Abstract
Purpose:
We examined hospital use of the 21-gene breast cancer test in the United States. We report state-level differences in utilization and propose a model for predicting implementation of guideline-recommended genomic testing.
Methods:
Genomic Health provided test orders for calendar year 2011.We summarized utilization at the hospital and state levels. Using logistic regression, we analyzed the association between the likelihood to order the test and the hospital’s institutional and regional characteristics.
Results:
In 2011, 45% of 4,712 acute-care hospitals ordered the test, which suggests that 25% of newly diagnosed invasive female breast cancer cases were tested. Significant predictors of testing included participation in National Cancer Institute (NCI) clinical research cooperative groups (odds ratio (OR) 3.73; 95% confidence interval, 2.96–4.70), advanced imaging (OR, 2.19; CI, 1.78–2.68), high-complexity laboratory (OR, 2.15; CI, 1.24–3.70), affiliation with a medical school (OR, 1.57; CI, 1.31–1.88), and reconstructive surgery (OR, 1.23; CI, 1.01–1.50). Significant regional predictors included metropolitan county (OR, 3.77; CI, 2.83–5.03), above-mean income (OR, 1.37; CI, 1.11–1.69), and education (OR, 1.26; CI, 1.03–1.54). Negative predictors included designation as a critical-access hospital (OR, 0.10; CI, 0.07–0.14) and distance from an NCI cancer center (OR, 0.998; CI, 0.997–0.999), with a 15% decrease in likelihood for every 100 miles.
Conclusion:
Despite considerable market penetration of the test, there are significant regional and site-of-care differences in implementation, particularly in rural states.
Genet Med 18 10, 982–990.
Similar content being viewed by others
Introduction
Breast cancer is a heterogeneous disease with a wide range of health outcomes not fully predicted by histological features of the breast tumors. Therefore, all modern approaches to breast cancer treatment rely on measurement of the expression of cancer-related genes in addition to histological evaluation of the tumor.1 Several molecular tests are recommended by clinical practice guidelines to inform diagnosis and treatment decisions. A growing number of laboratories are analyzing expression of cancer-related genes to estimate the risk of breast cancer recurrence and predict response to treatments. However, very few studies have analyzed population-level implementation of genomic tests. Such research is challenging to conduct due to limited availability of de-identified population-level data sets that accurately identify use of tests and due to the lack of gene or test-specific identifiers within medical claims data. These studies are crucial to allow further evaluation of clinical utility, equity in access, prevalence of cancer-related mutations, and health outcomes of genomic testing.
The development of molecular tumor profiling has made it possible to distinguish several clinically meaningful subtypes of breast cancer based on the expression of estrogen receptors (ER) and progesterone receptors (PR), collectively referred to as hormone receptors (HR), and human epidermal growth factor receptor 2 (HER2/neu). These subtypes are associated with different prognoses and are targets for different therapies.2 For ER+/HER2– patients in particular, adjuvant endocrine therapy is currently regarded as standard of care.3,4 The addition of cytotoxic chemotherapy to endocrine therapy results in a statistically significant, yet small, survival advantage.5 For a significant number of patients, additional chemotherapy offers little or no benefit, as demonstrated by several clinical trials.6 Until recently, the majority of early-stage, ER-positive (ER+) breast cancer patients were considered for adjuvant chemotherapy. Physicians had a limited ability to evaluate individual risks.6 Development of clinical tests to further stratify patients into risk groups to predict the benefit of chemotherapy is important.
Currently, several multigene assays are available that evaluate the likelihood of chemotherapy benefit and the probability of recurrence in cancer patients.7 Among these, the 21-gene recurrence score assay (Oncotype DX Breast Cancer Assay) developed and conducted by Genomic Health (Redwood City, CA) is the most widely used for management of ER+, lymph node–negative (LN–) early breast cancer.8,9 The predictive power of the 21-gene test has been validated in several large clinical trials, including a multicenter National Surgical Adjuvant Breast and Bowel Project clinical trial (B-14)10 and a population-based case-control study at Northern California Kaiser Permanente.11 Results from a prospective validation study (the Trial Assigning Individualized Options for Treatment, or TAILORx), launched in 2006, demonstrated remarkably low rates of recurrence with endocrine therapy alone for the low-risk subgroup (recurrence score of 0–10; representing 16% of the enrolled patients). At 5 years, the risk of the recurrence of breast cancer at a distant site was less than 1% and the risk of any recurrence was less than 2% (ref. 12).
