Key Points
-
Protein kinases and proteases are both major classes of cellular enzymes; the kinome and the degradome each include more than 500 genes.
-
Direct interactions between kinases and proteases are frequent, and more than 125 examples of kinases that undergo regulated processing by one or more proteases are currently known. Targets for proteases include kinases in all the major branches of the kinome, as well as atypical protein kinases. Conversely, approximately 150 proteases in all the main families in the degradome are known to be phosphorylated by one or more kinases.
-
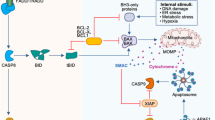
Bidirectional kinase–protease interactions have an important role in many cellular processes, including transmembrane signalling, apoptosis and cell migration. Prominent examples include kinase–caspase crosstalk in apoptosis and interactions between deubiquitylases or γ-secretase and receptor tyrosine kinases in cell proliferation and invasion.
-
Kinase–protease interactions are implicated as causal events in disease, and have a particularly important role in cancer. Cell migration and invasion are hallmarks of metastatic cancer, and many kinase–protease interactions are involved in activating these processes in a dysregulated fashion in cancer cells.
-
Clinical implications of kinase–protease crosstalk are emerging. Interplay between ERBB family receptor tyrosine kinases and a disintegrin and metalloproteinase (ADAM) proteases results in the shedding of ERBB2 soluble forms that blunt the efficiency of ERBB2-specific monoclonal antibody therapy in breast cancer. A clinical trial in patients with breast cancer is underway using a combination of an ADAM10 and ADAM17 inhibitor with an ERBB2-specific antibody.
-
Kinase–protease connections are being uncovered at a rapid pace, and their roles in cancer will undoubtedly become increasingly important and offer new opportunities for clinical intervention.
Abstract
Kinases and proteases are responsible for two fundamental regulatory mechanisms — phosphorylation and proteolysis — that orchestrate the rhythms of life and death in all organisms. Recent studies have highlighted the elaborate interplay between both post-translational regulatory systems. Many intracellular or pericellular proteases are regulated by phosphorylation, whereas multiple kinases are activated or inactivated by proteolytic cleavage. The functional consequences of this regulatory crosstalk are especially relevant in the different stages of cancer progression. What are the clinical implications derived from the fertile dialogue between kinases and proteases in cancer?
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Blume-Jensen, P. & Hunter, T. Oncogenic kinase signalling. Nature 411, 355–365 (2001).
Lopez-Otin, C. & Bond, J. S. Proteases: multifunctional enzymes in life and disease. J. Biol. Chem. 283, 30433–30437 (2008).
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Puente, X. S., Sanchez, L. M., Overall, C. M. & Lopez-Otin, C. Human and mouse proteases: a comparative genomic approach. Nature Rev. Genet. 4, 544–558 (2003).
Quesada, V., Ordonez, G. R., Sanchez, L. M., Puente, X. S. & Lopez-Otin, C. The Degradome database: mammalian proteases and diseases of proteolysis. Nucleic Acids Res. 37, D239–D243 (2009).
Murphy, G. The ADAMs: signalling scissors in the tumour microenvironment. Nature Rev. Cancer 8, 932–941 (2008).
Goll, D. E., Thompson, V. F., Li, H., Wei, W. & Cong, J. The calpain system. Physiol. Rev. 83, 731–801 (2003).
Kuninaka, S. et al. Serine protease Omi/HtrA2 targets WARTS kinase to control cell proliferation. Oncogene 26, 2395–2406 (2007).
Zhang, F., Paterson, A. J., Huang, P., Wang, K. & Kudlow, J. E. Metabolic control of proteasome function. Physiology (Bethesda) 22, 373–379 (2007).
Cursi, S. et al. Src kinase phosphorylates Caspase-8 on Tyr380: a novel mechanism of apoptosis suppression. EMBO J. 25, 1895–1905 (2006).
Gorr, I. H., Boos, D. & Stemmann, O. Mutual inhibition of separase and Cdk1 by two-step complex formation. Mol. Cell 19, 135–141 (2005).
Kurokawa, M. & Kornbluth, S. Caspases and kinases in a death grip. Cell 138, 838–854 (2009).
Sebbagh, M., Hamelin, J., Bertoglio, J., Solary, E. & Breard, J. Direct cleavage of ROCK II by granzyme B induces target cell membrane blebbing in a caspase-independent manner. J. Exp. Med. 201, 465–471 (2005).
Taylor, R. C., Cullen, S. P. & Martin, S. J. Apoptosis: controlled demolition at the cellular level. Nature Rev. Mol. Cell Biol. 9, 231–241 (2008).
Blobel, C. P. ADAMs: key components in EGFR signalling and development. Nature Rev. Mol. Cell Biol. 6, 32–43 (2005).
Selkoe, D. J. & Wolfe, M. S. Presenilin: running with scissors in the membrane. Cell 131, 215–221 (2007).
King, R. W., Deshaies, R. J., Peters, J. M. & Kirschner, M. W. How proteolysis drives the cell cycle. Science 274, 1652–1659 (1996). This is an excellent analysis of the functional relevance of proteolysis in the regulation of cell cycle progression.
Ciechanover, A. Proteolysis: from the lysosome to ubiquitin and the proteasome. Nature Rev. Mol. Cell Biol. 6, 79–87 (2005). An outstanding chronicle of the history of regulated proteolysis.
