Abstract
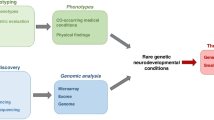
The role of genetic testing in the domain of neurodevelopmental and psychiatric disorders (NPDs) is gradually changing from providing etiological explanation for the presence of NPD phenotypes to also identifying young individuals at high risk of developing NPDs before their clinical manifestation. In clinical practice, the latter implies a shift towards the availability of individual genetic information predicting a certain liability to develop an NPD (e.g., autism, intellectual disability, psychosis etc.). The shift from mostly a posteriori explanation to increasingly a priori risk prediction is the by-product of the systematic implementation of whole exome or genome sequencing as part of routine diagnostic work-ups during the neonatal and prenatal periods. This rapid uptake of genetic testing early in development has far-reaching consequences for psychiatry: Whereas until recently individuals would come to medical attention because of signs of abnormal developmental and/or behavioral symptoms, increasingly, individuals are presented based on genetic liability for NPD outcomes before NPD symptoms emerge. This novel clinical scenario, while challenging, also creates opportunities for research on prevention interventions and precision medicine approaches. Here, we review why optimization of individual risk prediction is a key prerequisite for precision medicine in the sphere of NPDs, as well as the technological and statistical methods required to achieve this ambition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Carmignac V, Nambot S, Lehalle D, Callier P, Moortgat S, Benoit V, et al. Further delineation of the female phenotype with KDM5C disease causing variants: 19 new individuals and review of the literature. Clin Genet. 2020;98:43–55.
Vorstman JAS, Parr JR, Moreno-De-Luca D, Anney RJL, Nurnberger JI Jr, Hallmayer JF. Autism genetics: opportunities and challenges for clinical translation. Nat Rev Genet. 2017;18:362–76.
Trubetskoy V, Pardinas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Plomin R, von Stumm S. The new genetics of intelligence. Nat Rev Genet. 2018;19:148–59.
Yengo L, Vedantam S, Marouli E, Sidorenko J, Bartell E, Sakaue S, et al. A saturated map of common genetic variants associated with human height. Nature. 2022;610:704–12.
Brainstorm C, Anttila V, Bulik-Sullivan B, Finucane HK, Walters RK, Bras J, et al. Analysis of shared heritability in common disorders of the brain. Science. 2018;360:eaap8757.
Callahan KP, Radack J, Wojcik MH, Jenkins SM, Nye RT, Skraban C, et al. Hospital-level variation in genetic testing in children’s hospitals neonatal intensive care units from 2016 to 2021. Genet Med. 2023;25:100357.
Baribeau DA, Hoang N, Selvanayagam T, Stavropoulos DJ, Costain G, Scherer SW, et al. Developmental implications of genetic testing for physical indications. Eur J Hum Genet EJHG. 2022;30:1297–1300.
Summers J, Baribeau D, Perlman P, Hoang N, Cui S, Krakowski A, et al. An integrated clinical approach to children at genetic risk for neurodevelopmental and psychiatric conditions: interdisciplinary collaboration and research infrastructure. J Neurodev Disord. 2024;16:37.
McDonald-McGinn DM, Sullivan KE, Marino B, Philip N, Swillen A, Vorstman JA, et al. 22q11.2 deletion syndrome. Nat Rev Dis Primers. 2015;1:15071.
Schneider M, Debbane M, Bassett AS, Chow EW, Fung WL, van den Bree M, et al. Psychiatric disorders from childhood to adulthood in 22q11.2 deletion syndrome: results from the international consortium on brain and behavior in 22q11.2 deletion syndrome. Am J Psychiatry. 2014;171:627–39.
Huguet G, Schramm C, Douard E, Jiang L, Labbe A, Tihy F, et al. Measuring and estimating the effect sizes of copy number variants on general intelligence in community-based samples. JAMA Psychiatry. 2018;75:447–57.
Moreno-De-Luca A, Myers SM, Challman TD, Moreno-De-Luca D, Evans DW, Ledbetter DH. Developmental brain dysfunction: revival and expansion of old concepts based on new genetic evidence. Lancet Neurol. 2013;12:406–14.