The 21-gene test is the only multigene expression test for breast cancer included in the American Society of Clinical Oncology and National Comprehensive Cancer Network guidelines for early-stage ER+ tumors. The test is not recommended for patients with HER2+ or ER–/PR–/HER2– (triple-negative breast cancer) subtypes. Numerous publications report that the test has had a significant impact on treatment decisions, resulting in changing chemotherapy recommendations in 20.6 to 74.0% of cases, depending on the patient population studied and the guideline used for comparison.13,14 Research demonstrating cost-effectiveness of the test15,16 has been conducted in several countries, and it has achieved notable market penetration compared with other multigene assays in breast cancer.14 A regional study involving 7,375 patients at 17 cancer centers also showed a progressive increase in use of the 21-gene test from 2006 to 2008.17 In 2006, a California Medicare Administrative Contractor published a local coverage determination for reimbursement of the test for women with early-stage ER+, node-negative breast cancers.18 A recent retrospective analysis of the Surveillance, Epidemiology, and End Results (SEER) registry of Medicare-linked data for 70,802 beneficiaries revealed an increase in utilization of the test from 1.1% in 2005 to 10.1% in 2009 among Medicare beneficiaries.19 Other utilization studies were limited to small groups of several hundred patients.8,20,21
Although a growing number of publications have reported implementation of the 21-gene test, these studies are limited to specific patient groups. A comprehensive population-level analysis is needed to evaluate the current level of implementation. In this study, we measured nationwide utilization of the 21-gene test performed in calendar year 2011. Based on previously published research analyzing implementation of genomic medicine,8,20,21,22 we hypothesized that, despite rapid adoption of the 21-gene test,19 there would be regional and site-of-care variations, as well as use that departs from clinical practice guidelines. To test this hypothesis, we compared total test orders per county to the number of newly diagnosed breast cancer cases. We then analyzed the association between test orders and the hospital’s institutional and regional characteristics to identify factors that predict utilization. We evaluated concordance with guidelines by analyzing test utilization within the SEER registry data.
Materials and Methods
This was a retrospective cross-sectional study. We merged one proprietary data set with five public data sets. Genomic Health, the company that developed and conducts the 21-gene test, provided the name, city, state, zip code, and number of tests ordered by each hospital, laboratory, or outpatient clinic in the United States in calendar year 2011. Public data sets included the following:
-
1. Centers for Medicare and Medicaid Services (CMS) Provider of Services file, which contained hospital characteristics, including academic medical affiliation and services offered
-
2. CMS Clinical Laboratory Improvement Amendments (CLIA) specialty file, which identified laboratory accreditation to conduct cytogenetic testing
-
3. National Cancer Institute (NCI) Provider of Services file, which identified hospital participation in NCI clinical research cooperative groups
-
4. State-level and county-level counts of 2011 invasive female breast cancers provided by the Centers for Disease Control and Prevention’s (CDC) National Program of Cancer Registries (NPCR) or the NCI’s SEER program23
-
5. Health Resources and Services Administration Area Resource file, which provided county characteristics
We aggregated and summarized test orders at the hospital level, recording each hospital’s Medicare provider number from the CMS Provider of Services file. When possible, orders from independent pathology laboratories were matched to hospitals serviced. This matching was accomplished by using the laboratory CLIA number listed in the Provider of Services file and through Web-based searches of hospital pathology laboratories. It was not possible to perform such matching for regional or national reference laboratories. The aggregated data set was merged with public data sets using hospital number, zip code, and FIPS code as unique identifiers for each US hospital and county.
To calculate the state-level utilization, we obtained the number of newly diagnosed invasive female breast cancer cases per state from CDC NPCR and NCI SEER.23 Utilization was then calculated as the number of tests ordered divided by the number of new cases of invasive female breast cancer for each state.
