Abstract
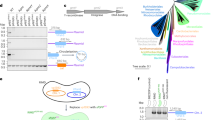
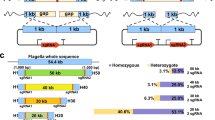
The variability in genome content among closely related strains of prokaryotes has been one of the most remarkable discoveries of genomics. One way to approach the description of this so-called pan-genome is to compare one reference strain genome with metagenomic sequences from the environment. We have applied this approach to one extreme aquatic habitat, saturated brines in a solar saltern. The genome of Haloquadratum walsbyi strain DSM 16790 was compared to an environmental metagenome obtained from the exact site of its isolation. This approach revealed that some regions of the strain genome were scarcely represented in the metagenome. Here we have analyzed these genomic islands (GI) in the genome of DSM 16790 and compared them with the complete sequence of some fosmids from the environmental library. Two of the islands, GI 2 and GI 4, overlapped with two large guanine and cytosine (GC)-rich regions that showed evidence of high variability through mobile elements. GI 3 seemed to be a phage or phage-remnant acquired by the reference genome, but not present in most environmental lineages. Most differential gene content was related to small molecule transport and detection, probably reflecting adaptation to different pools of organic nutrients. GI 1 did not possess traces of mobile elements and had normal GC content. This island contained the main cluster of cell envelope glycoproteins and the variability found was different from the other GIs. Rather than containing different genes it consisted of homologs with low similarity. This variation might reflect a phage evasion strategy.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Accession codes
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Angly FE, Felts B, Breitbart M, Salamon P, Edwards RA, Carlson C et al. (2006). The marine viromes of four oceanic regions. PLoS Biol 4: e368.
Blaurock AE, Stoeckenius W, Oesterhelt D, Scherfhof GL . (1976). Structure of the cell envelope of Halobacterium halobium. J Cell Biol 71: 1–22.
Bolhuis HH, Palm PP, Wende AA, Falb MM, Rampp MM, Rodriguez-Valera FF et al. (2006). The genome of the square archaeon Haloquadratum walsbyi: life at the limits of water activity. BMC Genomics 7: 169.
Bolhuis H, Poele EM, Rodriguez-Valera F . (2004). Isolation and cultivation of Walsby's square archaeon. Environ Microbiol 6: 1287–1291.
Breitbart M, Rohwer F . (2005). Here a virus, there a virus, everywhere the same virus? Trends Microbiol 13: 278–284.
Carver TJ, Rutherford KM, Berriman M, Rajandream MA, Barrell BG, Parkhill J . (2005). ACT: the artemis comparison tool. Bioinformatics 21: 3422–3423.
Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG et al. (2003). Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res 31: 3497–3500.
Coleman ML, Sullivan MB, Martiny AC, Steglich C, Barry K, Delong EF et al. (2006). Genomic islands and the ecology and evolution of prochlorococcus. Science 311: 1768–1770.
Delcher AL, Harmon D, Kasif S, White O, Salzberg SL . (1999). Improved microbial gene identification with GLIMMER. Nucleic Acids Res 27: 4636–4641.
Dempsey MP, Nietfeldt J, Ravel J, Hinrichs S, Crawford R, Benson AK . (2006). Paired-end sequence mapping detects extensive genomic rearrangement and translocation during divergence of Francisella tularensis subsp. tularensis and Francisella tularensis subsp. holarctica populations. J Bacteriol 188: 5904–5914.
Edgar RC . (2004). MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5: 113.
Gasol JM, Joint I, Kristine G, Gustavson K, Benlloch S, Díez B et al. (2004). Control of heterotrophic prokaryotic abundance and growth rate in hypersaline planktonic environments. Aquat Microb Ecol 34: 193–206.
Green J, Bohannan BJ . (2006). Spatial scaling of microbial biodiversity. Trends Ecol Evol 21: 501–507.
Guixa-Boixereu N . (1996). Viral lysis and bacterivory as prokaryotic loss factors along a salinity gradient. Aquat Microb Ecol 11: 213–227.
Guixa-Boixereu N, Lysnes K, Pedros-Alio C . (1999). Viral lysis and bacterivory during a phytoplankton bloom in a coastal water microcosm. Appl Environ Microbiol 65: 1949–1958.
He Q, Qiao D, Bai L, Zhang Q, Yang W, Li Q et al. (2007). Cloning and characterization of a plastidic glycerol 3-phosphate dehydrogenase cDNA from Dunaliella salina. J Plant Physiol 64: 214–220.
Hochhut B, Wilde C, Balling G, Middendorf B, Dobrindt U, Brzuszkiewicz E et al. (2006). Role of pathogenicity island-associated integrases in the genome plasticity of uropathogenic Escherichia coli strain 536. Mol Microbiol 61: 584–595.
Hughes D . (2000). Co-evolution of the tuf genes links gene conversion with the generation of chromosomal inversions. J Mol Biol 297: 355–364.
Kumar S, Tamura K, Nei M . (2004). MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5: 150–163.
Kurtz S, Phillippy A, Delcher AL, Smoot M, Shumway M, Antonescu C et al. (2004). Versatile and open software for comparing large genomes. Genome Biol 5: R12.
Larimer FW, Chain P, Hauser L, Lamerdin J, Malfatti S, Do L et al. (2004). Complete genome sequence of the metabolically versatile photosynthetic bacterium Rhodopseudomonas palustris. Nat Biotechnol 22: 55–61.
Legault BA, Lopez-Lopez A, Alba-Casado JC, Doolittle WF, Bolhuis H, Rodriguez-Valera F et al. (2006). Environmental genomics of ‘Haloquadratum walsbyi’ in a saltern crystallizer indicates a large pool of accessory genes in an otherwise coherent species. BMC Genomics 7: 171.
