Abstract
Purpose: In the United States, approximately 1/3,700 babies is born with cystic fibrosis each year. The >1,300 documented sequence variants pose a challenge for detection of cystic fibrosis through genetic screening. To investigate whether comprehensive characterization of the cystic fibrosis gene is feasible using dried newborn blood specimens, we modified the whole blood Ambry Test™: CF and determined its sensitivity by testing DNA from individuals with cystic fibrosis who still had unknown mutations after commercial mutation panel testing.
Methods: DNA from 42 archived newborn dried blood specimens of affected Hispanic, African-American and Caucasian individuals in California was analyzed by temporal temperature gradient electrophoresis screening and targeted sequencing, and by gross deletion analysis.
Results: Excluding two specimens that could not be analyzed due to poor DNA quality, we report a 100% sensitivity and clinical detection rate in the remaining 40 patients. Eighty-three mutations representing 40 different variants were detected, including 8 novel mutations.
Conclusions: This study demonstrates the feasibility of temporal temperature gradient electrophoresis-based full sequence analysis and targeted sequencing from DNA in newborn blood specimens. The Ambry Test™: CF, as an additional step in cystic fibrosis newborn screening models, can be used to dramatically reduce the number of cystic fibrosis carrier sweat test referrals.
Similar content being viewed by others
Main
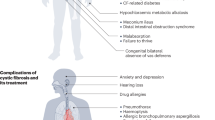
Cystic fibrosis (CF) is a common and generally severe autosomal recessive disorder caused by mutations in the CF transmembrane conductance regulator gene (CFTR, OMIM 602421). In the United States, 1/3,700 babies is born with CF, and there are approximately 10 million CF carriers and 30,000 diagnosed CF patients.1 CF causes abnormal sweat electrolytes, pulmonary disease, male infertility and pancreatic insufficiency. As of 2003, the median age of survival was 35.1 years.1 Since the cloning of the CFTR gene located on chromosome 7q31,2,3 more than 1,300 sequence variants have been identified and entered into the CF mutation database.4 The most common mutation in CF is delta F508, present in an estimated 20–80% of CF patients depending on ethnicity and demographics.2 Worldwide, only four other mutations have relative frequencies of 1.2–2.4%. The majority of mutations are rare, with most frequencies under 0.1%.
Given the diversity of known and novel CFTR mutations, prenatal and newborn screening success using commercially available panels is limited. Detection rates vary by ethnic background, with the highest detection rates being in non-Hispanic whites.5–7 This limitation is especially apparent in subpopulations, e.g., in Hispanics, among whom mutations are still not well characterized enough despite efforts to do so.8–13 Standard DNA sequencing thus far has been too labor-intensive and costly to be used as an initial screening tool. Another restriction of comprehensive sequencing for newborn screening has been the need for fresh whole blood, which rules out its use in screening programs that use newborn blood collected on filter paper.
This study aimed to investigate whether comprehensive characterization of the CFTR gene was feasible using dried newborn blood specimens. We modified the whole blood Ambry Test™: CF and tested the sensitivity and clinical detection rate using DNA from 42 archived newborn blood specimens of affected children in California with one or more unknown mutations.
MATERIALS AND METHODS
Study population
The 42 dried blood specimens analyzed were part of a larger study attempting to obtain genotype information on persons in California with an emphasis on those of Hispanic or African- American origins.14 The study population derives from the California CF Patient Registry, assembled from clinic (1995–2000), vital statistics-death (1982–2000), and Medi-Cal (1995–1999) records, and includes over 3,000 persons with CF. A description of the registry has been published previously.11 When genotype data were not complete from clinical records, dried blood specimens were sought from an archive of newborn blood maintained as part of the California Newborn Screening Program. The archive included California live births between 1982 and 2000. Dried blood specimens were first analyzed by the California Genetic Disease Laboratory using the ABI-31 mutation panel, and if one or more mutations were still not identified, the specimens were then sent to Genzyme Genetics for analysis using an 87-mutation panel. The subjects in this paper include 38 Hispanics, two African-Americans, and two Caucasians with CF, in which dried newborn blood specimens were available, and whose genotypes were incomplete after analysis with the Genzyme-87 panel. Ten of these subjects (cases 1, 2, 4–9, 12, and 13) were recruited by their CF Centers to provide fresh blood for TTGE and focused DNA sequencing testing by the Institute for Molecular and Human Genetics, Georgetown University Medical Center.11 IRB approval was obtained from the California Health and Human Services Agency Committee for the Protection of Human Subjects and from individual CF Center hospitals.