The unit of observation for multivariate analysis was the nonfederal, acute-care hospital, as defined by CMS. The outcome variable was whether a hospital ordered the 21-gene test; it was treated dichotomously and coded as 1 if it ordered one or more assays.
Independent variables that describe the hospital included institutional and regional characteristics. Institutional characteristics were affiliation with a medical school, designation as a critical-access hospital, distance to an NCI cancer center, member of NCI clinical research cooperative group, and types of services the hospital offers such as reconstructive surgery, positron emission tomography scans, and high-complexity laboratory services. (A high-complexity laboratory is defined as a clinical laboratory that conducts molecular testing and meets the federal regulatory standards as outlined in the CLIA of 1988.24) Regional (county) characteristics were education and income level, metropolitan status, percentage of the population that is black, and number of annual breast cancer cases. These variables were chosen because the proportion of eligible patients is influenced by sociodemographics. Education, race, and income impact the stage of breast diagnosis and tumor subtype.25,26
Most independent variables were dichotomous and coded as 1 if the hospital or region had the characteristic. Number of breast cancer cases per county, percentage of the county population that was black, and hospital distance from an NCI Cancer Center were treated as continuous variables. Hospital distance from an NCI Cancer Center was measured in miles and calculated using the longitude and latitude of each institution. To accurately interpret the odds ratio (OR) of distance, we obtained the logit coefficient and multiplied it by increments of 100 miles.
We conducted univariate and bivariate analyses, including t-tests for continuous variables and χ2 for categorical variables. Statistically significant explanatory variables (P values <0.05) were included in the logistic regression model. We performed multivariate logistic regression to identify characteristics associated with the ordering of the 21-gene test by a hospital. The statistics program STATA version 12.0 (STATA, College Station, TX) was used in the analyses.
To estimate the proportion of tests performed per guideline recommendations, we used data from the NCI SEER program. SEER has required the collection of the 21-gene test since 2010. We analyzed patients diagnosed with breast cancer in 2011 who had the 21-gene test ordered. Cases were categorized as meeting or not meeting the guideline criteria for testing.
Results
According to the CDC’s United States Cancer Statistics, in 2011 there were 221,663 new cases of invasive female breast cancer in the 50 states and the District of Columbia.23 In 2011, 2,461 institutions ordered 59,542 21-gene tests ( Table 1 ), representing 26.86% of all newly diagnosed invasive breast cancer cases. This percentage was slightly lower (24.81%) when we restricted analysis to only those tests that we could directly link to an acute-care hospital ( Table 2 ). Nonfederal acute-care hospitals accounted for 84.48% of tests ordered. NCI cancer centers and regional or national pathology laboratories, which constituted 2.25 and 7.43% of the institutions, ordered 8.03 and 6.04% of the tests, respectively. Various federal hospitals, hospitals in Puerto Rico, outpatient oncology clinics, and physicians’ offices accounted for the remaining tests.
Of 4,712 acute-care hospitals in the database, 45.06% ordered the test. The number of tests ordered by each hospital ranged from 1 to 697 tests (mean = 11.67, SD = 27.43). There were 144 hospitals that ordered only one test. Supplementary Table S1 online provides the hospital identifier and the number of tests ordered.
State-wise analysis
We limited subsequent analysis to hospitals and NCI cancer centers. Table 2 illustrates the state-level summary of the number of hospitals, NCI cancer centers, cytogenetic hospital laboratories, and the 21-gene test assays ordered in each state. The last column shows the percentage of newly diagnosed breast cancer cases for which the test was ordered. The map in Supplementary Figure S1 online provides a graphic illustration of the number of tests ordered within each state and the proportion of that number to the number of invasive female breast cancer cases per state.
