Abstract
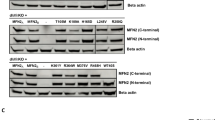
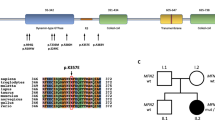
Charcot–Marie–Tooth disease (CMT) is a group of hereditary peripheral neuropathies. The dominantly inherited axonal CMT2 displays striking genetic heterogeneity, with 17 presently known disease genes. The large number of candidate genes, combined with lack of genotype–phenotype correlations, has made genetic diagnosis in CMT2 time-consuming and costly. In Finland, 25% of dominant CMT2 is explained by either a GDAP1 founder mutation or private MFN2 mutations but the rest of the families have remained without molecular diagnosis. Whole-exome and genome sequencing are powerful techniques to find disease mutations for CMT patients but they require large amounts of sequencing to confidently exclude heterozygous variants in all candidate genes, and they generate a vast amount of irrelevant data for diagnostic needs. Here we tested a targeted next-generation sequencing approach to screen the CMT2 genes. In total, 15 unrelated patients from dominant CMT2 families from Finland, in whom MFN2 and GDAP1 mutations had been excluded, participated in the study. The targeted approach produced sufficient sequence coverage for 95% of the 309 targeted exons, the rest we excluded by Sanger sequencing. Unexpectedly, the screen revealed a disease mutation only in one family, in the HSPB1 gene. Thus, new disease genes underlie CMT2 in the remaining families, indicating further genetic heterogeneity. We conclude that targeted next-generation sequencing is an efficient tool for genetic screening in CMT2 that also aids in the selection of patients for genome-wide approaches.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Szigeti K, Lupski JR : Charcot-Marie-Tooth disease. Eur J Hum Genet 2009; 17: 703–710.
Lupski JR, de Oca-Luna RM, Slaugenhaupt S et al: DNA duplication associated with Charcot-Marie-Tooth disease type 1A. Cell 1991; 66: 219–232.
Shy ME, Patzko A : Axonal Charcot-Marie-Tooth disease. Curr Opin Neurol 2011; 24: 475–483.
Saporta AS, Sottile SL, Miller LJ, Feely SM, Siskind CE, Shy ME : Charcot-Marie-Tooth disease subtypes and genetic testing strategies. Ann Neurol 2011; 69: 22–33.
Auranen M, Ylikallio E, Toppila J, Somer M, Kiuru-Enari S, Tyynismaa H : Dominant GDAP1 founder mutation is a common cause of axonal Charcot-Marie-Tooth disease in Finland. Neurogenetics 2013; 14: 123–132.
Choi BO, Koo SK, Park MH et al: Exome sequencing is an efficient tool for genetic screening of Charcot-Marie-Tooth Disease. Hum Mutat 2012; 33: 1610–1615.
Lupski JR, Reid JG, Gonzaga-Jauregui C et al: Whole-genome sequencing in a patient with Charcot-Marie-Tooth neuropathy. N Engl J Med 2010; 362: 1181–1191.
Weedon MN, Hastings R, Caswell R et al: Exome sequencing identifies a DYNC1H1 mutation in a large pedigree with dominant axonal Charcot-Marie-Tooth disease. Am J Hum Genet 2011; 89: 308–312.
Sulonen AM, Ellonen P, Almusa H et al: Comparison of solution-based exome capture methods for next generation sequencing. Genome Biol 2011; 12: R94.
Engelfried K, Vorgerd M, Hagedorn M et al: Charcot-Marie-Tooth neuropathy type 2A: novel mutations in the mitofusin 2 gene (MFN2). BMC Med Genet 2006; 7: 53.
Braathen GJ, Sand JC, Lobato A, Hoyer H, Russell MB : MFN2 point mutations occur in 3.4% of Charcot-Marie-Tooth families. An investigation of 232 Norwegian CMT families. BMC Med Genet 2010; 11: 48.
Albulym OM, Zhu D, Reddel S, Kennerson M, Nicholson G : The MFN2 V705I variant is not a disease-causing mutation: a segregation analysis in a CMT2 family. J Neurodegenerative Dis 2013; 2013: doi:10.1155/2013/495873.
Evgrafov OV, Mersiyanova I, Irobi J et al: Mutant small heat-shock protein 27 causes axonal Charcot-Marie-Tooth disease and distal hereditary motor neuropathy. Nat Genet 2004; 36: 602–606.
Osman C, Merkwirth C, Langer T : Prohibitins and the functional compartmentalization of mitochondrial membranes. J Cell Sci 2009; 122: 3823–3830.
Faber CG, Hoeijmakers JG, Ahn HS et al: Gain of function Nanu1.7 mutations in idiopathic small fiber neuropathy. Ann Neurol 2012; 71: 26–39.
Estacion M, Han C, Choi JS et al: Intra- and interfamily phenotypic diversity in pain syndromes associated with a gain-of-function variant of NaV1.7. Mol Pain 2011; 7: 92.
Persson AK, Liu S, Faber CG, Merkies IS, Black JA, Waxman SG : Neuropathy-associated Nav1.7 variant I228M impairs integrity of dorsal root ganglion neuron axons. Ann Neurol 2013; 73: 140–145.
Patzko A, Shy ME : Update on Charcot-Marie-Tooth disease. Curr Neurol Neurosci Rep 2011; 11: 78–88.
Sathirapongsasuti JF, Lee H, Horst BA et al: Exome sequencing-based copy-number variation and loss of heterozygosity detection: ExomeCNV. Bioinformatics 2011; 27: 2648–2654.
Braathen GJ, Sand JC, Lobato A, Hoyer H, Russell MB : Genetic epidemiology of Charcot-Marie-Tooth in the general population. Eur J Neurol 2011; 18: 39–48.
Acknowledgements
We thank the patients and their families for their participation in the study. Riitta Lehtinen is thanked for technical help. We also acknowledge the target enrichment, sequencing, and variant calling pipeline analysis performed by the Institute for Molecular Medicine Finland (FIMM). We thank the following funding sources for support: Sigrid Jusélius Foundation, the Finnish Neuromuscular Disorders Association, University of Helsinki and the Academy of Finland, Arvid and Greta Olin’s Foundation, and Finska Läkaresällskapet.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Ylikallio, E., Johari, M., Konovalova, S. et al. Targeted next-generation sequencing reveals further genetic heterogeneity in axonal Charcot–Marie–Tooth neuropathy and a mutation in HSPB1. Eur J Hum Genet 22, 522–527 (2014). https://doi.org/10.1038/ejhg.2013.190
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ejhg.2013.190
Keywords
This article is cited by
-
Small heat shock proteins in neurodegenerative diseases
Cell Stress and Chaperones (2020)
-
A PRPH splice-donor variant associates with reduced sural nerve amplitude and risk of peripheral neuropathy
Nature Communications (2019)
-
The Variant p.(Arg183Trp) in SPTLC2 Causes Late-Onset Hereditary Sensory Neuropathy
NeuroMolecular Medicine (2016)
-
Dominant transmission of de novo KIF1A motor domain variant underlying pure spastic paraplegia
European Journal of Human Genetics (2015)
-
Ser135Phe mutation in HSPB1 (HSP27) from Charcot–Marie–Tooth disease type 2F families
Genes & Genomics (2015)