Subjects were not notified of the Ambry Genetics test results from the newborn specimens. The specimens of these subjects were identified through record linkage of historic data sources, and, except for the ten subjects who gave whole blood, contact information and vital status were unknown. Results from the whole blood genetic analyses performed by Georgetown University were previously reported to the ten subjects through their CF care provider.
Genetic analysis
The dried newborn blood specimens were collected on Schleicher and Schuell 903 Specimen Collection Paper and then banked for varying lengths of time at −20°C after testing was completed for a number of state-mandated screening disorders. Coded and blinded specimens were sent to Ambry Genetics. DNA was extracted using the recommended protocol by Schleicher and Schuell based on a procedure used at the National Institute for Standards and Technology.15 Genomic DNA was then used in the Ambry Test™: CF.
The Ambry Test™: CF is a full-mutation scan of the CFTR gene by TTGE followed by dye terminator DNA sequencing of suspect regions. All CFTR exons as well as relevant intronic regions were amplified using polymerase chain reaction (PCR) and proprietary primers. Standard PCR amplification was performed using HotStarTaq Master Mix (Qiagen) with 10–30 ng input gDNA per reaction. Typical PCR conditions were 1 cycle: 95°C for 15 minutes, 39 cycles: 94°C for 30 seconds, 54°C for 30 seconds, 72°C for 30 seconds, 1 cycle: 72°C for 10 minutes. Annealing temperatures differed depending on primer pair. Prior to Ambry gel analysis, the PCR products were denatured and slowly cooled to allow for maximal heteroduplex formation. For a subset of CFTR regions, DNA was mixed with known wild-type DNA to facilitate detection of homozygous mutations. PCR products were processed for TTGE on DCode gels (BioRad) in adherence with the Ambry Test™ technology. Polyacrylamide gels were analyzed for the presence of mutations following staining in ethidium bromide (EtBr) and image capture under UV using the Gel Doc 1000 system (BioRad). Gel analysis was performed by two technicians and fragments were scored against known controls.
Regions indicating the presence of a mutation by TTGE were processed for sequencing. Apparently affected exons were amplified with a unique primer set using Taq PCR Master Mix (Qiagen, Valencia, CA). Typical PCR conditions were 1 cycle: 95°C for 5 minutes, 35 cycles: 94°C for 30 seconds, 54°C for 30 seconds, 72°C for 30 seconds, 1 cycle: 72°C for 10 minutes. Annealing temperatures differed depending on the primer pair. PCR products were analyzed via agarose gel electrophoresis, followed by treatment with ExoSAP-It (USB) according to manufacturer recommendations. Standard dye terminator cycle sequencing DTCS (Beckman Coulter) was conducted followed by loading onto a CEQ8000 sequencer. Exons were always sequenced in both sense and antisense directions. All reported mutations follow the numbering of Genbank entry NM_000492, the CF database4 and recommendations by den Dunnen and Antonarakis for novel variants.16
Two specimens in which only one mutation was detected with TTGE were reflex-tested with gross deletion analysis utilizing multiplex ligation-dependent probe amplification (MLPA) (MRC –Holland).17 DNA was analyzed with the CF MLPA kit. Briefly, genomic DNA was hybridized overnight with a CF and control probe mixture. Each exon's probe set consists of two probes of immediately adjoining sequence that hybridize to the same strand. The probes each contain a flanking universal primer region at their respective 5′ ends. Ligation of the probes was followed by amplification with the universal primers and separation and sizing of products by automated gel capillary electrophoresis on a CEQ8000. Data were obtained with regression-enhanced MLPA analysis based on a method by Mavrogiannis and Cockburn.18 In summary, calculations of amplicon-size corrected peak heights were performed for each experimental and control subject and dosage quotients were calculated. The dosage quotients between experimentals and controls allow for scoring of each exon's allelic state, indicating whether or not exons are present, deleted or duplicated. In addition to two reflex-tested specimens, 22 specimens in which two mutations had been identified were also processed for gross deletion analysis (all were negative).