Vorstman JA, Ophoff RA. Genetic causes of developmental disorders. Curr Opin Neurol. 2013;26:128–36.
Fiksinski AM, Hoftman GD, Vorstman JAS, Bearden CE. A genetics-first approach to understanding autism and schizophrenia spectrum disorders: the 22q11.2 deletion syndrome. Mol Psychiatry. 2023;28:341–53.
Fiksinski AM, Schneider M, Zinkstok J, Baribeau D, Chawner S, Vorstman JAS. Neurodevelopmental trajectories and psychiatric morbidity: lessons learned from the 22q11.2 deletion syndrome. Curr Psychiatry Rep. 2021;23:13.
Lee PH, Feng YA, Smoller JW. Pleiotropy and cross-disorder genetics among psychiatric disorders. Biol Psychiatry. 2021;89:20–31.
Moreau CA, Kumar K, Harvey A, Huguet G, Urchs SGW, Schultz LM, et al. Brain functional connectivity mirrors genetic pleiotropy in psychiatric conditions. Brain. 2023;146:1686–96.
Sommer IE, Bearden CE, van Dellen E, Breetvelt EJ, Duijff SN, Maijer K, et al. Early interventions in risk groups for schizophrenia: what are we waiting for? NPJ Schizophr. 2016;2:16003.
Watanabe K, Stringer S, Frei O, Umicevic Mirkov M, de Leeuw C, Polderman TJC, et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat Genet. 2019;51:1339–48.
Negi SK, Guda C. Global gene expression profiling of healthy human brain and its application in studying neurological disorders. Sci Rep. 2017;7:897.
Jacquemont S, Huguet G, Klein M, Chawner S, Donald KA, van den Bree MBM, et al. Genes to mental health (G2MH): a framework to map the combined effects of rare and common variants on dimensions of cognition and psychopathology. Am J Psychiatry. 2022;179:189–203.
Chung WK, Berg JS, Botkin JR, Brenner SE, Brosco JP, Brothers KB, et al. Newborn screening for neurodevelopmental diseases: are we there yet? Am J Med Genet C Semin Med Genet. 2022;190:222–30.
Baribeau D, Vorstman J, Anagnostou E. Novel treatments in autism spectrum disorder. Curr Opin Psychiatry. 2022;35:101–10.
Lin BD, Pinzon-Espinosa J, Blouzard E, van der Horst MZ, Okhuijsen-Pfeifer C, van Eijk KR, et al. Associations between polygenic risk score loading, psychosis liability, and clozapine use among individuals with schizophrenia. JAMA Psychiatry. 2023;80:181–5.
Rolland T, Cliquet F, Anney RJL, Moreau C, Traut N, Mathieu A, et al. Phenotypic effects of genetic variants associated with autism. Nat Med. 2023;29:1671–80.
Chung WK, Roberts TP, Sherr EH, Snyder LG, Spiro JE. 16p11.2 deletion syndrome. Curr Opin Genet Dev. 2021;68:49–56.
Mollon J, Almasy L, Jacquemont S, Glahn DC. The contribution of copy number variants to psychiatric symptoms and cognitive ability. Mol Psychiatry. 2023;28:1480–93.
Mulle JG, Gambello MJ, Sanchez Russo R, Murphy MM, Burrell TL, Klaiman C et al. 3q29 Recurrent Deletion. In: Adam MP, Mirzaa GM, Pagon RA, Wallace SE, Bean LJH, Gripp KW et al. (eds). GeneReviews((R)): Seattle (WA), 1993.
McCarthy SE, Makarov V, Kirov G, Addington AM, McClellan J, Yoon S, et al. Microduplications of 16p11.2 are associated with schizophrenia. Nat Genet. 2009;41:1223–7.
Richards C, Jones C, Groves L, Moss J, Oliver C. Prevalence of autism spectrum disorder phenomenology in genetic disorders: a systematic review and meta-analysis. Lancet Psychiatry. 2015;2:909–16.
Grove J, Ripke S, Als TD, Mattheisen M, Walters RK, Won H, et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 2019;51:431–44.
Demontis D, Walters RK, Martin J, Mattheisen M, Als TD, Agerbo E, et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat Genet. 2019;51:63–75.