Lu, Z. & Hunter, T. Degradation of activated protein kinases by ubiquitination. Annu. Rev. Biochem. 78, 435–475 (2009).
Mosesson, Y., Mills, G. B. & Yarden, Y. Derailed endocytosis: an emerging feature of cancer. Nature Rev. Cancer 8, 835–850 (2008).
Dephoure, N. et al. A quantitative atlas of mitotic phosphorylation. Proc. Natl Acad. Sci. USA 105, 10762–10767 (2008). One of several key papers demonstrating the feasibility of global analysis of phosphoproteomes.
Wei, N., Serino, G. & Deng, X. W. The COP9 signalosome: more than a protease. Trends Biochem. Sci. 33, 592–600 (2008).
Tomoda, K. et al. The Jab1/COP9 signalosome subcomplex is a downstream mediator of Bcr-Abl kinase activity and facilitates cell-cycle progression. Blood 105, 775–783 (2005).
Richardson, K. S. & Zundel, W. The emerging role of the COP9 signalosome in cancer. Mol. Cancer Res. 3, 645–653 (2005).
Adler, A. S. et al. CSN5 isopeptidase activity links COP9 signalosome activation to breast cancer progression. Cancer Res. 68, 506–515 (2008).
Komander, D., Clague, M. J. & Urbe, S. Breaking the chains: structure and function of the deubiquitinases. Nature Rev. Mol. Cell Biol. 10, 550–563 (2009).
Niendorf, S. et al. Essential role of ubiquitin-specific protease 8 for receptor tyrosine kinase stability and endocytic trafficking in vivo. Mol. Cell. Biol. 27, 5029–5039 (2007).
Alwan, H. A. & van Leeuwen, J. E. UBPY-mediated epidermal growth factor receptor (EGFR) de-ubiquitination promotes EGFR degradation. J. Biol. Chem. 282, 1658–1669 (2007).
Janssen, J. W., Schleithoff, L., Bartram, C. R. & Schulz, A. S. An oncogenic fusion product of the phosphatidylinositol 3-kinase p85β subunit and HUMORF8, a putative deubiquitinating enzyme. Oncogene 16, 1767–1772 (1998).
McCullough, J., Clague, M. J. & Urbe, S. AMSH is an endosome-associated ubiquitin isopeptidase. J. Cell Biol. 166, 487–492 (2004).
Duex, J. E. & Sorkin, A. RNA interference screen identifies Usp18 as a regulator of epidermal growth factor receptor synthesis. Mol. Biol. Cell 20, 1833–1844 (2009).
Zhao, B., Schlesiger, C., Masucci, M. G. & Lindsten, K. The ubiquitin specific protease 4 (Usp4) is a new player in the Wnt signalling pathway. J. Cell. Mol. Med. 13, 1886–1895 (2009).
Al-Hakim, A. K. et al. Control of AMPK-related kinases by USP9X and atypical Lys29/Lys33-linked polyubiquitin chains. Biochem. J. 411, 249–260 (2008).
Zhang, D., Zaugg, K., Mak, T. W. & Elledge, S. J. A role for the deubiquitinating enzyme USP28 in control of the DNA-damage response. Cell 126, 529–542 (2006). An excellent example of functional interplay between DUBs and kinases in response to DNA damage.
Yamaguchi, T., Kimura, J., Miki, Y. & Yoshida, K. The deubiquitinating enzyme USP11 controls an IκB kinase α (IKKα)-p53 signaling pathway in response to tumor necrosis factor α (TNFα). J. Biol. Chem. 282, 33943–33948 (2007).
van Leuken, R. J., Luna-Vargas, M. P., Sixma, T. K., Wolthuis, R. M. & Medema, R. H. Usp39 is essential for mitotic spindle checkpoint integrity and controls mRNA-levels of aurora B. Cell Cycle 7, 2710–2719 (2008).
Sun, S. C. CYLD: a tumor suppressor deubiquitinase regulating NF-κB activation and diverse biological processes. Cell Death Differ. 17, 25–34 (2010).
Stegmeier, F. et al. The tumor suppressor CYLD regulates entry into mitosis. Proc. Natl Acad. Sci. USA 104, 8869–8874 (2007).
Fernandez-Montalvan, A. et al. Biochemical characterization of USP7 reveals post-translational modification sites and structural requirements for substrate processing and subcellular localization. FEBS J. 274, 4256–4270 (2007).
Joo, H. Y. et al. Regulation of cell cycle progression and gene expression by H2A deubiquitination. Nature 449, 1068–1072 (2007).
Stegmeier, F. et al. Anaphase initiation is regulated by antagonistic ubiquitination and deubiquitination activities. Nature 446, 876–881 (2007).
Matsuoka, S. et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 316, 1160–1166 (2007).
Mu, J. J. et al. A proteomic analysis of ataxia telangiectasia-mutated (ATM)/ATM-Rad3-related (ATR) substrates identifies the ubiquitin-proteasome system as a regulator for DNA damage checkpoints. J. Biol. Chem. 282, 17330–17334 (2007).
Hutti, J. E. et al. IκB kinaseβ phosphorylates the K63 deubiquitinase A20 to cause feedback inhibition of the NF-κB pathway. Mol. Cell. Biol. 27, 7451–7461 (2007).