These data demonstrate that hospitals in every state in the country ordered the test. If we presume that all patients tested had newly diagnosed invasive cases, then we can estimate the percentage of patients tested per state using the invasive female breast cancer incidence data reported in CDC NPCR as the denominator. The percentage of cases tested within a state ranged from 11.96 to 73.00% (mean = 24.53%, SD = 8.13). The highest utilization of the test (73%) was in the District of Columbia; however, this was an outlier. One hospital, a community affiliate of NCI cancer center Johns Hopkins, ordered nearly half the tests in the District of Columbia. These orders probably reflected patients migrating across borders to obtain care at this hospital. Pennsylvania, which has five NCI-designated cancer centers and 15 hospitals with high-complexity laboratories, had the second highest utilization (30.83%), followed closely by New York at 30.75%. Mississippi, which has no NCI-designated cancer center and only one hospital with a high-complexity laboratory, had the lowest utilization (11.96%).
County analysis
There are 2,496 counties in the United States with acute-care hospitals. Hospitals in 1,001 (40.00%) of these counties ordered the 21-gene test. Ninety-six counties (3.84%) have hospitals accredited to conduct high-complexity testing; 50 (2.00%) have NCI cancer centers. Most hospitals that ordered the 21-gene test were located within metropolitan counties (2,048, 96.47%); 1,652 (67.81%) were located in counties with above average income levels; 1,589 (74.84%) were located in counties with above average education levels above the mean. Analyzed independently, there was a positive association between the percentage of blacks living within a country and the likelihood that a hospital ordered the 21-gene test. However, that is probably confounded by the hospital’s location in a metropolitan county.
Bivariate analysis of hospital characteristics
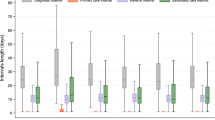
Based on a previous study that analyzed implementation of a lung cancer molecular test,22 it was expected that the hospital characteristics, including affiliation with an academic medical center, participation in an NCI cooperative group, availability of positron emission tomography scan services, and having a CLIA-accredited high-complexity laboratory, would be positively associated with likelihood to order genomic tests. For this analysis, we added availability of reconstructive surgery to the model because hospitals offering this service are likely to perform breast cancer mastectomies and reconstruction. Figure 1a illustrates that the institutional characteristic most associated with ordering of the 21-gene test was the hospital having a CLIA-accredited high-complexity laboratory; 88% of hospitals that have this service ordered the test. Participation in NCI-funded cooperative groups, medical school affiliation, and reconstructive surgery were also associated with ordering the test. Only 48 (4.09%) hospitals designated as critical-access hospitals ordered the 21-gene assay.
It was expected that the highest percentage of hospitals ordering the assay would be within NCI cooperative groups conducting breast cancer clinical trials. Figure 1b shows that Radiation Therapy Oncology Group and Southwest Oncology Group had the highest percentage of members (90%) ordering the assay. All other NCI cooperative groups had high levels of participation in testing (≥79%), including 88% of the hospitals within the National Surgical Adjuvant Breast and Bowel Project, which conducted the first clinical trial of the 21-gene assay.10
Prediction model
Multivariate logistic regression analysis identified a number of independent variables that predicted ordering the 21-gene test. Table 3 summarizes the OR and 95% confidence intervals (CI) for each variable. Significant institutional predictors included participation in an NCI clinical research cooperative group (OR, 3.73; 95% CI, 2.96–4.70), advanced imaging (OR, 2.19; CI, 1.78–2.68), high-complexity laboratory (OR, 2.15; CI, 1.24–3.70), affiliation with a medical school (OR, 1.57; CI, 1.31–1.88), and reconstructive surgery (OR, 1.23; CI, 1.01–1.50). Significant regional predictors included metropolitan county (OR, 3.77; CI, 2.83–5.03), above the mean income (OR, 1.37; CI, 1.11–1.69), and education (OR, 1.26; CI, 1.03–1.54). Negative predictors included designation as a critical-access hospital (OR, 0.10; CI, 0.07–0.14) and distance from an NCI cancer center (OR, 0.998; CI, 0.997–0.999), with a 15% decrease in likelihood for every 100 miles.
Concordance with clinical guidelines
The nationwide hospital-level analysis described above did not demonstrate whether the test was performed in accordance with clinical practice guidelines. SEER data allowed us to look at characteristics of patients with the 21-gene test in light of guidelines. Table 4 presents the results of this analysis. Of all patients who had the test reported to SEER (N = 8,673), 79–84% were per-guideline recommendations. The majority of the tests performed outside the guideline recommendations were in LN+ patients (9.4%).