Positive gross deletions were also verified with a second method using quantitative real-time PCR. CF exon 2-, 3-, 17A-, and 17B-specific probes were used to analyze allelic state on a Chromo4 system (BioRad). Breakpoints were also determined for the CFTRdele2,3 according to the method described by Dork et al.19
RESULTS
Comprehensive CFTR TTGE screening and sequencing was successful for 39 of 42 specimens. One sample was processed partially; however, two mutations were still identified. Two of the specimens were not included in the analysis because DNA amplification was poor. The cause of the DNA amplification problems could not be determined, and repeat extraction from dried blood in another spot on the same newborn collection paper did not improve results significantly.
Genotype results from two panels (ABI-31 and Genzyme-87), the Ambry Test™: CF with gross deletion analysis, as well as the TTGE results (using whole blood from ten patients) are shown in Table 1. The initial ABI-31 and Genzyme panels had correctly identified 32 mutations in the 40 patients with sufficiently amplified DNA. Using the Ambry Test™: CF, a total of 81 mutations was initially identified: 35 specimens had two mutations detected, 3 had three mutations detected, and 2 had only one mutation identified. Subsequent analysis with the gross deletion assay revealed the presence of gross deletions in the two specimens that only had one mutation detected, bringing the detection rate (clinical sensitivity) to 100% for these 40 specimens and the total number of mutations to 83. Nineteen of 21 mutations identified using the Ambry Test™: CF in ten patients were also confirmed by the Georgetown University laboratory using fresh blood (Table 1).
Altogether, these 83 mutations represent 40 different variations. Apart from delta F508, nine other mutations were present more than once in this data set, with several mutations occurring 4–5 times (1288insTA, 2055del9insA, 406-1G>A, G542X and H199Y). Using the ACMG/ACOG-23 panel in this study population, only 30 mutations (36.1%) representing five different mutations (12.5%) would have been detected. Using the current expanded Genzyme-97 panel, 51 mutations (61.4%) representing 15 different mutations (37.5%) would have been detected.
Moreover, novel mutations and gross deletions contributed significantly to the genotypes. We identified eight novel CFTR mutations in seven persons in the study population (Table 2): two contained stop codons (E257X, Y914X), two were small deletions (3940delG, 2289del10ins5), two were amino acid substitutions (M150K, T465N), one was a gross deletion of exons 17A and 17B, and one was a base pair substitution (3849 + 72G>A).
Clinical data for the seven patients with novel mutations are shown in Table 2. All of the patients with novel mutations had elevated immunoreactive trypsinogen (IRT) and/or sweat tests and are clinically diagnosed with CF; however, the patient with the amino acid substitution T465N (and a known deleterious mutation at 711 + 1G>T) died in childhood and no additional clinical information is available.
DISCUSSION
Utilizing standard procedures of DNA extraction from dried newborn blood specimens and PCR amplification, combined with TTGE screening, DNA sequencing procedures, and gross deletion analysis, we were able to reliably detect two or more mutations in 100% of a group of 40 CF patients with diverse backgrounds. DNA amplification in two other specimens was problematic and it was impossible to request a repeat specimen due to the retrospective study design, unlike in clinical or newborn screening settings.