Hutti, J. E. et al. Phosphorylation of the tumor suppressor CYLD by the breast cancer oncogene IKKɛ promotes cell transformation. Mol. Cell 34, 461–472 (2009).
Reiley, W., Zhang, M., Wu, X., Granger, E. & Sun, S. C. Regulation of the deubiquitinating enzyme CYLD by IκB kinase γ-dependent phosphorylation. Mol. Cell. Biol. 25, 3886–3895 (2005).
Mizuno, E., Kitamura, N. & Komada, M. 14-3-3-dependent inhibition of the deubiquitinating activity of UBPY and its cancellation in the M phase. Exp. Cell Res. 313, 3624–3634 (2007).
Cao, Z., Wu, X., Yen, L., Sweeney, C. & Carraway, K. L. Neuregulin-induced ErbB3 downregulation is mediated by a protein stability cascade involving the E3 ubiquitin ligase Nrdp1. Mol. Cell. Biol. 27, 2180–2188 (2007).
Olsen, J. V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648 (2006).
Diella, F., Gould, C. M., Chica, C., Via, A. & Gibson, T. J. Phospho.ELM: a database of phosphorylation sites — update 2008. Nucleic Acids Res. 36, D240–D244 (2008).
Gnad, F. et al. PHOSIDA (phosphorylation site database): management, structural and evolutionary investigation, and prediction of phosphosites. Genome Biol. 8, R250 (2007).
Sun, Y. et al. Separase is recruited to mitotic chromosomes to dissolve sister chromatid cohesion in a DNA-dependent manner. Cell 137, 123–132 (2009).
Zou, H., McGarry, T. J., Bernal, T. & Kirschner, M. W. Identification of a vertebrate sister-chromatid separation inhibitor involved in transformation and tumorigenesis. Science 285, 418–422 (1999).
Stemmann, O., Zou, H., Gerber, S. A., Gygi, S. P. & Kirschner, M. W. Dual inhibition of sister chromatid separation at metaphase. Cell 107, 715–726 (2001).
Zhang, N. et al. Overexpression of Separase induces aneuploidy and mammary tumorigenesis. Proc. Natl Acad. Sci. USA 105, 13033–13038 (2008).
Meyer, R. et al. Overexpression and mislocalization of the chromosomal segregation protein separase in multiple human cancers. Clin. Cancer Res. 15, 2703–2710 (2009).
Barila, D. et al. Caspase-dependent cleavage of c-Abl contributes to apoptosis. Mol. Cell. Biol. 23, 2790–2799 (2003).
Matsuura, K., Wakasugi, M., Yamashita, K. & Matsunaga, T. Cleavage-mediated activation of Chk1 during apoptosis. J. Biol. Chem. 283, 25485–25491 (2008).
Fischer, U., Stroh, C. & Schulze-Osthoff, K. Unique and overlapping substrate specificities of caspase-8 and caspase-10. Oncogene 25, 152–159 (2006).
Coleman, M. L. et al. Membrane blebbing during apoptosis results from caspase-mediated activation of ROCK I. Nature Cell Biol. 3, 339–345 (2001).
Sebbagh, M. et al. Caspase-3-mediated cleavage of ROCK I induces MLC phosphorylation and apoptotic membrane blebbing. Nature Cell Biol. 3, 346–352 (2001).
Cheung, W. L. et al. Apoptotic phosphorylation of histone H2B is mediated by mammalian sterile twenty kinase. Cell 113, 507–517 (2003). References 60–62 describe how caspase-mediated cleavage of certain kinases contributes to morphological changes that are characteristic of apoptotic cells.
Arnold, R. et al. Sustained JNK signaling by proteolytically processed HPK1 mediates IL-3 independent survival during monocytic differentiation. Cell Death Differ. 14, 568–575 (2007).
Brenner, D. et al. Caspase-cleaved HPK1 induces CD95L-independent activation-induced cell death in T and B lymphocytes. Blood 110, 3968–3977 (2007).
Bachelder, R. E. et al. p53 inhibits α6β4 integrin survival signaling by promoting the caspase 3-dependent cleavage of AKT/PKB. J. Cell Biol. 147, 1063–1072 (1999).
Wen, L. P. et al. Cleavage of focal adhesion kinase by caspases during apoptosis. J. Biol. Chem. 272, 26056–26061 (1997).
Baek, K. H. et al. Caspases-dependent cleavage of mitotic checkpoint proteins in response to microtubule inhibitor. Oncol. Res. 15, 161–168 (2005).
McGinnis, K. M., Whitton, M. M., Gnegy, M. E. & Wang, K. K. Calcium/calmodulin-dependent protein kinase IV is cleaved by caspase-3 and calpain in SH-SY5Y human neuroblastoma cells undergoing apoptosis. J. Biol. Chem. 273, 19993–20000 (1998).
Morley, S. J., Jeffrey, I., Bushell, M., Pain, V. M. & Clemens, M. J. Differential requirements for caspase-8 activity in the mechanism of phosphorylation of eIF2α, cleavage of eIF4GI and signaling events associated with the inhibition of protein synthesis in apoptotic Jurkat T cells. FEBS Lett. 477, 229–236 (2000).
Smith, G. C., d' Adda di Fagagna, F., Lakin, N. D. & Jackson, S. P. Cleavage and inactivation of ATM during apoptosis. Mol. Cell. Biol. 19, 6076–6084 (1999).