Sanders SJ, Murtha MT, Gupta AR, Murdoch JD, Raubeson MJ, Willsey AJ, et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature. 2012;485:237–41.
Singh T, Poterba T, Curtis D, Akil H, Al Eissa M, Barchas JD, et al. Rare coding variants in ten genes confer substantial risk for schizophrenia. Nature. 2022;604:509–16.
Al-Mubarak BR, Omar A, Baz B, Al-Abdulaziz B, Magrashi AI, Al-Yemni E, et al. Whole exome sequencing in ADHD trios from single and multi-incident families implicates new candidate genes and highlights polygenic transmission. Eur J Hum Genet EJHG. 2020;28:1098–110.
Malhotra D, Sebat J. CNVs: harbingers of a rare variant revolution in psychiatric genetics. Cell. 2012;148:1223–41.
Szatmari P, Paterson AD, Zwaigenbaum L, Roberts W, Brian J, Liu XQ, et al. Mapping autism risk loci using genetic linkage and chromosomal rearrangements. NatGenet. 2007;39:319–28.
Jiang YH, Yuen RK, Jin X, Wang M, Chen N, Wu X, et al. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am J Hum Genet. 2013;93:249–63.
Trost B, Engchuan W, Nguyen CM, Thiruvahindrapuram B, Dolzhenko E, Backstrom I, et al. Genome-wide detection of tandem DNA repeats that are expanded in autism. Nature. 2020;586:80–86.
Demontis D, Walters GB, Athanasiadis G, Walters R, Therrien K, Nielsen TT, et al. Genome-wide analyses of ADHD identify 27 risk loci, refine the genetic architecture and implicate several cognitive domains. Nat Genet. 2023;55:198–208.
Trost, Thiruvahindrapuram B, Chan AJS B, Engchuan W, Higginbotham EJ, Howe JL, et al. Genomic architecture of autism from comprehensive whole-genome sequence annotation. Cell. 2022;185:4409–27 e4418.
Antaki D, Guevara J, Maihofer AX, Klein M, Gujral M, Grove J, et al. A phenotypic spectrum of autism is attributable to the combined effects of rare variants, polygenic risk and sex. Nat Genet. 2022;54:1284–92.
Bergen SE, Ploner A, Howrigan D, Group CNVA, the Schizophrenia Working Group of the Psychiatric Genomics C, O’Donovan MC, et al. Joint contributions of rare copy number variants and common SNPs to risk for schizophrenia. Am J Psychiatry. 2019;176:29–35.
Weiner DJ, Wigdor EM, Ripke S, Walters RK, Kosmicki JA, Grove J, et al. Polygenic transmission disequilibrium confirms that common and rare variation act additively to create risk for autism spectrum disorders. Nat Genet. 2017;49:978–85.
Tansey KE, Rees E, Linden DE, Ripke S, Chambert KD, Moran JL, et al. Common alleles contribute to schizophrenia in CNV carriers. Mol Psychiatry. 2016;21:1085–9.
Calle Sanchez X, Helenius D, Bybjerg-Grauholm J, Pedersen C, Hougaard DM, Borglum AD, et al. Comparing copy number variations in a Danish case cohort of individuals with psychiatric disorders. JAMA Psychiatry. 2022;79:59–69.
Wright CF, Eberhardt RY, Constantinou P, Hurles ME, FitzPatrick DR, Firth HV, et al. Evaluating variants classified as pathogenic in ClinVar in the DDD study. Genet Med. 2021;23:571–5.
Girirajan S, Rosenfeld JA, Cooper GM, Antonacci F, Siswara P, Itsara A, et al. A recurrent 16p12.1 microdeletion supports a two-hit model for severe developmental delay. Nat Genet. 2010;42:203–9.
Williams HJ, Monks S, Murphy KC, Kirov G, O’Donovan MC, Owen MJ. Schizophrenia two-hit hypothesis in velo-cardio facial syndrome. Am J Med Genet B Neuropsychiatr Genet. 2013;162B:177–82.