Discussion
This is the first study to analyze national implementation of the 21-gene breast cancer test across all payer groups. This study also illustrated the robustness of our model, which uses institutional and regional factors to analyze implementation of cancer genomics. Our analysis demonstrates that there has been considerable market penetration of the 21-gene breast cancer test. Nearly half of all acute-care hospitals in the United States have ordered the test.
When compared with lung cancer molecular testing,22 implementation of the 21-gene breast cancer test is widely distributed across the country. Although the percentage of patients tested ranged from 11.96 to 73.00%, this range is not particularly broad, especially given potential variations in age, comorbidities, stage, and incidence of ER+ breast cancer. Despite widespread use of the test, there remain important regional and site-of-care variations that require further analysis. Breast cancer patients who obtain care at hospitals affiliated with NCI and/or medical schools that offer specialized services such as positron emission tomography scans and reconstructive surgeries and have CLIA-accredited laboratories are most likely to get tested. Breast cancer patients who obtain care at critical-access hospitals or in rural states are less likely to be tested. There also may be some barriers to access for patients who live in states that lack NCI cancer centers or hospitals participating in cooperative group clinical trials, and that lack hospitals with high-complexity laboratories. Within the Veterans Healthcare Administration, we have made a proactive effort to ensure equity in access to genomics for veterans in rural areas. We offer telemedicine services, virtual tumor boards, and a centralized genomic medicine program that provides genetic consultation services to 66 pathology laboratories across the country. Third-party payers may consider offering similar services for patients accessing care in rural hospitals.
This hospital-level analysis did not enable us to determine whether individual cases met the eligibility criteria (i.e., ER+/HER2–, LN– patients, for whom the test result would affect clinical decisions). Thus, it is not possible to calculate actual utilization rates per eligible cases. We can, however, calculate the upper limit of possible utilization by using the estimated number of guideline-recommended cases as the denominator. The 2007 American Society of Clinical Oncology guidelines, which were applicable during the time covered by our study, recommended that all ER+/HER2–, LN– patients undergo testing. The population that met these criteria can be estimated based on the data from a large study by Howlader et al.,27 who showed that 44.14% of all breast cancer cases with known breast cancer subtypes were both lymph node–negative and HR+/HER2–. HR+ breast cancer subtype is presumed to be ER+ because PR expression requires active ER, and ER–/PR+ breast cancer cases are extremely rare.28,29 The 44.14% rate applied to all newly diagnosed invasive cases in 2011 yields an estimated number of 97,842 eligible patients. Thus, if the 54,985 tests identified in our study were ordered only for guideline-recommended patients, then it suggests that 56.20% of eligible patients were tested. The actual rate is probably lower, given that some LN+ as well as ER– or HER2+ patients could have been tested. The 2011 SEER registry data gave us the opportunity to estimate the proportion of tests performed per guidelines, which varied from 79 to 84% depending on the selection criteria.
Our SEER registry-based analysis demonstrated that testing was mostly concordant with clinical practice guidelines because there were relatively few patients undergoing the test who were not eligible. The largest group tested outside the guidelines was LN+ patients. However, guidelines have been evolving and the latest National Comprehensive Cancer Network recommendation excludes LN– patients with the smallest low-grade tumors but includes a subset of LN+ patients with limited (1–3) lymph node involvement.30 There has been site-of-care and regional variations in implementation of these guidelines. Therefore, some of the utilization that we identified in our hospital-level analysis probably corresponds to hospitals that followed the broader National Comprehensive Cancer Network guidelines rather than the American Society of Clinical Oncology guidelines. As more clinical data on test validation are collected, the proportion of LN-positive patients to whom multigene expression testing is applicable will probably increase.