Han, Z. et al. DNA-dependent protein kinase is a target for a CPP32-like apoptotic protease. J. Biol. Chem. 271, 25035–25040 (1996).
Ancot, F., Foveau, B., Lefebvre, J., Leroy, C. & Tulasne, D. Proteolytic cleavages give receptor tyrosine kinases the gift of ubiquity. Oncogene 28, 2185–2195 (2009).
Strohecker, A. M., Yehiely, F., Chen, F. & Cryns, V. L. Caspase cleavage of HER-2 releases a Bad-like cell death effector. J. Biol. Chem. 283, 18269–18282 (2008).
Cardone, M. H. et al. Regulation of cell death protease caspase-9 by phosphorylation. Science 282, 1318–1321 (1998). This was the first experimental demonstration that caspases are directly regulated by phosphorylation.
Allan, L. A. et al. Inhibition of caspase-9 through phosphorylation at Thr25 by ERK MAPK. Nature Cell Biol. 5, 647–654 (2003).
Laguna, A. et al. The protein kinase DYRK1A regulates caspase-9-mediated apoptosis during retina development. Dev. Cell 15, 841–853 (2008).
Seifert, A. & Clarke, P. R. p38α- and DYRK1A-dependent phosphorylation of caspase-9 at an inhibitory site in response to hyperosmotic stress. Cell Signal. 21, 1626–1633 (2009).
McDonnell, M. A. et al. Phosphorylation of murine caspase-9 by the protein kinase casein kinase 2 regulates its cleavage by caspase-8. J. Biol. Chem. 283, 20149–20158 (2008).
Martin, M. C. et al. Protein kinase A regulates caspase-9 activation by Apaf-1 downstream of cytochrome c. J. Biol. Chem. 280, 15449–15455 (2005).
Allan, L. A. & Clarke, P. R. Phosphorylation of caspase-9 by CDK1/cyclin B1 protects mitotic cells against apoptosis. Mol. Cell 26, 301–310 (2007).
Senft, J., Helfer, B. & Frisch, S. M. Caspase-8 interacts with the p85 subunit of phosphatidylinositol 3-kinase to regulate cell adhesion and motility. Cancer Res. 67, 11505–11509 (2007).
Jia, S. H., Parodo, J., Kapus, A., Rotstein, O. D. & Marshall, J. C. Dynamic regulation of neutrophil survival through tyrosine phosphorylation or dephosphorylation of caspase-8. J. Biol. Chem. 283, 5402–5413 (2008).
Alvarado-Kristensson, M. et al. p38-MAPK signals survival by phosphorylation of caspase-8 and caspase-3 in human neutrophils. J. Exp. Med. 199, 449–458 (2004).
Thome, M. et al. Identification of CARDIAK, a RIP-like kinase that associates with caspase-1. Curr. Biol. 8, 885–888 (1998).
Voss, O. H., Kim, S., Wewers, M. D. & Doseff, A. I. Regulation of monocyte apoptosis by the protein kinase Cδ-dependent phosphorylation of caspase-3. J. Biol. Chem. 280, 17371–17379 (2005).
Raina, D. et al. c-Abl tyrosine kinase regulates caspase-9 autocleavage in the apoptotic response to DNA damage. J. Biol. Chem. 280, 11147–11151 (2005).
Shi, M. et al. DNA-PKcs-PIDDosome: a nuclear caspase-2-activating complex with role in G2/M checkpoint maintenance. Cell 136, 508–520 (2009). This paper identifies a new kinase–protease complex that is implicated in the cellular response to DNA damage.
Carragher, N. O., Fincham, V. J., Riley, D. & Frame, M. C. Cleavage of focal adhesion kinase by different proteases during SRC-regulated transformation and apoptosis. Distinct roles for calpain and caspases. J. Biol. Chem. 276, 4270–4275 (2001). This is one of several key papers demonstrating that proteases that are distinct from caspases are involved in mutual interactions with kinases during apoptotic processes.
Andrade, F. et al. Granzyme B directly and efficiently cleaves several downstream caspase substrates: implications for CTL-induced apoptosis. Immunity 8, 451–460 (1998).
Loeb, C. R., Harris, J. L. & Craik, C. S. Granzyme B proteolyzes receptors important to proliferation and survival, tipping the balance toward apoptosis. J. Biol. Chem. 281, 28326–28335 (2006).
Gervais, F. G., Thornberry, N. A., Ruffolo, S. C., Nicholson, D. W. & Roy, S. Caspases cleave focal adhesion kinase during apoptosis to generate a FRNK-like polypeptide. J. Biol. Chem. 273, 17102–17108 (1998).
Yang, L. et al. Akt attenuation of the serine protease activity of HtrA2/Omi through phosphorylation of serine 212. J. Biol. Chem. 282, 10981–10987 (2007).
Barbero, S. et al. Caspase-8 association with the focal adhesion complex promotes tumor cell migration and metastasis. Cancer Res. 69, 3755–3763 (2009).
Xu, L. & Deng, X. Suppression of cancer cell migration and invasion by protein phosphatase 2A through dephosphorylation of mu- and m-calpains. J. Biol. Chem. 281, 35567–35575 (2006). This paper demonstrates that the interaction between kinases and calpains could have important consequences for cancer cell invasion.
