Abstract
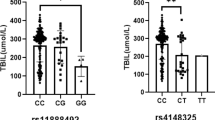
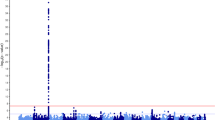
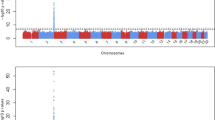
With dense single-nucleotide polymorphism (SNP) maps for 199 drug-related genes, we examined associations between 4190 SNPs and 38 commonly measured quantitative traits using data from 752 healthy Japanese subjects. On analysis, we observed a strong association between five SNPs within the uridine diphosphate glucuronosyltransferase 1A1 (UGT1A1) gene and serum total bilirubin levels (minimum P-value in Mann–Whitney test=1.82 × 1010). UGT1A1 catalyzes the conjugation of bilirubin with glucuronic acid, thus enhancing bilirubin elimination. This enzyme is known to play an important role in the variation of serum bilirubin levels. The five SNPs, including a nonsynonymous SNP—rs4148323 (211G>A or G71R variant allele known as UGT1A1*6)—showed strong linkage disequilibrium with each other. No other genes were clearly associated with serum total bilirubin levels. Results of linear multiple regression analysis on serum total bilirubin levels followed by analysis of variance showed that at least 13% of the variance in serum total bilirubin levels could be explained by three haplotype-tagging SNPs in the UGT1A1 gene.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
References
Lesko, L. J. & Woodcock, J. Translation of pharmacogenomics and pharmacogenetics: a regulatory perspective. Nat. Rev. Drug Discov. 3, 763–769 (2004).
Wilke, R. A., Lin, D. W., Roden, D. M., Watkins, P. B., Flockhart, D., Zineh, I. et al. Identifying genetic risk factors for serious adverse drug reactions: current progress and challenges. Nat. Rev. Drug Discov. 6, 904–916 (2007).
The Wellcome Trust Case Control Consortium. Genome-wide association study of 14 000 cases of seven common diseases and 3000 shared controls. Nature 447, 661–678 (2007).
Kruglyak, L. The road to genome-wide association studies. Nat. Rev. Genet. 9, 314–318 (2008).
Kamatani, N., Sekine, A., Kitamoto, T., Iida, A., Saito, S., Kogame, A. et al. Large-scale single-nucleotide polymorphism (SNP) and haplotype analyses, using dense SNP Maps, of 199 drug-related genes in 752 subjects: the analysis of the association between uncommon SNPs within haplotype blocks and the haplotypes constructed with haplotype-tagging SNPs. Am. J. Hum. Genet. 75, 190–203 (2004).
Frayling, T. M., Timpson, N. J., Weedon, M. N., Zeggini, E., Freathy, R. M., Lindgren, C. M. et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316, 889–894 (2007).
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Ohnishi, Y., Tanaka, T., Ozaki, K., Yamada, R., Suzuki, H. & Nakamura, Y. A high-throughput SNP typing system for genome-wide association studies. J. Hum. Genet. 46, 471–477 (2001).
Price, A. L., Patterson, N. J., Plenge, R. M., Weinblatt, M. E., Shadick, N. A. & Reich, D. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Purcell, S., Neale, B., Todd-Brown, K., Thomas, L., Ferreira, M. A., Bender, D. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
De Bakker, P. I., Yelensky, R., Pe'er, I., Gabriel, S. B., Daly, M. J. & Altshuler, D. Efficiency and power in genetic association studies. Nat. Genet. 37, 1217–1223 (2005).
Shibata, K., Ito, T., Kitamura, Y., Iwasaki, N., Tanaka, H. & Kamatani, N. Simultaneous estimation of haplotype frequencies and quantitative trait parameters: applications to the test of association between phenotype and diplotype configuration. Genetics 168, 525–539 (2004).
Tukey, R. H. & Strassburg, C. P. Human UDP-glucuronosyltransferases: metabolism, expression, and disease. Annu. Rev. Pharmacol. Toxicol. 40, 581–616 (2000).
Kadakol, A., Ghosh, S. S., Sappal, B. S., Sharma, G., Chowdhury, J. R. & Chowdhury, N. R. Genetic lesions of bilirubin uridine-diphosphoglucuronate glucuronosyltransferase (UGT1A1) causing Crigler–Najjar and Gilbert syndromes: correlation of genotype to phenotype. Hum. Mutat. 16, 297–306 (2000).
Hall, D., Ybazeta, G., Destro-Bisol, G., Petzl-Erler, M. L. & Di Rienzo, A. Variability at the uridine diphosphate glucuronosyltransferase 1A1 promoter in human populations and primates. Pharmacogenetics 9, 591–599 (1999).
Akaba, K., Kimura, T., Sasaki, A., Tanabe, S., Ikegami, T., Hashimoto, M. et al. Neonatal hyperbilirubinemia and mutation of the bilirubin uridine diphosphate-glucuronosyltransferase gene: a common missense mutation among Japanese, Koreans and Chinese. Biochem. Mol. Biol. Int. 46, 21–26 (1998).
Yamamoto, K., Sato, H., Fujiyama, Y., Doida, Y. & Bamba, T. Contribution of two missense mutations (G71R and Y486D) of the bilirubin UDP glycosyltransferase (UGT1A1) gene to phenotypes of Gilbert's syndrome and Crigler–Najjar syndrome type II. Biochim. Biophys. Acta. 1406, 267–273 (1998).
Iyer, L., Das, S., Janisch, L., Wen, M., Ramirez, J., Karrison, T. et al. UGT1A1*28 polymorphism as a determinant of irinotecan disposition and toxicity. Pharmacogenomics J. 2, 43–47 (2002).
