Abstract
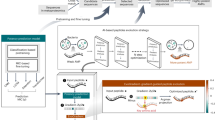
The rapid rise of antibiotic-resistant bacteria is one of the major concerns in modern medicine. Therefore, to treat bacterial infections, there is an urgent need for new antibacterials–preferably directed against alternative bacterial targets. One such potential target is the preprotein translocation motor SecA. SecA is a peripheral membrane ATPase and a key component of the Sec secretion pathway, the major route for bacterial protein export across or into the cytoplasmic membrane. As SecA is essential for bacterial viability, ubiquitous and highly conserved in bacteria, but not present in eukaryotic cells, it represents an attractive antibacterial target. Using an in silico approach, we have defined several potentially druggable and conserved pockets on the surface of SecA. We show that three of these potentially druggable sites are important for SecA function. A starting collection of ~500 000 commercially available small-molecules was virtually screened against a predicted druggable pocket in the preprotein-binding domain of Escherichia coli SecA using a multi-step virtual ligand screening protocol. The 1040 top-scoring molecules were tested in vitro for inhibition of the translocation ATPase activity of E. coli SecA. Five inhibitors of the translocation ATPase, and not of basal or membrane ATPase, were identified with IC50 values <65 μm. The most potent inhibitor showed an IC50 of 24 μm. The antimicrobial activity was determined for the five most potent SecA inhibitors. Two compounds were found to possess weak antibacterial activity (IC50 ~198 μm) against E. coli, whereas some compounds showed moderate antibacterial activity (IC50 ~100 μm) against Staphylococcus aureus.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Rodriguez-Rojas, A., Rodriguez-Beltran, J., Couce, A. & Blazquez, J. Antibiotics and antibiotic resistance: a bitter fight against evolution. Int. J. Med. Microbiol. 303, 293–297 (2013).
Nordmann, P., Naas, T., Fortineau, N. & Poirel, L. Superbugs in the coming new decade; multidrug resistance and prospects for treatment of Staphylococcus aureus, Enterococcus spp. and Pseudomonas aeruginosa in 2010. Curr. Opin. Microbiol. 10, 436–440 (2007).
Berdy, J. Thoughts and facts about antibiotics: where we are now and where we are heading. J. Antibiot. (Tokyo) 65, 441 (2012).
Cooper, M. A. & Shlaes, D. Fix the antibiotics pipeline. Nature 472, 32 (2011).
Stephens, C. & Shapiro, L. Bacterial protein secretion—a target for new antibiotics? Chem. Biol. 4, 637–641 (1997).
Economou, A. Sec, drugs and rock'n'roll: antibiotic targeting of bacterial protein translocation. Expert Opin. Ther. Targets 5, 141–153 (2001).
Rao, C. V. S., De Waelheyns, E., Economou, A. & Anne, J. Antibiotic targeting of the bacterial secretory pathway. Biochim. Biophys. Acta 1843, 1762–1783 (2014).
Kusters, I. & Driessen, A. J. SecA, a remarkable nanomachine. Cell. Mol. Life Sci. 68, 2053–2066 (2011).
Chatzi, K. E., Sardis, M. F., Economou, A. & Karamanou, S. SecA-mediated targeting and translocation of secretory proteins. Biochim. Biophys. Acta 1843, 1466–1474 (2014).
Gouridis, G. et al. Quaternary dynamics of the SecA motor drive translocase catalysis. Mol. Cell 52, 655–666 (2013).
Lill, R., Dowhan, W. & Wickner, W. The ATPase activity of SecA is regulated by acidic phospholipids, SecY, and the leader and mature domains of precursor proteins. Cell 60, 271–280 (1990).
Caruthers, J. M. & McKay, D. B. Helicase structure and mechanism. Curr. Opin. Struct. Biol. 12, 123–133 (2002).
Sianidis, G. et al. Cross-talk between catalytic and regulatory elements in a DEAD motor domain is essential for SecA function. EMBO J. 20, 961–970 (2001).
Hunt, J. F. et al. Nucleotide control of interdomain interactions in the conformational reaction cycle of SecA. Science 297, 2018–2026 (2002).
Papanikolau, Y. et al. Structure of dimeric SecA, the Escherichia coli preprotein translocase motor. J. Mol. Biol. 366, 1545–1557 (2007).