Lu, X. et al. ADAMTS1 and MMP1 proteolytically engage EGF-like ligands in an osteolytic signaling cascade for bone metastasis. Genes Dev. 23, 1882–1894 (2009). This is an excellent example of the crosstalk between different metalloproteinases and RTK signalling during tumour invasion and metastasis.
Barbero, S. et al. Identification of a critical tyrosine residue in caspase 8 that promotes cell migration. J. Biol. Chem. 283, 13031–13034 (2008).
Finlay, D., Howes, A. & Vuori, K. Critical role for caspase-8 in epidermal growth factor signaling. Cancer Res. 69, 5023–5029 (2009).
Franco, S. J. & Huttenlocher, A. Regulating cell migration: calpains make the cut. J. Cell Sci. 118, 3829–3838 (2005).
Carragher, N. O. & Frame, M. C. Focal adhesion and actin dynamics: a place where kinases and proteases meet to promote invasion. Trends Cell Biol. 14, 241–249 (2004).
Hajimohammadreza, I. et al. Neuronal nitric oxide synthase and calmodulin-dependent protein kinase IIα undergo neurotoxin-induced proteolysis. J. Neurochem. 69, 1006–1013 (1997).
Kishimoto, A. et al. Limited proteolysis of protein kinase C subspecies by calcium-dependent neutral protease (calpain). J. Biol. Chem. 264, 4088–4092 (1989).
Goni-Oliver, P., Lucas, J. J., Avila, J. & Hernandez, F. N-terminal cleavage of GSK-3 by calpain: a new form of GSK-3 regulation. J. Biol. Chem. 282, 22406–22413 (2007).
Glading, A. et al. Epidermal growth factor activates m-calpain (calpain II), at least in part, by extracellular signal-regulated kinase-mediated phosphorylation. Mol. Cell. Biol. 24, 2499–2512 (2004).
Xu, L. & Deng, X. Protein kinase Cι promotes nicotine-induced migration and invasion of cancer cells via phosphorylation of micro- and m-calpains. J. Biol. Chem. 281, 4457–4466 (2006).
Shiraha, H., Glading, A., Chou, J., Jia, Z. & Wells, A. Activation of m-calpain (calpain II) by epidermal growth factor is limited by protein kinase A phosphorylation of m-calpain. Mol. Cell. Biol. 22, 2716–2727 (2002).
Bai, D. S. et al. Capn4 overexpression underlies tumor invasion and metastasis after liver transplantation for hepatocellular carcinoma. Hepatology 49, 460–470 (2009).
Libertini, S. J. et al. Cyclin E both regulates and is regulated by calpain 2, a protease associated with metastatic breast cancer phenotype. Cancer Res. 65, 10700–10708 (2005).
Thomas, G. Furin at the cutting edge: from protein traffic to embryogenesis and disease. Nature Rev. Mol. Cell Biol. 3, 753–766 (2002).
Scamuffa, N. et al. Selective inhibition of proprotein convertases represses the metastatic potential of human colorectal tumor cells. J. Clin. Invest. 118, 352–363 (2008). This is an excellent example of the relevance of furin–kinase interactions for tumour cell metastasis.
Christianson, T. A. et al. NH2-terminally truncated HER-2/neu protein: relationship with shedding of the extracellular domain and with prognostic factors in breast cancer. Cancer Res. 58, 5123–5129 (1998).
Steiner, H., Fluhrer, R. & Haass, C. Intramembrane proteolysis by γ-secretase. J. Biol. Chem. 283, 29627–29631 (2008).
Ni, C. Y., Murphy, M. P., Golde, T. E. & Carpenter, G. γ-secretase cleavage and nuclear localization of ErbB-4 receptor tyrosine kinase. Science 294, 2179–2181 (2001). This paper reports the discovery that RTKs undergo RIPping events mediated by γ-secretase.
Naresh, A. et al. The ERBB4/HER4 intracellular domain 4ICD is a BH3-only protein promoting apoptosis of breast cancer cells. Cancer Res. 66, 6412–6420 (2006).
McElroy, B., Powell, J. C. & McCarthy, J. V. The insulin-like growth factor 1 (IGF-1) receptor is a substrate for gamma-secretase-mediated intramembrane proteolysis. Biochem. Biophys. Res. Commun. 358, 1136–1141 (2007).
Kasuga, K., Kaneko, H., Nishizawa, M., Onodera, O. & Ikeuchi, T. Generation of intracellular domain of insulin receptor tyrosine kinase by gamma-secretase. Biochem. Biophys. Res. Commun. 360, 90–96 (2007).
Levi, E. et al. Matrix metalloproteinase 2 releases active soluble ectodomain of fibroblast growth factor receptor 1. Proc. Natl Acad. Sci. USA 93, 7069–7074 (1996).
Lin, K. T., Sloniowski, S., Ethell, D. W. & Ethell, I. M. Ephrin-B2-induced cleavage of EphB2 receptor is mediated by matrix metalloproteinases to trigger cell repulsion. J. Biol. Chem. 283, 28969–28979 (2008).
Butler, G. S., Dean, R. A., Tam, E. M. & Overall, C. M. Pharmacoproteomics of a metalloproteinase hydroxamate inhibitor in breast cancer cells: dynamics of membrane type 1 matrix metalloproteinase-mediated membrane protein shedding. Mol. Cell. Biol. 28, 4896–4914 (2008).