The CF subjects studied here represent those whose mutations were still not identified after testing with commercial panels, and as such, were selected to have greater mutational heterogeneity than a random sample of CF cases. Our methods were able to identify eight novel mutations in seven individuals. While the interpretation of novel mutations often poses a challenge as phenotypic impact is uncertain, five of the novel mutations are likely to be deleterious: two contained stop codons (E257X, Y914X), two were small deletions (3940delG, 2289del10ins5), and one was a gross deletion of exons 17A and 17B. Although Niel et al.20 recently described a CFTRdele17A,17B, we classify our dele 17A, B as a novel mutation since precise breakpoints could not be compared due to lack of DNA. Various breakpoints have been shown to occur involving introns 16 and 18.21–25 The 2289del10ins5 mutation was novel to Ambry at the time of analysis, but was subsequently published.11
The novel variation identified at 3849 + 72G>A may not have phenotypic significance if disease is manifest due to the other mutations present, 3120 + 1G>A and Q989X, occurring in trans. But, the same intron has a known deleterious mutation at 3849 + 40 bp, and deep intronic at 3849 + 10 kb, so conceivably this intronic variant could be a contributing factor. Unfortunately, in this study we did not have the ability to conduct family studies to determine the phase of the mutations, which is always recommended clinically. The two other novel variants were amino acid substitutions (M150K, T465N). The patient with M150K also had the gross deletion of exons 17A and 17B. The other patient with the amino acid substitution T465N (and a known deleterious mutation at 711 + 1G>T) died in childhood, and no additional clinical information could be obtained.
One other gross deletion warrants special mention. CFTRdele2,3 (21kb) was detected in study subject 8 which also carried a delta F508. This gross deletion occurred at the exact documented breakpoints.19 Interestingly, this gross deletion is present at high frequencies (1.1–6.4% of CF chromosomes) in many Eastern and Western Slavic populations, suggesting its inclusion on first tier mutation panels.
The ability of the procedures described in this study to be used on dried newborn blood specimens along with the relatively quick turn-around time (procedures can be performed in 2–4 weeks), make their usefulness of particular importance to newborn screening programs. Methods for CF newborn screening differ widely from state to state.26 All testing starts with a phenotypic test that measures IRT in dried blood specimens. Specimens with elevated IRT measurements (typically top 4% or less) undergo further tests. In states with two specimen models, another IRT is typically measured in the second specimen. In states with single specimen models, original specimens are tested for the presence of the most common mutation, delta F508, and/or other deleterious mutations using mutation panels. In all states, newborns with a positive screening test are followed-up with a diagnostic sweat chloride assessment and genetic counseling. Typically, for each CF case identified 10 or more screen-positive newborns are found not to have CF.26
The highly sensitive laboratory procedures adapted for dried blood specimens used in this study may provide an alternative to sweat testing all babies with one identified mutation or with a second high IRT level. The Ambry Test™: CF, or comparable methods of comprehensive DNA sequencing,27 could be added as a step in the screening model prior to sweat testing so that only those newborns with two or more mutations identified would then be referred for sweat testing. The benefits of this new approach are several. These include on the immediate analytical side: fewer false-positive results, sweat tests and repeat sweat tests, fewer referrals and an expedited diagnosis allowing for earlier care. It is documented that babies diagnosed through newborn screening have fewer complications carrying into adolescence than those later diagnosed through symptomatic disease.28 For parents, one would expect a reduced burden in terms of taking time off from work for repeat doctors visits, since all initial analysis (IRT and DNA) can be routed from dried blood collected at birth. In addition, an earlier definite diagnosis would reduce the overall emotional strain on the parents from the knowledge that the newborn screened positive before the results were complete.29 From the care provider side, diagnostic centers would incur less burden of sweat testing false-positive babies and counseling their families, allowing for resources to be better directed to those in need. Genetic counseling could target affected families as well, supplying more certainty in counseling those with borderline sweat test results, and allowing for greater ease in referring family members for genetic testing. Also, data obtained from using such a testing paradigm will unveil more genetic information and add value for future CF research and therapies. Comprehensive genetic information will also give the state newborn screening program the ability to monitor and adapt mutation panels to changing mutation frequencies in the population.