Institutional and regional-level analyses are valuable tools to obtain a broad overview of implementation of health-care innovations. Site-of-care analysis can identify specific hospitals that have either overutilization or underutilization of specific health-care innovations. However, there are several limitations to a data set that does not capture patient-level characteristics. It is not possible to calculate the number of breast cancer patients being served by each hospital. Therefore, there is no way to calculate a hospital-level utilization rate. Although it is possible to calculate county-level utilization using this method, there is no way to identify whether patients being served by the various hospitals within the county are actually residents of that county and therefore counted in the cancer registry data. Further, without population-level clinical data, it is not possible to separate delays in translation due to hospital and regional factors and appropriately lower levels of implementation due to variations in patient characteristics.
Numerous reports have demonstrated that breast cancer subtype distribution varies by a number of factors, including age at diagnosis, race/ethnicity, and socioeconomic status.31,32,33,34 Howlader et al.27 found that the proportion of the HR+/HER2– subtype among non-Hispanic black patients was 60.2% compared with 75.5% among non-Hispanic whites. Also, in that study, ~12% of cases overall had unknown HR/HER2 status, but that proportion varied significantly across SEER registries, ranging from 4.5 to 22.1%, and was higher among blacks, Hispanics, older patients, and those diagnosed with more advanced disease. Kohler et al.25 expanded this analysis to include data covering 84% of the US female population. Consistent with previous results, they found that breast cancer rates of the HR+/HER2– subtype, eligible for the 21-gene test, were highest among non-Hispanic white women. Conversely, incidence rates of triple-negative breast cancer, which is not an appropriate use of the test, were almost double among the non-Hispanic blacks than among non-Hispanic whites (27.2 vs. 14.4 per 100,000 women). Similar results were reported by a recent study of the California Cancer Registry.26 Thus, the actual proportion of eligible patients may vary across regions and hospital types. For example, the black population in Mississippi is 37.40%. A portion of the low testing rate in Mississippi is very likely explained by a higher rate of triple-negative breast cancer.
Our data are in agreement with several other studies. A study of 7,375 women diagnosed from 2006 to 2008 with stage I to stage III HR+ breast cancer at a large number of hospitals in different geographic areas and practice settings found that utilization of gene expression profile testing varies within different patient populations.17 For example, black women and those with less education (high school degree or less) were 36 and 35% less likely to undergo testing than white non-Hispanic women and those with more than a high school education, respectively. Also, gene expression testing was less common among community-based hospitals compared with comprehensive cancer centers. Our previous research identified substantial underuse of genomic tests among hospitals located in nonmetropolitan counties that are distant from NCI cancer centers.22 Medicaid coverage for the 21-gene test became available in 2007. However, a study conducted in New York City found that site of care significantly influenced utilization, and women treated at municipal hospitals were unlikely to get the 21-gene test even after Medicaid coverage should have ensured equal access.21
Although our analysis identifies variations in testing that are potentially disparities in access, caution is needed in this interpretation. Kohler et al. showed that breast cancer rates of the HR+/HER2– subtype decreased with increasing poverty for every racial and ethnic group.25 Also, incidence rates of HR+/HER2– breast cancer are highest for early-stage cases, which increase with advances in early detection. These authors point out that access to mammography, more common among affluent populations, appears to drive HR+/HER2– breast cancer rates.25 Thus, we can expect that less testing among patients of low socioeconomic status can be partially explained by their clinical characteristics (which are most likely the result of both biologic and social factors) and not simply by lack of access to genomic testing. More research using patient-level data documenting individual clinical characteristics will help clarify these issues. Collectively, these findings indicate that interventions are needed to ensure that all patients are obtaining access to guideline-recommended genomic testing.
The landscape of genomic testing and implementation science is rapidly evolving. There is a concerted effort among cancer registrars to collect and report molecular test data. NCI has increased funding for research on implementation of genomic medicine. In 2013, the American Medical Association published 101 new gene- and test-specific billing codes,35 making it possible to identify individual tests in claims records. Although our analysis of 2011 data is the first comprehensive insight into nationwide implementation of the 21-gene test, these changes will enable much more timely analyses.
Disclosure
Genomic Health, Inc., is currently funding a multisite clinical utility study of their prostate gene expression test within the VA; J.A.L. is the principal investigator of this study. The other authors declare no conflict of interest.