Darragh, M. R., Bhatt, A. S. & Craik, C. S. MT-SP1 proteolysis and regulation of cell-microenvironment interactions. Front. Biosci. 13, 528–539 (2008).
Chen, M., Chen, L. M., Lin, C. Y. & Chai, K. X. The epidermal growth factor receptor (EGFR) is proteolytically modified by the Matriptase-Prostasin serine protease cascade in cultured epithelial cells. Biochim. Biophys. Acta 1783, 896–903 (2008).
Izumi, Y. et al. A metalloprotease-disintegrin, MDC9/meltrin-γ/ADAM9 and PKCδ are involved in TPA-induced ectodomain shedding of membrane-anchored heparin-binding EGF-like growth factor. EMBO J. 17, 7260–7272 (1998).
Janes, P. W. et al. Cytoplasmic relaxation of active Eph controls Ephrin shedding by ADAM10. PLoS Biol. 7, e1000215 (2009). This paper identifies a new molecular mechanism linking kinase and sheddase activities.
Stautz, D. et al. ADAM12 localizes with c-Src to actin-rich structures at the cell periphery and regulates Src kinase activity. Exp. Cell Res. 316, 55–67 (2010).
Poghosyan, Z. et al. Phosphorylation-dependent interactions between ADAM15 cytoplasmic domain and Src family protein-tyrosine kinases. J. Biol. Chem. 277, 4999–5007 (2002).
Maretzky, T. et al. Src stimulates fibroblast growth factor receptor-2 shedding by an ADAM15 splice variant linked to breast cancer. Cancer Res. 69, 4573–4576 (2009).
Diaz-Rodriguez, E., Montero, J. C., Esparis-Ogando, A., Yuste, L. & Pandiella, A. Extracellular signal-regulated kinase phosphorylates tumor necrosis factor α-converting enzyme at threonine 735: a potential role in regulated shedding. Mol. Biol. Cell 13, 2031–2044 (2002).
Sariahmetoglu, M. et al. Regulation of matrix metalloproteinase-2 (MMP-2) activity by phosphorylation. FASEB J. 21, 2486–2495 (2007).
Nyalendo, C. et al. Src-dependent phosphorylation of membrane type I matrix metalloproteinase on cytoplasmic tyrosine 573: role in endothelial and tumor cell migration. J. Biol. Chem. 282, 15690–15699 (2007).
Moss, N. M. et al. Epidermal growth factor receptor-mediated membrane type 1 matrix metalloproteinase endocytosis regulates the transition between invasive versus expansive growth of ovarian carcinoma cells in three-dimensional collagen. Mol. Cancer Res. 7, 809–820 (2009).
Uemura, K. et al. GSK3β activity modifies the localization and function of presenilin 1. J. Biol. Chem. 282, 15823–15832 (2007).
Prager, K. et al. A structural switch of presenilin 1 by glycogen synthase kinase 3β-mediated phosphorylation regulates the interaction with β-catenin and its nuclear signaling. J. Biol. Chem. 282, 14083–14093 (2007).
Kuo, L. H. et al. Tumor necrosis factor-α-elicited stimulation of γ-secretase is mediated by c-Jun N-terminal kinase-dependent phosphorylation of presenilin and nicastrin. Mol. Biol. Cell 19, 4201–4212 (2008).
Walter, J., Schindzielorz, A., Grunberg, J. & Haass, C. Phosphorylation of presenilin-2 regulates its cleavage by caspases and retards progression of apoptosis. Proc. Natl Acad. Sci. USA 96, 1391–1396 (1999).
Fluhrer, R., Friedlein, A., Haass, C. & Walter, J. Phosphorylation of presenilin 1 at the caspase recognition site regulates its proteolytic processing and the progression of apoptosis. J. Biol. Chem. 279, 1585–1593 (2004).
Foveau, B. et al. Down-regulation of the met receptor tyrosine kinase by presenilin-dependent regulated intramembrane proteolysis. Mol. Biol. Cell 20, 2495–2507 (2009).
Swendeman, S. et al. VEGF-A stimulates ADAM17-dependent shedding of VEGFR2 and crosstalk between VEGFR2 and ERK signaling. Circ. Res. 103, 916–918 (2008).
Baselga, J. & Swain, S. M. Novel anticancer targets: revisiting ERBB2 and discovering ERBB3. Nature Rev. Cancer 9, 463–475 (2009).
Zhang, J., Yang, P. L. & Gray, N. S. Targeting cancer with small molecule kinase inhibitors. Nature Rev. Cancer 9, 28–39 (2009). References 137 and 138 review the current status and recent developments of strategies for targeting kinases in cancer.
Orlowski, R. Z. & Kuhn, D. J. Proteasome inhibitors in cancer therapy: lessons from the first decade. Clin. Cancer Res. 14, 1649–1657 (2008).
Overall, C. M. & Lopez-Otin, C. Strategies for MMP inhibition in cancer: innovations for the post-trial era. Nature Rev. Cancer 2, 657–672 (2002).
Borgono, C. A. & Diamandis, E. P. The emerging roles of human tissue kallikreins in cancer. Nature Rev. Cancer 4, 876–890 (2004).
Palermo, C. & Joyce, J. A. Cysteine cathepsin proteases as pharmacological targets in cancer. Trends Pharmacol. Sci. 29, 22–28 (2008).