While this approach will lower the number of false-positives dramatically, there will be some individuals identified, with one clear mutation and one novel or questionable variant or with two mutations that occur in trans, who will be found to be false-positives upon sweat testing. Family studies, which could not be performed in this study, typically are very helpful in clarifying these questions. Finally, due to the many variables involved in establishing a newborn screening protocol, detailed financial implications of this approach are beyond the scope of this paper; however, we anticipate the additional cost per newborn specimen will be offset by the benefits of the new screening model and will ultimately provide a cost savings and health benefits.
References
CFF patient registry annual data report 2004. Available at: http://www.cff.org/publications/#Patient_Registry. Accessed on June 20, 2006.
Kerem B, Rommens JM, Buchanan JA, Markiewicz D, et al. Identification of the cystic fibrosis gene: genetic analysis. Science 1989; 245: 1073–1080.
Riordan JR, Rommens JM, Kerem B, Alon N, et al. Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science 1989; 245: 1066–1073. Erratum in: Science 1989;245:1437.
Cystic Fibrosis Mutation Database. Available at: www.genet.sickkids.on.ca/cftr/. Accessed June 20, 2006.
Bobadilla JL, Macek M Jr, Fine JP, Farrell PM . Cystic fibrosis: a worldwide analysis of CFTR mutations—correlation with incidence data and application to screening. Hum Mutat 2002; 19: 575–606.
Heim RA, Sugarman EA, Allitto BA . Improved detection of cystic fibrosis mutations in the heterogeneous U.S. population using an expanded, pan-ethnic mutation panel. Genet Med 2001; 3: 168–176.
McCormick J, Green MW, Mehta G, Culross F, et al. Demographics of the UK cystic fibrosis population: implications for neonatal screening. Eur J Hum Genet 2002; 10: 583–590.
Orozco L, Zielenski J, Markiewicz D, Villarreal T, et al. Two novel frameshift deletions (1924del7, 2055del9–>A) in the CFTR gene in Mexican cystic fibrosis patients. Hum Mutat 1997; 10: 239–240.
Orozco L, Velazquez R, Zielenski J, Tsui LC, et al. Spectrum of CFTR mutations in Mexican cystic fibrosis patients: identification of five novel mutations (W1098C, 846delT, P750L, 4160insGGGG and 297-1G–>A). Hum Genet 2000; 106: 360–365.
Wong LJ, Wang J, Zhang YH, et al. Improved detection of CFTR mutations in southern California Hispanic CF patients. Hum Mutat 2002; 19: 79.
Alper OM, Wong LJ, Young S, Pearl M, et al. Identification of novel and rare mutations in California Hispanic and African-American cystic fibrosis patients. Hum Mutat 2004; 24: 353.
Alper OM, Wong LJ, Young S, Pearl M, et al. Erratum: Identification of novel and rare mutations in California Hispanic and African-American cystic fibrosis patients. Hum Mutat 2005; 25: 223.
Schrijver I, Ramalingam S, Sankaran R, Swanson S, et al. Diagnostic testing by CFTR gene mutation analysis in a large group of Hispanics: novel mutations and assessment of a population-specific mutation spectrum. J Mol Diagn 2005; 7: 289–299.
California Genetics Disease Branch and the Public Health Institute, 2004. Available at: http://www.dhs.ca.gov/pcfh/gdb/html/PDE/CFStudy.htm. Accessed on June 20, 2006.
Walsh PS, Metzger DA, Higuchi R . Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 1991; 10: 506–513.
den Dunnen JT, Antonarakis SE . Mutation nomenclature extensions and suggestions to describe complex mutations: a discussion. Hum Mutat 2000; 15: 7–12. Erratum in: Hum Mutat 2002;20:403.
