Abstract
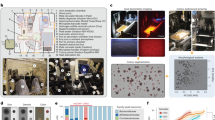
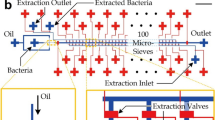
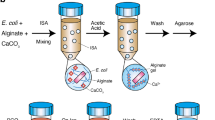
Assembling a complete genome from a single bacterial cell, termed single-cell genomics, is challenging with current technologies. Recovery rates of complete genomes from fragmented assemblies of single-cell templates significantly vary. Although increasing the amount of genomic template material by standard cultivation improves recovery, most bacteria are unfortunately not amenable to traditional cultivation, possibly owing to the lack of unidentified, yet necessary, growth signals and/or specific symbiotic influences. To overcome this limitation, we adopted and modified the method of cocultivation of single-captured bacterial cells in gel microdroplets (GMDs) to improve full genomic sequence recovery. By completing multiple genomes of two novel species derived from single cells, we demonstrated its efficacy on diverse bacterial species using human oral and gut microbiome samples. Here we describe a detailed protocol for capturing single bacterial cells, cocultivating them in medium and isolating microcolonies in GMDs with flow cytometry. Beginning with preliminary studies, obtaining GMDs with single microcolonies for whole-genome amplification may take ∼4 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lasken, R. Single-cell genomic sequencing using multiple displacement amplification. Curr. Opin. Microbiol. 10, 510–516 (2007).
Marcy, Y. et al. Dissecting biological 'dark matter' with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth. Proc. Natl. Acad. Sci. USA 104, 11889–11894 (2007).
Podar, M. et al. Targeted access to the genomes of low-abundance organisms in complex microbial communities. Appl. Environ. Microbiol. 73, 3205–3214 (2007).
Stepanauskas, R. & Sieracki, M.E. Matching phylogeny and metabolism in the uncultured marine bacteria, one cell at a time. Proc. Natl. Acad. Sci. USA 104, 9052–9057 (2007).
Rodrigue, S. et al. Whole genome amplification and de novo assembly of single bacterial cells. PLoS ONE 4, e6864 (2009).
Woyke, T. et al. Assembling the marine metagenome, one cell at a time. PLoS ONE 4, e5299 (2009).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Ballantyne, K.N., Oorschot, R.A., Muharamc, I., Daal, A. & Mitchell, R.J. Decreasing amplification bias associated with multiple displacement amplification and short tandem repeat genotyping. Anal. Biochem. 368, 222–229 (2007).
Pan, X. et al. A procedure for highly specific, sensitive, and unbiased whole-genome amplification. Proc. Natl. Acad. Sci. USA 105, 15499–15504 (2008).
Lasken, R.S. & Stockwell, T.B. Mechanism of chimera formation during the multiple displacement amplification reaction. BMC Biotechnol. 7, 1–11 (2007).
Fitzsimons, M.S. et al. Nearly finished genomes produced using gel microdroplet culturing reveals substantial intraspecies diversity within the human microbiome. Genome Res. 23, 878–888 (2013).
Seth-Smith, H.M.B. et al. Whole-genome sequences of Chlamydia trachomatis directly from clinical samples without culture. Genome Res. 23, 855–866 (2013).
Blainey, P.C. & Quake, S.R. Digital MDA for enumeration of total nucleic acid contamination. Nucleic Acids Res. 39, e19 (2010).
Woyke, T. et al. Decontamination of MDA reagents for single cell whole genome amplification. PLoS ONE 6, e26161 (2011).
Dichosa, A.E.K. et al. Artificial polyploidy improves bacterial single cell genome recovery. PLoS ONE 7, e37387 (2012).
Woyke, T. et al. One bacterial cell, one complete genome. PLoS ONE 5, e10314 (2010).
Seth-Smith, H.M.B. et al. Generating whole bacterial genome sequences of low-abundance species from complex samples with IMS-MDA. Nat. Protoc. 8, 2404–2412 (2013).
McLean, J.S. et al. Candidate phylum TM6 genome recovered from a hospital sink biofilm provides genomic insights into this uncultivated phylum. Proc. Natl. Acad. Sci. USA 110, E2390–E2399 (2013).
Weaver, J.C., Williams, G.B., Klibanov, A. & Demain, A.L. Gel microdroplets: rapid detection and enumeration of individual microorganisms by their metabolic activity. Nat. Biotechnol. 6, 1084–1089 (1988).