Zimmer, J., Nam, Y. & Rapoport, T. A. Structure of a complex of the ATPase SecA and the protein-translocation channel. Nature 455, 936–943 (2008).
Gelis, I. et al. Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell 131, 756–769 (2007).
Sugie, Y. et al. CJ-21,058, a new SecA inhibitor isolated from a fungus. J. Antibiot. (Tokyo) 55, 25–29 (2002).
Vrontou, E., Karamanou, S., Baud, C., Sianidis, G. & Economou, A. Global co-ordination of protein translocation by the SecA IRA1 switch. J. Biol. Chem. 279, 22490–22497 (2004).
Segers, K., Klaassen, H., Economou, A., Chaltin, P. & Anne, J. Development of a high-throughput screening assay for the discovery of small-molecule SecA inhibitors. Anal. Biochem. 413, 90–96 (2011).
Segers, K. & Anne, J. Traffic jam at the bacterial sec translocase: targeting the SecA nanomotor by small-molecule inhibitors. Chem. Biol. 18, 685–698 (2011).
Jang, M. Y., De Jonghe, S., Segers, K., Anne, J. & Herdewijn, P. Synthesis of novel 5-amino-thiazolo[4,5-d]pyrimidines as E. coli and S. aureus SecA inhibitors. Bioorg. Med. Chem. 19, 702–714 (2011).
Huang, Y. J. et al. Fluorescein analogues inhibit SecA ATPase: the first sub-micromolar inhibitor of bacterial protein translocation. ChemMedChem 7, 571–577 (2012).
Parish, C. A. et al. Antisense-guided isolation and structure elucidation of pannomycin, a substituted cis-decalin from Geomyces pannorum. J. Nat. Prod 72, 59–62 (2009).
Alksne, L. E. et al. Identification and analysis of bacterial protein secretion inhibitors utilizing a SecA-LacZ reporter fusion system. Antimicrob. Agents Chemother. 44, 1418–1427 (2000).
Li, M., Huang, Y. J., Tai, P. C. & Wang, B. Discovery of the first SecA inhibitors using structure-based virtual screening. Biochem. Biophys. Res. Commun. 368, 839–845 (2008).
Chen, W. et al. The first low microM SecA inhibitors. Bioorg. Med. Chem. 18, 1617–1625 (2010).
Akula, N., Zheng, H., Han, F. Q. & Wang, N. Discovery of novel SecA inhibitors of Candidatus Liberibacter asiaticus by structure based design. Bioorg. Med. Chem. Lett. 21, 4183–4188 (2011).
Akula, N., Trivedi, P., Han, F. Q. & Wang, N. Identification of small molecule inhibitors against SecA of Candidatus Liberibacter asiaticus by structure based design. Eur. J. Med. Chem. 54, 919–924 (2012).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Schwede, T., Kopp, J., Guex, N. & Peitsch, M. C. SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res. 31, 3381–3385 (2003).
An, J., Totrov, M. & Abagyan, R. Pocketome via comprehensive identification and classification of ligand binding envelopes. Mol. Cell. Proteomics 4, 752–761 (2005).
Miteva, M. A., Lee, W. H., Montes, M. O. & Villoutreix, B. O. Fast structure-based virtual ligand screening combining FRED, DOCK, and Surflex. J. Med. Chem. 48, 6012–6022 (2005).
Du, J., Bleylevens, I. W., Bitorina, A. V., Wichapong, K. & Nicolaes, G. A. Optimization of compound ranking for structure-based virtual ligand screening using an established FRED-Surflex consensus approach. Chem. Biol. Drug Des. 83, 37–51 (2014).
Lagorce, D., Sperandio, O., Galons, H., Miteva, M. A. & Villoutreix, B. O. FAF-Drugs2: free ADME/tox filtering tool to assist drug discovery and chemical biology projects. BMC Bioinformatics 9, 396 (2008).
Hawkins, P. C., Skillman, A. G., Warren, G. L., Ellingson, B. A. & Stahl, M. T. Conformer generation with OMEGA: algorithm and validation using high quality structures from the Protein Databank and Cambridge Structural Database. J. Chem. Inf. Model 50, 572–584 (2010).
McGann, M. FRED pose prediction and virtual screening accuracy. J. Chem. Inf. Model. 51, 578–596 (2011).