Lerat E, Daubin V, Ochman H, Moran NA . (2005). Evolutionary origins of genomic repertoires in bacteria. PLoS Biol 3: e130.
Mackiewicz P, Mackiewicz D, Kowalczuk M, Cebrat S . (2001). Flip-flop around the origin and terminus of replication in prokaryotic genomes. Genome Biol 2: interactions 1004.1–1004.4.
Mengele R, Sumper M . (1992). Drastic differences in glycosylation of related S-layer glycoproteins from moderate and extreme halophiles. J Biol Chem 267: 8182–8185.
Pedros-Alio C, Calderon-Paz JI, MacLean MH, Medina G, Marrase C, Gasol JM . (2000). The microbial food web along salinity gradients. FEMS Microbiol Ecol 32: 143–155.
Petrosino JF, Xiang Q, Karpathy SE, Jiang H, Yerrapragada S, Liu Y et al. (2006). Chromosome rearrangement and diversification of Francisella tularensis revealed by the type B (OSU18) genome sequence. J Bacteriol 188: 6977–6985.
Phadwal K, Singh PK . (2003). Effect of nutrient depletion on beta-carotene and glycerol accumulation in two strains of Dunaliella sp. Bioresour Technol 90: 55–58.
Read TD, Ussery DW . (2006). Opening the pan-genomics box. Curr Opin Microbiol 9: 496–498.
Rice P, Longden I, Bleasby A . (2000). EMBOSS: the European molecular biology open software suite. Trends Genet 16: 276–277.
Schaffer C, Messner P . (2001). Glycobiology of surface layer proteins. Biochimie 83: 591–599.
Schaffer C, Graninger M, Messner P . (2001). Prokaryotic glycosylation. Proteomics 1: 248–261.
Schauer R . (2000). Achievements and challenges of sialic acid research. Glycoconj J 17: 485–499.
Sullivan MB, Coleman ML, Weigele P, Rohwer F, Chisholm SW . (2005). Three Prochlorococcus cyanophage genomes: signature features and ecological interpretations. PLoS Biol 3: e144.
Tettelin H, Masignani V, Cieslewicz MJ, Donati C, Medini D, Ward NL et al. (2005). Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial ‘pan-genome’. Proc Natl Acad Sci USA 102: 13950–13955.
Thingstad T . (2000). Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic ecosystems. Limnol Oceanogr 45: 1320–1328.
Thompson JD, Higgins DG, Gibson TJ . (1994). CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22: 4673–4680.
Thompson JR, Pacocha S, Pharino C, Klepac-Ceraj V, Hunt DE, Benoit J et al. (2005). Genotypic diversity within a natural coastal bacterioplankton population. Science 307: 1311–1313.
Trachtenberg S, Pinnick B, Kessel M . (2000). The cell surface glycoprotein layer of the extreme halophile Halobacterium salinarum and its relation to Haloferax volcanii: cryo-electron tomography of freeze-substituted cells and projection studies of negatively stained envelopes. J Struct Biol 130: 10–26.
Tyson GW, Chapman J, Hugenholtz P, Allen EE, Ram RJ, Richardson PM et al. (2004). Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428: 37–43.
Walsby AE . (2005). Archaea with square cells. Trends Microbiol 13: 193–195.
Wang L, Andrianopoulos K, Liu D, Popoff MY, Reeves PR . (2002). Extensive variation in the O-antigen gene cluster within one Salmonella enterica serogroup reveals an unexpected complex history. J Bacteriol 184: 1669–1677.
Willenbrock H, Petersen A, Sekse C, Kiil K, Wasteson Y, Ussery DW . (2006). Design of a seven-genome Escherichia coli microarray for comparative genomic profiling. J Bacteriol 188: 7713–7721.
Xiang SH, Hobbs M, Reeves PR . (1994). Molecular analysis of the rfb gene cluster of a group D2 Salmonella enterica strain: evidence for its origin from an insertion sequence-mediated recombination event between group E and D1 strains. J Bacteriol 176: 4357–4365.
Acknowledgements
SCO acknowledges CAPES Foundation (Brazil) for the postdoctoral fellowship received. ABMC is supported by a MEC (Spain) Postdoctoral Fellowship. OZ is supported through a CIHR Postdoctoral Fellowship and is an honorary Killam Postdoctoral Fellow at Dalhousie University. This work was funded by the GEMINI (QLK3-CT-2002–02056) project of the European Commission, and MEC (Spain) (CTM2005–04564) ‘Metagenomic study of bacterioplankton of deep Mediterranean Sea’ project.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Rights and permissions
About this article
Cite this article
Cuadros-Orellana, S., Martin-Cuadrado, AB., Legault, B. et al. Genomic plasticity in prokaryotes: the case of the square haloarchaeon. ISME J 1, 235–245 (2007). https://doi.org/10.1038/ismej.2007.35
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ismej.2007.35
Keywords
This article is cited by
-
Distinct ecotypes within a natural haloarchaeal population enable adaptation to changing environmental conditions without causing population sweeps
The ISME Journal (2021)
-
Genomic variation and biogeography of Antarctic haloarchaea
Microbiome (2018)
-
Genomic variation in microbial populations inhabiting the marine subseafloor at deep-sea hydrothermal vents
Nature Communications (2017)
-
New insights into marine group III Euryarchaeota, from dark to light
The ISME Journal (2017)
-
Gene-specific selective sweeps in bacteria and archaea caused by negative frequency-dependent selection
BMC Biology (2015)