatory mutations of the ribosomal RNAs of Drosophila melanogaster. Mol Biol Evol, 5: 393–414.
Higgins, D. G., Bleasby, A. J. and Fuchs, R. (1992). Clustal-W: improved software for multiple sequence alignment. Comput Appl Biosci, 8: 189–191.
Holzmann, M., Piller, W. and Pawlowski, J. (1996). Sequence variations in the large-subunit RNA gene of Ammonia (Foraminifera, Protozoa) and their evolutionary implications. J Mol Evol, 43: 145–151.
Kolosha, V. O. and Fodor, I. (1990). Nucleotide sequence of Citrus limon 26S rRNA gene and secondary structure model of its RNA. Plant Mol Biol, 14: 147–161.
Kuzoff, R. K., Swere, J. A., Soltis, D. E. and Zimmer, E. A. (1998). The phylogentic potential of the entire 26S rDNA sequences in plants. Mol Biol Evol, 15: 251–263.
Kvarnheden, A., Albert, V. A. and Engstroem, P. (1998). Molecular evolution of cdc2 pseudogenes in spruce (Picea). Plant Mol Biol, 36: 767–774.
Leffers, H. and Andersen, A. H. (1993). The sequence of 28S ribosomal RNA varies within and between human cell lines. Nucl Acids Res, 21: 1449–1455.
Levinson, G. and Gutman, G. A. (1987). Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol Biol Evol, 4: 203–221.
Litvaitis, M. K., Nunn, G., Thomas, W. K. and Kocher, T. D. (1994). A molecular approach for the identification of meiofaunal turbellarians (Platyhelminthes, Turbellaria). Mar Biol, 120: 437–422.
Lobuglio, K. F., Pitt, J. I. and Taylor, J. W. (1993). Phylogenetic analysis of two ribosomal DNA regions indicates multiple independent losses of a sexual Talaromyces state among asexual Penicillium species in subgenus Biverticillium. Mycologia, 85: 592–604.
Mclain, D. K., Li, J., Oliver, J. H. Jr (2001). Interspecific and geographic variation in the sequence of rDNA expansion segment D3 of Ixodes ticks (Acari: Ixodidae). Heredity, 86: 234–242, 10.1046/j.1365-2540.2001.00835.x.
Mclain, D. K., Wesson, D. M., Collins, F. H., Oliver, J. H. Jr (1995). Evolution of the rDNA spacer, ITS 2, in the ticks Ixodes scapularis and I. pacificus (Acari. Ixodidae). Heredity, 75: 303–319.
Mercure, S., Rougeau, N., Montplaisir, S. and Lemay, G. (1993). The nucleotide sequence of the 25S rRNA-encoding gene from Cadida albicans. Nucl Acids Res, 21: 1490–1490.
Michot, B. and Bachellerie, J. -P. (1987). Comparisons of large sub-unit rRNAs reveal some eukaryote-specific elements of secondary structure. Biochimie, 69: 11–23.
Michot, B., Qu, L. -H. and Bachellerie, J. -P. (1990). Evolution of large sub-unit rRNA structure. The diversification of divergent D3 domain among major phylogenetic groups. Eur J Biochem, 188: 219–229.
Morton, B. R. (1997). The influence of neighboring base composition on substitutions in plant chloroplast coding sequences. Mol Biol Evol, 14: 189–194.
Mugridge, N. B., Morrison, D. A., Jäkel, T., Heckroth, A. R., Tenter, A. M. and Johnson, A. M. (2000). Effects of sequence alignment and structural domains of ribosomal DNA on phylogeny reconstruction for the protozoan family Sarcosystidae. Mol Biol Evol, 17: 1842–1853.
Nunn, G. B., Theisen, B. F., Christensen, B. and Arctander, P. (1996). Simplicity-correlated size growth of the nuclear 28S ribosomal RNA D3 expansion segment in the crustacean order Isopoda. J Mol Evol, 42: 211–223.
Peattie, D. A., Douthwaite, S., Garrett, R. A. and Noller, H. F. (1981). A ‘bulged’ double helix in a RNA-protein contact site. Proc Natl Acad Sci USA, 78: 7331–7335.
Petrov, D. A. and Hartl, D. L. (1999). Patterns of nucleotide substitution in Drosophila and mammalian genomes. Proc Natl Acad Sci USA, 96: 1475–1479.
Rice, W. R. (1989). Analyzing tables of statistical tests. Evolution, 43: 223–225.
Smith, S. W., Overbeek, R., Woese, C. R., Gilbert, W. and Gillvet, P. M. (1994). The genetic data environment and expandable GUI for multiple sequence analysis. CABIOS, 10: 671–675.
Swofford, D. L. (1999). PAUP: phylogenetic analysis using parsimony. Illinois Natural History Survey, Champaign, IL.
Vawter, L. and Brown, W. M. (1993). Rates and patterns of base change in the small subunit ribosomal RNA gene. Genetics, 134: 597–608.
Vogler, A. P., Welsh, A. and Hancock, J. M. (1997). Phylogenetic analysis of slippage-like sequence variation in the V4 rRNA expansion segment in tiger beetles (Cicindelidae). Mol Biol Evol, 14: 6–19.
Wada, A. and Suyama, A. (1986). Local stability of DNA and RNA secondary structure and its relation to biological function. Prog Biophys Mol Biol, 47: 113–157.
Waters, A. P., White, W. and McCutchan, T. F. (1995). The structure of the large subunit rRNA expressed in blood stages of Plasmodium falciparum. Mol Biochem Parasitol, 72: 227–237.
Zarlenga, D. S. and Dame, J. B. (1992). The identification and characterization of a break within the large subunit ribosomal RNA of Trichinella spiralis: comparison of gap sequences within the genus. Mol Biochem Parasitol, 51: 281–290.
Zarlenga, D. S., Aschenbrenner, R. A. and Lichtenfels, J. R. (1996). Variations in microsatellite sequences provide evidence for population differences and multiple ribosomal gene repeats within Trichinella pseudospiralis. J Parasitol, 82: 534–538.
Zuker, M. and Steigler, P. (1981). Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucl Acids Res, 9: 133–148.
Acknowledgements
This work was supported by National Institutes of Health Grant 1R15 AI 34136–01. I thank referee Dr J. M. Hancock for advise on developing plausible transcript structures and for recommendations for data analysis.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mclain, D. Evolution of transcript structure and base composition of rDNA expansion segment D3 in ticks. Heredity 87, 544–557 (2001). https://doi.org/10.1046/j.1365-2540.2001.00943.x
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1046/j.1365-2540.2001.00943.x