Schouten JP, McElgunn CJ, Waaijer R, Zwijnenburg D, et al. Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res 2002; 30: E57.
Mavrogiannis and Cockburn REX-MLPA User guide, 2004. Available at: http://leedsdna.info/science/dosage/REX-MLPA/REX-MLPA_analysis_User_Guide.pdf. Accessed on June 20, 2006.
Dork T, Macek M, Mekus F, Tummler B, et al. Characterization of a novel 21-kb deletion, CFTRdele2,3(21 kb), in the CFTR gene: a cystic fibrosis mutation of Slavic origin common in Central and East Europe. Hum Genet 2000; 106: 259–268.
Niel F, Martin J, Dastot-Le Moal F, Costes B, et al. Rapid detection of CFTR gene rearrangements impacts on genetic counselling in cystic fibrosis. J Med Genet 2004; 41: E118.
Lerer I, Laufer-Cahana A, Rivlin JR, Augarten A, et al. A large deletion mutation in the CFTR gene (3120+1Kbdel8.6Kb): a founder mutation in the Palestinian Arabs. Mutation in brief no. 231. Online. Hum Mutat 1999; 13: 337.
Feuillet-Fieux MN, Ferrec M, Gigarel N, Thuillier L, et al. Novel CFTR mutations in black cystic fibrosis patients. Clin Genet 2004; 65: 284–287.
Kilinc MO, Ninis VN, Dagli E, Demirkol M, et al. Highest heterogeneity for cystic fibrosis: 36 mutations account for 75% of all CF chromosomes in Turkish patients. Am J Med Genet 2002; 113: 250–257.
Shrimpton AE, Borowitz D, Swender P . Cystic fibrosis mutation frequencies in upstate New York. Hum Mutat 1997; 10: 436–442.
Chevalier-Porst F, Souche G, Bozon D . Identification and characterization of three large deletions and a deletion/polymorphism in the CFTR gene. Hum Mutat 2005 May; 25: 504.
Grosse SD, Boyle CA, Botkin JR, Comeau AM, et al. CDC. Newborn screening for cystic fibrosis: Evaluation of benefits and risks and recommendations for state newborn screening programs. MMWR Recomm Rep 2004; 53: 1–36.
Chou LZ, Lyon E, Wittwer CT . A comparison of high-resolution melting analysis with denaturing high-performance liquid chromatography for mutation scanning. Am J Clin Path 2005; 124: 330–338.
Accurso FJ, Sontag MK, Wagener JS . Complications associated with symptomatic diagnosis in infants with cystic fibrosis. J Pediatr 2005; 147: S37–S41.
Kharrazi M, Kharrazi LD . Delayed diagnosis of cystic fibrosis and the family perspective. J Pediatr 2005; 147: S21–S25.
Acknowledgements
The authors thank the patients and clinicians for their participation in this study. Judy Bolstad provided assistance with sample collection. Mutation analysis at the Genetic Disease Laboratory was conducted by Dr. Helen Chow under the direction of Drs. John Sherwin and John Eastman. We also wish to thank the staff at Ambry Genetics for their expert technical assistance.
Supported by: CF Foundation Therapeutics, Inc. (M.K. grant # KHARRA01A0); Cystic Fibrosis Research, Inc. (CFRI) (M.K. grant # 00-003) to the Public Health Institute.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Kammesheidt, A., Kharrazi, M., Graham, S. et al. Comprehensive genetic analysis of the cystic fibrosis transmembrane conductance regulator from dried blood specimens – Implications for newborn screening. Genet Med 8, 557–562 (2006). https://doi.org/10.1097/01.gim.0000237793.19868.97
Received:
Accepted:
Issue date:
DOI: https://doi.org/10.1097/01.gim.0000237793.19868.97
Keywords
This article is cited by
-
A comprehensive overview of the cystic fibrosis on the island of São Miguel (Azores, Portugal)
BMC Pediatrics (2020)
-
Whole genome microarray analysis, from neonatal blood cards
BMC Genetics (2009)