Jain, A. N. Surflex: fully automatic flexible molecular docking using a molecular similarity-based search engine. J. Med. Chem. 46, 499–511 (2003).
Mitchell, C. & Oliver, D. Two distinct ATP-binding domains are needed to promote protein export by Escherichia coli SecA ATPase. Mol. Microbiol. 10, 483–497 (1993).
Klein, M., Meens, J. & Freudl, R. Functional characterization of the Staphylococcus carnosus SecA protein in Escherichia coli and Bacillus subtilis secA mutant strains. FEMS Microbiol. Lett. 131, 271–277 (1995).
Anagnostopoulos, C. & Spizizen, J. Requirements for Transformation in Bacillus subtilis. J. Bacteriol. 81, 741–746 (1961).
Karamanou, S. et al. A molecular switch in SecA protein couples ATP hydrolysis to protein translocation. Mol. Microbiol. 34, 1133–1145 (1999).
Gouridis, G., Karamanou, S., Koukaki, M. & Economou, A. In vitro assays to analyze translocation of the model secretory preprotein alkaline phosphatase. Methods Mol. Biol. 619, 157–172 (2010).
Gouridis, G., Karamanou, S., Gelis, I., Kalodimos, C. G. & Economou, A. Signal peptides are allosteric activators of the protein translocase. Nature 462, 363–367 (2009).
Abagyan, R., Totrov, M. & Kuznetsov, D. ICM: a new method for protein modeling and design: applications to docking and structure prediction from the distorted native conformation. J. Comp. Chem. 15, 488–506 (1994).
Sardis, M. F. & Economou, A. SecA: a tale of two protomers. Mol. Microbiol. 76, 1070–1081 (2010).
Lill, R. et al. SecA protein hydrolyzes ATP and is an essential component of the protein translocation ATPase of Escherichia coli. EMBO J. 8, 961–966 (1989).
Cunningham, K. et al. SecA protein, a peripheral protein of the Escherichia coli plasma membrane, is essential for the functional binding and translocation of proOmpA. EMBO J. 8, 955–959 (1989).
Auclair, S. M. et al. Mapping of the signal peptide-binding domain of Escherichia coli SecA using Forster resonance energy transfer. Biochemistry 49, 782–792 (2010).
Kourtz, L. & Oliver, D. Tyr-326 plays a critical role in controlling SecA-preprotein interaction. Mol. Microbiol. 37, 1342–1356 (2000).
Segers, K. et al. Design of protein membrane interaction inhibitors by virtual ligand screening, proof of concept with the C2 domain of factor V. Proc. Natl Acad. Sci. USA 104, 12697–12702 (2007).
Chatzigeorgiou, A. et al. Blocking CD40-TRAF6 signaling is a therapeutic target in obesity-associated insulin resistance. Proc. Natl Acad. Sci. USA 111, 2686–2691 (2014).
Alvarez, J. & Shoichet, B. Virtual Screening in Drug Discovery, CRC Press, (2005).
Acknowledgements
We are grateful to T. D’huys for his contribution to the cloning of the saSecA1 Site I mutants; M. Kulharia for technical assistance during the initial phase of the virtual screening process; L. Van Berckelaer, J. Balzarini and D. Schols for cytotoxicity tests; and P. Chaltin and H. Klaassen for useful discussions. This work was supported by grants SBO IWT 50146; FWO (Krediet aan Navorsers) 1500912N and ESCMID Grant 2012; the Cardiovascular Research Institute Maastricht (to GAFN); the Transnational University Limburg (to GAFN) and KUL-Spa (Onderzoekstoelagen 2013; Bijzonder Onderzoeksfonds; KU Leuven; to AE); RiMembR (Vlaanderen Onderzoeksprojecten; #G0C6814N; FWO; to AE); an Excellence grant (#1473 to AE; Greek General Secretariat of Research).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Supplementary information
Rights and permissions
About this article
Cite this article
De Waelheyns, E., Segers, K., Sardis, M. et al. Identification of small-molecule inhibitors against SecA by structure-based virtual ligand screening. J Antibiot 68, 666–673 (2015). https://doi.org/10.1038/ja.2015.53
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/ja.2015.53
This article is cited by
-
Inhibitors of protein translocation across membranes of the secretory pathway: novel antimicrobial and anticancer agents
Cellular and Molecular Life Sciences (2018)