Abstract
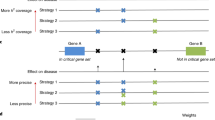
Many associated single-nucleotide polymorphisms (SNPs) have been identified by association studies for numerous diseases. However, the association between a SNP and a disease can result from a causal variant in linkage disequilibrium (LD) with the considered SNP. Assuming that the true causal variant is among the genotyped SNPs, other authors demonstrated that the power to discriminate between it and other SNPs in LD is low. Here, we propose to take advantage of the information provided by family data to improve the inference on the causal variant: we exploit the linkage information provided by affected sib pairs to discriminate the causal variant from the associated SNPs. The family-based approach improves discrimination power requiring up to five times less individuals than its case–control equivalent. However, the main advantage of family design is the possibility to carry out the procedure one step further: the linkage information allows inference on causal variants, which are not genotyped but in LD with tag-SNPs displaying association, which is impossible with case–control design. By means of Bayesian methods, we estimate the LD between the observed SNPs and an unobserved causal variant, as well as the allelic odds ratio at the unobserved causal variant. The proposed procedure is illustrated on a multiple sclerosis (MS) family data set including genotypes of SNPs in IL2RA, confirming the advantage of using a family design to identify causal variants. The results of our method on this data suggest the existence of two distinct causal variants in this gene for the MS.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Risch N, Merikangas K : The future of genetic studies of complex human diseases. Science 1996; 273: 1516–1517.
Manolio T : Genomewide association studies and assessment of the risk of disease. N Engl J Med 2010; 363: 166–176.
Udler MS, Tyrer J, Easton DF : Evaluating the power to discriminate between highly correlated snps in genetic association studies. Genet Epidemiol 2010; 34: 463–468.
Perdry H, Müller-Myhsok B, Clerget-Darpoux F : Using affected sib-pairs to uncover rare disease variants. Hum Hered 2012; 74: 129–141.
Compston A, Confavreux C, Lassmann H et al McAlpine’s Multiple Sclerosis. Churchill Livingstone Elsevier, 2005.
Goris A, Pauwels I, Dubois B : Progress in multiple sclerosis genetics. Curr Genomics 2012; 13: 646.
Babron MC, Perdry H, Handel AE et al: Determination of the real effect of genes identified in GWAS: the example of IL2RA in multiple sclerosis. Am J Hum Genet 2011; 20: 321–325.
Matesanz F, Caro-Maldonado A, Fedetz M et al: IL2RA/CD2 polymorphisms contribute to multiple sclerosis susceptibility. J Neurol 2007; 254: 682–684.
Ramagopalan SV, Anderson C, Sadovnick AD, Ebers GC, Matesanz F et al: Genomewide study of multiple sclerosis. N Engl J Med 2007; 357: 2199–2200.
Hafler DA, Compston A, Sawcer S et al: Risk alleles for multiple sclerosis identified by a genomewide study. N Engl J Med 2007; 357: 851–862.
Ramagopalan SV, Anderson C, Sadovnick AD, Ebers GC : Genomewide study of multiple sclerosis. N Engl J Med 2007; 357: 2199–2200.
Rubio J, Stankovich J, Field J et al: Replication of KIAA0350, IL2RA, RPL5 and CD58 as multiple sclerosis susceptibility genes in australians. Genes Immun 2008; 9: 624–630.
Weber F, Fontaine B, Cournu-Rebeix I et al: IL2RA and IL7RA genes confer susceptibility for multiple sclerosis in two independent european populations. Genes Immun 2008; 9: 259–263.
Armitage P : Tests for linear trends in proportions and frequencies. Biometrics 1955; 11: 375–386.
Sasieni PD : From genotypes to genes: doubling the sample size. Biometrics 1997; 53: 1253–1261.
Slager S, Schaid D : Case-control studies of genetic markers: Power and sample size approximations for armitage’s test for trend. Hum Hered 2001; 52: 149–153.
Robert CP, Casella G : Monte Carlo Statistical Methods.. New York: Springer, 2004, Vol 319.
Poser CM, Paty DW, Scheinberg L et al: New diagnostic criteria for multiple sclerosis: guidelines for research protocols. Ann Neurol 1983; 13: 227–231.
Thomson G : Mapping disease genes: family-based association studies. Am J Hum Genets 1995; 57: 487.
Abecasis GR, Cherny SS, Cookson WO, Cardon LR : Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 2001; 30: 97–101.
Tröster AI : Refining genetic associations in multiple sclerosis. Neurology 2006; 66: 1830–1836.
Alcina A, Fedetz M, Ndagire D et al: IL2RA/CD25 gene polymorphisms: uneven association with multiple sclerosis (ms) and type 1 diabetes (t1d). PLoS One 2009; 4: e4137.
Dendrou CA, Plagnol V, Fung E et al: Cell-specific protein phenotypes for the autoimmune locus IL2RA using a genotype-selectable human bioresource. Nat Genet 2009; 41: 1011–1015.
Maier LM, Anderson DE, Severson CA et al: Soluble IL2RA levels in multiple sclerosis subjects and the effect of soluble IL-2RA on immune responses. J Immunol 2009; 182: 1541–1547.
Maier LM, Lowe CE, Cooper J et al: IL2RA genetic heterogeneity in multiple sclerosis and type 1 diabetes susceptibility and soluble interleukin-2 receptor production. PLoS Genet 2009; 5: e1000322.
Binet J, Auquier A, Dighiero G et al: A new prognostic classification of chronic lymphocytic leukemia derived from a multivariate survival analysis. Cancer 1981; 48: 198–206.
Bourgey M, Perdry H, Clerget-Darpoux F : Modeling the effect of PTPN22 in rheumatoid arthritis. BMC Proc 2007;1:S37.
Clerget-Darpoux F, Babron M, Prum B, Lathrop G, Deschamps I, Hors J : A new method to test genetic models in HLA associated diseases: the MASC method. Ann Hum Genet 1988; 52: 247–258.
Bochdanovits Z, Simón-Sánchez J, Jonker M, Hoogendijk WJ, van der Vaart A, Heutink P : Accurate prediction of a minimal region around a genetic association signal that contains the causal variant. Eur J Hum Genet 2013; 22: 238–242.
Acknowledgements
We wish to thank gratefully Françoise Clerget-Darpoux for many fruitful discussions. We thank all the patients who generously participated in and the physicians constituting the REFGENSEP (Réseau français de la génétique de la sclérose en plaques). We also thank all the authors of Babron et al7 for their help on example data set, in particular Marie-Claude Babron. We also thank Rémi Kazma for a careful reading of the manuscript. Hervé Perdry was partly funded by a grant from ARSEP fondation (Fondation pour l’Aide à la Recherche sur la Sclérose en Plaques).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Dandine-Roulland, C., Perdry, H. Where is the causal variant? On the advantage of the family design over the case–control design in genetic association studies. Eur J Hum Genet 23, 1357–1363 (2015). https://doi.org/10.1038/ejhg.2014.284
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ejhg.2014.284
This article is cited by
-
The genetics of venous thromboembolism: a systematic review of thrombophilia families
Journal of Thrombosis and Thrombolysis (2021)