Lopez-Otin, C. & Matrisian, L. M. Emerging roles of proteases in tumour suppression. Nature Rev. Cancer 7, 800–808 (2007).
Scaltriti, M. et al. Expression of p95HER2, a truncated form of the HER2 receptor, and response to anti-HER2 therapies in breast cancer. J. Natl Cancer Inst. 99, 628–638 (2007).
Zhou, B. B. et al. Targeting ADAM-mediated ligand cleavage to inhibit HER3 and EGFR pathways in non-small cell lung cancer. Cancer Cell 10, 39–50 (2006).
Witters, L. et al. Synergistic inhibition with a dual epidermal growth factor receptor/HER-2/neu tyrosine kinase inhibitor and a disintegrin and metalloprotease inhibitor. Cancer Res. 68, 7083–7089 (2008). One of several key papers demonstrating the feasibility of anticancer therapies based on the combined inhibition of kinases and metalloproteinases.
Merchant, N. B. et al. TACE/ADAM-17: a component of the epidermal growth factor receptor axis and a promising therapeutic target in colorectal cancer. Clin. Cancer Res. 14, 1182–1191 (2008).
Guaiquil, V. et al. ADAM9 is involved in pathological retinal neovascularization. Mol. Cell. Biol. 29, 2694–2703 (2009).
Hussain, S., Zhang, Y. & Galardy, P. J. DUBs and cancer: the role of deubiquitinating enzymes as oncogenes, non-oncogenes and tumor suppressors. Cell Cycle 8, 1688–1697 (2009).
An, J. & Rettig, M. B. Epidermal growth factor receptor inhibition sensitizes renal cell carcinoma cells to the cytotoxic effects of bortezomib. Mol. Cancer Ther. 6, 61–69 (2007).
Lorch, J. H., Thomas, T. O. & Schmoll, H. J. Bortezomib inhibits cell-cell adhesion and cell migration and enhances epidermal growth factor receptor inhibitor-induced cell death in squamous cell cancer. Cancer Res. 67, 727–734 (2007).
Cascone, T. et al. Synergistic anti-proliferative and pro-apoptotic activity of combined therapy with bortezomib, a proteasome inhibitor, with anti-epidermal growth factor receptor (EGFR) drugs in human cancer cells. J. Cell. Physiol. 216, 698–707 (2008).
Ayala, G. et al. Bortezomib-mediated inhibition of steroid receptor coactivator-3 degradation leads to activated Akt. Clin. Cancer Res. 14, 7511–7518 (2008).
Chen, K. F. et al. Down-regulation of phospho-Akt is a major molecular determinant of bortezomib-induced apoptosis in hepatocellular carcinoma cells. Cancer Res. 68, 6698–6707 (2008).
Sloss, C. M. et al. Proteasome inhibition activates epidermal growth factor receptor (EGFR) and EGFR-independent mitogenic kinase signaling pathways in pancreatic cancer cells. Clin. Cancer Res. 14, 5116–5123 (2008).
Navas, T. A. et al. Inhibition of p38α MAPK enhances proteasome inhibitor-induced apoptosis of myeloma cells by modulating Hsp27, Bcl-XL, Mcl-1 and p53 levels in vitro and inhibits tumor growth in vivo. Leukemia 20, 1017–1027 (2006). References 150–156 describe how the combination of proteasome and RTK inhibitors can be effective in some tumour types.
Osipo, C. et al. ErbB-2 inhibition activates Notch-1 and sensitizes breast cancer cells to a γ-secretase inhibitor. Oncogene 27, 5019–5032 (2008).
Palomero, T. et al. Mutational loss of PTEN induces resistance to NOTCH1 inhibition in T-cell leukemia. Nature Med. 13, 1203–1210 (2007).
Foster, F. M. et al. Targeting inhibitor of apoptosis proteins in combination with ErbB antagonists in breast cancer. Breast Cancer Res. 11, R41 (2009).
Ziegler, D. S. et al. Resistance of human glioblastoma multiforme cells to growth factor inhibitors is overcome by blockade of inhibitor of apoptosis proteins. J. Clin. Invest. 118, 3109–3122 (2008).
Fischer, E. H. & Krebs, E. G. Conversion of phosphorylase b to phosphorylase a in muscle extracts. J. Biol. Chem. 216, 121–132 (1955). This was the first observation of the need for proteolysis in kinase activation.
Johnson, S. A. & Hunter, T. Kinomics: methods for deciphering the kinome. Nature Methods 2, 17–25 (2005).
Overall, C. M. & Blobel, C. P. In search of partners: linking extracellular proteases to substrates. Nature Rev. Mol. Cell Biol. 8, 245–257 (2007).
Lopez-Otin, C. & Overall, C. M. Protease degradomics: a new challenge for proteomics. Nature Rev. Mol. Cell Biol. 3, 509–519 (2002).
Alvarado-Kristensson, M. & Andersson, T. Protein phosphatase 2A regulates apoptosis in neutrophils by dephosphorylating both p38 MAPK and its substrate caspase 3. J. Biol. Chem. 280, 6238–6244 (2005).
Dessauge, F. et al. Identification of PP1α as a caspase-9 regulator in IL-2 deprivation-induced apoptosis. J. Immunol. 177, 2441–2451 (2006).