References
Yersal O, Barutca S. Biological subtypes of breast cancer: prognostic and therapeutic implications. World J Clin Oncol 2014;5:412–424.
Cadoo KA, Fornier MN, Morris PG. Biological subtypes of breast cancer: current concepts and implications for recurrence patterns. Q J Nucl Med Mol Imaging 2013;57:312–321.
National Cancer Institute. Breast Cancer Treatment—for health professionals (PDQ®). 2014. http://www.cancer.gov/cancertopics/pdq/treatment/breast/healthprofessional. Accessed 6 January 2015.
Stuart-Harris R, Davis A. Optimal adjuvant endocrine therapy for early breast cancer. Womens Health (Lond Engl) 2010;6:383–398.
Peto R, Davies C, Godwin J, et al.; Early Breast Cancer Trialists’ Collaborative Group (EBCTCG). Comparisons between different polychemotherapy regimens for early breast cancer: meta-analyses of long-term outcome among 100,000 women in 123 randomised trials. Lancet 2012;379:432–444.
Bedard PL, Cardoso F. Can some patients avoid adjuvant chemotherapy for early-stage breast cancer? Nat Rev Clin Oncol 2011;8:272–279.
Cobain EF, Hayes DF. Indications for prognostic gene expression profiling in early breast cancer. Curr Treat Options Oncol 2015;16:23.
Ademuyiwa FO, Miller A, O’Connor T, et al. The effects of oncotype DX recurrence scores on chemotherapy utilization in a multi-institutional breast cancer cohort. Breast Cancer Res Treat 2011;126:797–802.
Biroschak JR, Schwartz GF, Palazzo JP, et al. Impact of oncotype DX on treatment decisions in ER-positive, node-negative breast cancer with histologic correlation. Breast J 2013;19:269–275.
Paik S, Shak S, Tang G, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 2004;351:2817–2826.
Habel LA, Shak S, Jacobs MK, et al. A population-based study of tumor gene expression and risk of breast cancer death among lymph node-negative patients. Breast Cancer Res 2006;8:R25.
Sparano JA, Gray RJ, Makower DF, et al. Prospective validation of a 21-gene expression assay in breast cancer. N Engl J Med 2015;373:2005–2014.
Hornberger J, Alvarado MD, Rebecca C, Gutierrez HR, Yu TM, Gradishar WJ. Clinical validity/utility, change in practice patterns, and economic implications of risk stratifiers to predict outcomes for early-stage breast cancer: a systematic review. J Natl Cancer Inst 2012;104:1068–1079.
Issa AM, Chaudhari VS, Marchant GE. The value of multigene predictors of clinical outcome in breast cancer: an analysis of the evidence. Expert Rev Mol Diagn 2014:1–10.
Lamond NW, Skedgel C, Younis T. Is the 21-gene recurrence score a cost-effective assay in endocrine-sensitive node-negative breast cancer? Expert Rev Pharmacoecon Outcomes Res 2013;13:243–250.
Rouzier R, Pronzato P, Chéreau E, Carlson J, Hunt B, Valentine WJ. Multigene assays and molecular markers in breast cancer: systematic review of health economic analyses. Breast Cancer Res Treat 2013;139:621–637.
Hassett MJ, Silver SM, Hughes ME, et al. Adoption of gene expression profile testing and association with use of chemotherapy among women with breast cancer. J Clin Oncol 2012;30:2218–2226.
Burken MI, Wilson KS, Heller K, Pratt VM, Schoonmaker MM, Seifter E. The interface of Medicare coverage decision-making and emerging molecular-based laboratory testing. Genet Med 2009;11:225–231.
Dinan MA, Mi X, Reed SD, Hirsh BR, Lyman GH, Curtis LH. Initial trends in the use of the 21-gene recurrence score assay for patients with breast cancer in the Medicare Population, 2005–2009. JAMA Oncol 2015;1:158–166.
DeFrank JT, Salz T, Reeder-Hayes K, Brewer NT. Who gets genomic testing for breast cancer recurrence risk? Public Health Genomics 2013;16:215–222.