Jinno, H., Tanaka-Kagawa, T., Hanioka, N., Saeki, M., Ishida, S., Nishimura, T. et al. Glucuronidation of 7-ethyl-10-hydroxycamptothecin (SN-38), an active metabolite of irinotecan (CPT-11), by human UGT1A1 variants, G71R, P229Q, and Y486D. Drug. Metab. Dispos. 31, 108–113 (2003).
Ando, Y., Saka, H., Ando, M., Sawa, T., Muro, K., Ueoka, H. et al. Polymorphisms of UDP-glucuronosyltransferase gene and irinotecan toxicity: a pharmacogenetic analysis. Cancer Res. 60, 6921–6926 (2000).
Innocenti, F., Undevia, S. D., Iyer, L., Chen, P. X., Das, S., Kocherginsky, M. et al. Genetic variants in the UDP-glucuronosyltransferase 1A1 gene predict the risk of severe neutropenia of irinotecan. J. Clin. Oncol. 22, 1382–1388 (2004).
Marcuello, E., Altes, A., Menoyo, A., Del Rio, E., Gomez-Pardo, M. & Baiget, M. UGT1A1 gene variations and irinotecan treatment in patients with metastatic colorectal cancer. Br. J. Cancer 91, 678–682 (2004).
Rouits, E., Boisdron-Celle, M., Dumont, A., Guerin, O., Morel, A. & Gamelin, E. Relevance of different UGT1A1 polymorphisms in irinotecan-induced toxicity: a molecular and clinical study of 75 patients. Clin. Cancer Res. 10, 5151–5159 (2004).
Sai, K., Saeki, M., Saito, Y., Ozawa, S., Katori, N., Jinno, H. et al. UGT1A1 haplotypes associated with reduced glucuronidation and increased serum bilirubin in irinotecan-administered Japanese patients with cancer. Clin. Pharmacol. Ther. 75, 501–515 (2004).
Kaniwa, N., Kurose, K., Jinno, H., Tanaka-Kagawa, T., Saito, Y., Saeki, M. et al. Racial variability in haplotype frequencies of UGT1A1 and glucuronidation activity of a novel single nucleotide polymorphism 686C>T (P229L) found in an African-American. Drug Metab. Dispos. 33, 458–465 (2005).
Minami, H., Sai, K., Saeki, M., Saito, Y., Ozawa, S., Suzuki, K. et al. Irinotecan pharmacokinetics/pharmacodynamics and UGT1A genetic polymorphisms in Japanese: roles of UGT1A1*6 and *28. Pharmacogenet. Genomics 17, 497–504 (2007).
Ransohoff, D. F. & Feinstein, A. R. Problems of spectrum and bias in evaluating the efficacy of diagnostic tests. N. Engl. J. Med. 299, 926–930 (1978).
Tosteson, A. N. & Begg, C. B A general regression methodology for ROC curve estimation. Med. Decis. Making 8, 204–215 (1988).
Pepe, M. S. A regression modelling framework for receiver operating characteristic curve in medical diagnostic testing. Biometrika 84, 595–608 (1997).
Pepe, M. S. Three approaches to regression analysis of receiver operating characteristic curves for continuous test results. Biometrics 54, 124–135 (1998).
Pepe, M. S. An interpretation for the ROC curve and inference using GLM procedures. Biometrics 56, 352–359 (2000).
McClish, D. K. Analyzing a portion of the ROC curve. Med. Decis. Making 9, 190–195 (1989).
Dorfman, D. D. & Alf, E. Maximum likelihood estimation of parameters of signal detection theory and determination of confidence intervals-rating method data. J. Math. Psychol. 6, 487–496 (1969).
Metz, C. E., Herman, B. A. & Shen, J. H. Maximum likelihood estimation of receiver operating characteristic (ROC) curves from continuously-distributed data. Stat. Med. 17, 1033–1053 (1998).
Ieiri, I., Suzuki, H., Kimura, M., Takane, H., Nishizato, Y., Irie, S. et al. Influence of common variants in the pharmacokinetic genes (OATP-C, UGT1A1, and MRP2) on serum bilirubin levels in healthy subjects. Hepatol. Res. 30, 91–95 (2004).
Huang, C. S., Huang, M. J., Lin, M. S., Yang, S. S., Teng, H. C. & Tang, K. S. Genetic factors related to unconjugated hyperbilirubinemia amongst adults. Pharmacogenet. Genomics 15, 43–50 (2005).
Ho, R. H., Choi, L., Lee, W., Mayo, G., Schwarz, U. I., Tirona, R. G. et al. Effect of drug transporter genotypes on pravastatin disposition in European and African-American participants. Pharmacogenet. Genomics 17, 647–656 (2007).
Acknowledgements
This research was supported by Health and Labour Sciences Research Grants from the Ministry of Health, Labour, and Welfare of Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website (http://www.nature.com/jhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Saito, A., Kawamoto, M. & Kamatani, N. Association study between single-nucleotide polymorphisms in 199 drug-related genes and commonly measured quantitative traits of 752 healthy Japanese subjects. J Hum Genet 54, 317–323 (2009). https://doi.org/10.1038/jhg.2009.31
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/jhg.2009.31
Keywords
This article is cited by
-
Pharmacogenetics of tamoxifen therapy in Asian populations: from genetic polymorphism to clinical outcomes
European Journal of Clinical Pharmacology (2021)
-
Interaction of genetic markers associated with serum alkaline phosphatase levels in the Japanese population
Human Genome Variation (2015)