Mazars, A. et al. A caspase-dependent cleavage of CDC25A generates an active fragment activating cyclin-dependent kinase 2 during apoptosis. Cell Death Differ. 16, 208–218 (2009).
Torres, J. et al. Phosphorylation-regulated cleavage of the tumor suppressor PTEN by caspase-3: implications for the control of protein stability and PTEN-protein interactions. J. Biol. Chem. 278, 30652–30660 (2003).
Trumpler, A., Schlott, B., Herrlich, P., Greer, P. A. & Bohmer, F. D. Calpain-mediated degradation of reversibly oxidized protein-tyrosine phosphatase 1B. FEBS J. 276, 5622–5633 (2009).
Anders, L. et al. Furin-, ADAM 10-, and γ-secretase-mediated cleavage of a receptor tyrosine phosphatase and regulation of β-catenin's transcriptional activity. Mol. Cell. Biol. 26, 3917–3934 (2006).
Burgoyne, A. M. et al. Proteolytic cleavage of protein tyrosine phosphatase mu regulates glioblastoma cell migration. Cancer Res. 69, 6960–6968 (2009).
Queralt, E., Lehane, C., Novak, B. & Uhlmann, F. Downregulation of PP2A(Cdc55) phosphatase by separase initiates mitotic exit in budding yeast. Cell 125, 719–732 (2006).
Song, M. S. et al. The deubiquitinylation and localization of PTEN are regulated by a HAUSP-PML network. Nature 455, 813–817 (2008).
Acknowledgements
We thank all members of our laboratories for their helpful comments on the manuscript and apologize for the omission of relevant works owing to space constraints. C.L-O. is supported by grants from Ministerio de Ciencia e Innovación-Spain, European Union-Microenvimet and Fundación M. Botín. T.H. is supported by grants from the National Cancer Institute and is a Frank and Else Schilling American Cancer Society Research Professor. The Instituto Universitario de Oncología is supported by Obra Social Cajastur and Acción Transversal del Cáncer RTICC.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information S1 (table)
Proteases phosphorylated by kinases (PDF 312 kb)
Supplementary information S2 (table)
Kinases targeted by proteases (PDF 525 kb)
Supplementary information S3 (figure)
Global representation of the crosstalk between the human kinome and the human degradome. (PDF 166 kb)
Related links
Glossary
- Proteasome
-
Multimeric and multicatalytic protease that degrades multiple intracellular proteins in a ubiquitin- and ATP-dependent manner and has a key role in many events of cellular regulation mediated by protein degradation.
- Shedding
-
Protease-mediated process that involves the release of the extracellular portion of a membrane protein.
- RIPping
-
Protease-mediated process in which a receptor is cleaved in the transmembrane domain by intramembrane-cleaving proteases (I-CLiPs), a group of proteases including the presenilins that are part of a larger complex called γ-secretase.
- Signalosome
-
Multi-subunit protease that promotes the activity of cullin-containing ubiquitin E3 ligases and has important roles in the cell cycle, DNA damage response, nuclear transport, gene transcription and kinase signalling.
- Deubiquitylases
-
A large family of enzymes that comprises cysteine proteases and metalloproteinases, which remove conjugated ubiquitin from proteins and are linked to the regulation and termination of ubiquitin-mediated signalling.
- E3 ligase
-
Enzyme that catalyses the final step of the process that activates and transfers ubiquitin and ubiquitin-like molecules to substrate proteins.
- Isopeptidase
-
Enzyme that cleaves amide bonds that are not present in the main polypeptide chain of a protein, such as those formed between the ɛ-amino group of lysine side chains of ubiquitin-targeted proteins and the terminal carboxyl group of ubiquitin.
- Dependence receptors
-
Structurally unrelated receptors that can induce cell death by apoptosis when unoccupied by ligand, and so create cellular dependence on their ligands. In the presence of ligand, these receptors mediate survival, differentiation and migration.
- Apoptosome
-
Multiprotein complex that mediates the activation of initiator caspases at the onset of apoptosis.
- Furin
-
Member of a group of propotein convertases that catalyse the maturation of precursors for many secreted and transmembrane proteins by cleavage at Arg-X-Lys/Arg-Arg sites.
- Trastuzumab
-
Humanized monoclonal antibody that interferes with ERBB2 signalling and is currently used for the treatment of breast cancer.
Rights and permissions
About this article
Cite this article
López-Otín, C., Hunter, T. The regulatory crosstalk between kinases and proteases in cancer. Nat Rev Cancer 10, 278–292 (2010). https://doi.org/10.1038/nrc2823
Published:
Issue date:
DOI: https://doi.org/10.1038/nrc2823
This article is cited by
-
Phosphorylation-dependent deubiquitinase OTUD3 regulates YY1 stability and promotes colorectal cancer progression
Cell Death & Disease (2024)
-
Enzyme-catalyzed synthesis of selenium-doped manganese phosphate for synergistic therapy of drug-resistant colorectal cancer
Journal of Nanobiotechnology (2023)
-
Phosphorylation of USP27X by GSK3β maintains the stability and oncogenic functions of CBX2
Cell Death & Disease (2023)
-
Community assessment to advance computational prediction of cancer drug combinations in a pharmacogenomic screen
Nature Communications (2019)
-
Phosphorylation by protein kinase A disassembles the caspase-9 core
Cell Death & Differentiation (2018)