Guth AA, Fineberg S, Fei K, Franco R, Bickell NA. Utilization of oncotype DX in an inner city population: race or place? Int J Breast Cancer 2013;2013:653805.
Lynch JA, Khoury MJ, Borzecki A, et al. Utilization of epidermal growth factor receptor (EGFR) testing in the United States: a case study of T3 translational research. Genet Med 2013;15:630–638.
US Cancer Statistics Working Group. United States Cancer Statistics: 1999–2011 Incidence and Mortality Data. US Department of Health and Human Services, Centers for Disease Control and Prevention and National Cancer Institute, 2014. http://www.cdc.gov/uscs or http://statecancerprofiles.cancer.gov/index.html.
Centers for Medicare & Medicaid Services. Clinical Laboratory Improvement Amendments (CLIA). https://www.cms.gov/Regulations-and-Guidance/Legislation/CLIA/index.html.
Kohler BA, Sherman RL, Howlader N, et al. Annual report to the nation on the status of cancer, 1975–2011, featuring incidence of breast cancer subtypes by race/ethnicity, poverty, and state. J Natl Cancer Inst 2015;107:djv048.
Parise CA, Caggiano V. The influence of socioeconomic status on racial/ethnic disparities among the ER/PR/HER2 breast cancer subtypes. J Cancer Epidemiol 2015;2015:813456.
Howlader N, Altekruse SF, Li CI, et al. US incidence of breast cancer subtypes defined by joint hormone receptor and HER2 status. J Natl Cancer Inst 2014;106.
Nadji M, Gomez-Fernandez C, Ganjei-Azar P, Morales AR. Immunohistochemistry of estrogen and progesterone receptors reconsidered: experience with 5,993 breast cancers. Am J Clin Pathol 2005;123:21–27.
Hefti MM, Hu R, Knoblauch NW, et al. Estrogen receptor negative/progesterone receptor positive breast cancer is not a reproducible subtype. Breast Cancer Res 2013;15:R68.
Chung C, Christianson M. Predictive and prognostic biomarkers with therapeutic targets in breast, colorectal, and non-small cell lung cancers: a systemic review of current development, evidence, and recommendation. J Oncol Pharm Pract 2014;20:11–28.
Carey LA, Perou CM, Livasy CA, et al. Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA 2006;295:2492–2502.
Bauer KR, Brown M, Cress RD, Parise CA, Caggiano V. Descriptive analysis of estrogen receptor (ER)-negative, progesterone receptor (PR)-negative, and HER2-negative invasive breast cancer, the so-called triple-negative phenotype: a population-based study from the California cancer Registry. Cancer 2007;109:1721–1728.
Kwan ML, Kushi LH, Weltzien E, et al. Epidemiology of breast cancer subtypes in two prospective cohort studies of breast cancer survivors. Breast Cancer Res 2009;11:R31.
Chuang E, Paul C, Flam A, et al. Breast cancer subtypes in Asian-Americans differ according to Asian ethnic group. J Immigr Minor Health 2012;14:754–758.
American Medical Association. Coding With CPT® for Proper Reimbursement. http://www.ama-assn.org/ama/pub/physician-resources/solutions-managing-your-practice/coding-billing-insurance/cpt.page?. Accessed 2 October 2015.
Acknowledgements
This research was conducted while J.A.L. was a postdoctoral fellow within the Center for Healthcare Organization and Implementation Research (CHOIR), a Veterans Administration Health Services Research & Development Center of Excellence. She and B.B. are funded by the National Cancer Institute through an Interagency Agreement.
The opinions in this article reflect those of the authors and do not necessarily reflect the official position of the Department of Health and Human Services or the Veterans Health Administration. Genomic Health, Inc., did not sponsor this study. It provided its sales data but had no role in data analysis, interpretation, or manuscript writing.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figure
(EPS 4707 kb)
Supplementary Table
(XLS 184 kb)
Rights and permissions
About this article
Cite this article
Lynch, J., Berse, B., Petkov, V. et al. Implementation of the 21-gene recurrence score test in the United States in 2011. Genet Med 18, 982–990 (2016). https://doi.org/10.1038/gim.2015.218
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/gim.2015.218