Williams, G.B., Weaver, J.C. & Demain, A.L. Rapid microbial detection and enumeration using gel microdroplets and colorimetric or fluorescence indicator systems. J. Clin. Microbiol. 28, 1002–1008 (1990).
Borth, N., Zeyda, M. & Katinge, H. Efficient selection of high-producing subclones during gene amplification of recombinant Chinese hamster ovary cells by flow cytometry and cell sorting. Biotechnol. Bioeng. 71, 266–273 (2000).
Katsuragi, T., Tanaka, S., Nagahiro, S. & Tani, Y. Gel microdroplet technique leaving microorganisms alive for sorting by flow cytometry. J. Microbiol. Methods 42, 81–86 (2000).
Manome, A. et al. Application of gel microdroplet and flow cytometry techniques to selective enrichment of non-growing bacterial cells. FEMS Microbiol. Lett. 197, 29–33 (2001).
Cox, D.L., Akins, D.R., Porcella, S.F., Norgard, M.V. & Radolf, J.D. Treponema pallidum in gel microdroplets: a novel strategy for investigation of treponemal molecular architecture. Mol. Microbiol. 15, 1151–1164 (1995).
Cox, D.L. et al. Limited surface exposure of Borrelia burgdorferi outer surface lipoproteins. Proc. Natl. Acad. Sci. USA 93, 7973–7978 (1996).
Ryan, C., Nguyen, B.-T. & Sullivan, S.J. Rapid assay for mycobacterial growth and antibiotic susceptibility using gel microdrop encapsulation. J. Clin. Microbiol. 33, 1720–1726 (1995).
Powell, K.T. & Weaver, J.C. Gel microdroplets and flow cytometry: rapid determination of antibody secretion by individual cells within a cell population. Nat. Biotechnol. 8, 333–337 (1990).
Zengler, K. et al. Cultivating the uncultured. Proc. Natl. Acad. Sci. USA 99, 15681–15686 (2002).
One Cell Systems, Inc. Gel microdrop assay systems, GMD bacterial growth assay: experimental guidelines and protocols. Doc. no. 9005. version 004133 (2003).
Edwards, U., Rogall, T., Blöcker, H., Emde, M. & Böttger, E.C. Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res. 17, 7843–7853 (1989).
Turner, S., Pryer, K.M., Miao, V.P.W. & Palmer, J.D. Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. J. Eukaryotic Microbiol. 46, 327–338 (1999).
Wilson, K.H., Blitchington, R.B. & Greene, R.C. Amplification of bacterial 16S ribosomal DNA with polymerase chain reaction. J. Clin. Microbiol. 28, 1942–1946 (1990).
Baker, G.C. & Cowan, D.A. 16 S rDNA primers and the unbiased assessment of thermophile diversity. Biochem. Soc. Transac. 32, 218–221 (2004).
Berney, M., Hammes, F. & Bosshard, F. Assessment and interpretation of bacterial viability by using the LIVE/DEAD BacLight kit in combination with flow cytometry. Appl. Environ. Microbiol. 73, 3283–3290 (2007).
Gasol, J.M. & Giorgio, P.A.d. Using flow cytometry for counting natural planktonic bacteria and understanding the structure of planktonic bacterial communities. Scientia Marina 64, 197–224 (2000).
Hammes, F. et al. Flow-cytometric total bacterial cell counts as a descriptive microbiological parameter for drinking water treatment processes. Water Res. 42, 269–277 (2008).
Acknowledgements
This work was supported by Los Alamos National Laboratory through a Directed Research program with project code 20110034DR.
Author information
Authors and Affiliations
Contributions
C.S.H., M.S.F. and A.E.K.D. conceived the GMD-SCG strategies. M.S.F., A.E.K.D., A.R.D. and K.G.R. performed the experiments. The authors analyzed the data. A.E.K.D., A.R.D., K.G.R. and C.S.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Dichosa, A., Daughton, A., Reitenga, K. et al. Capturing and cultivating single bacterial cells in gel microdroplets to obtain near-complete genomes. Nat Protoc 9, 608–621 (2014). https://doi.org/10.1038/nprot.2014.034
Published:
Issue date:
DOI: https://doi.org/10.1038/nprot.2014.034
This article is cited by
-
Double emulsions as a high-throughput enrichment and isolation platform for slower-growing microbes
ISME Communications (2023)
-
High-throughput cultivation based on dilution-to-extinction with catalase supplementation and a case study of cultivating acI bacteria from Lake Soyang
Journal of Microbiology (2020)
-
The trajectory of microbial single-cell sequencing
Nature Methods (2017)