Pizzo L, Jensen M, Polyak A, Rosenfeld JA, Mannik K, Krishnan A, et al. Rare variants in the genetic background modulate cognitive and developmental phenotypes in individuals carrying disease-associated variants. Genet Med. 2019;21:816–25.
Hill M, Hammond J, Lewis C, Mellis R, Clement E, Chitty LS. Delivering genome sequencing for rapid genetic diagnosis in critically ill children: parent and professional views, experiences and challenges. Eur J Hum Genet EJHG. 2020;28:1529–40.
Dimmock DP, Clark MM, Gaughran M, Cakici JA, Caylor SA, Clarke C, et al. An RCT of rapid genomic sequencing among seriously Ill infants results in high clinical utility, changes in management, and low perceived harm. Am J Hum Genet. 2020;107:942–52.
https://www.genomicsengland.co.uk/initiatives/newborns/ethics
Spiekerkoetter U, Bick D, Scott R, Hopkins H, Krones T, Gross ES, et al. Genomic newborn screening: are we entering a new era of screening? J Inherit Metab Dis. 2023;46:778–95.
Scocchia A, Wigby KM, Masser-Frye D, Del Campo M, Galarreta CI, Thorpe E, et al. Clinical whole genome sequencing as a first-tier test at a resource-limited dysmorphology clinic in Mexico. NPJ Genom Med. 2019;4:5.
Lionel AC, Costain G, Monfared N, Walker S, Reuter MS, Hosseini SM, et al. Improved diagnostic yield compared with targeted gene sequencing panels suggests a role for whole-genome sequencing as a first-tier genetic test. Genet Med. 2018;20:435–43.
Stavropoulos DJ, Merico D, Jobling R, Bowdin S, Monfared N, Thiruvahindrapuram B, et al. Whole genome sequencing expands diagnostic utility and improves clinical management in pediatric medicine. NPJ Genom Med. 2016;1:15012.
Gymrek M. A genomic view of short tandem repeats. Curr Opin Genet Dev. 2017;44:9–16.
Brandler WM, Antaki D, Gujral M, Kleiber ML, Whitney J, Maile MS, et al. Paternally inherited cis-regulatory structural variants are associated with autism. Science. 2018;360:327–31.
Brandler WM, Antaki D, Gujral M, Noor A, Rosanio G, Chapman TR, et al. Frequency and complexity of de novo structural mutation in autism. Am J Hum Genet. 2016;98:667–79.
Battle SL, Puiu D, Group TOmW, Verlouw J, Broer L, Boerwinkle E, et al. A bioinformatics pipeline for estimating mitochondrial DNA copy number and heteroplasmy levels from whole genome sequencing data. NAR Genom Bioinform. 2022;4:lqac034.
An JY, Lin K, Zhu L, Werling DM, Dong S, Brand H, et al. Genome-wide de novo risk score implicates promoter variation in autism spectrum disorder. Science. 2018;362:eaat6576.
Polygenic Risk Score Task Force of the International Common Disease A. Responsible use of polygenic risk scores in the clinic: potential benefits, risks and gaps. Nat Med. 2021;27:1876–84.
Inouye M, Abraham G, Nelson CP, Wood AM, Sweeting MJ, Dudbridge F, et al. Genomic risk prediction of coronary artery disease in 480,000 adults: implications for primary prevention. J Am Coll Cardiol. 2018;72:1883–93.
Elliott J, Bodinier B, Bond TA, Chadeau-Hyam M, Evangelou E, Moons KGM, et al. Predictive accuracy of a polygenic risk score-enhanced prediction model vs a clinical risk score for coronary artery disease. JAMA. 2020;323:636–45.
Abraham G, Havulinna AS, Bhalala OG, Byars SG, De Livera AM, Yetukuri L, et al. Genomic prediction of coronary heart disease. Eur Heart J. 2016;37:3267–78.
Riveros-Mckay F, Weale ME, Moore R, Selzam S, Krapohl E, Sivley RM, et al. Integrated polygenic tool substantially enhances coronary artery disease prediction. Circ Genom Precis Med. 2021;14:e003304.
O’Sullivan JW, Raghavan S, Marquez-Luna C, Luzum JA, Damrauer SM, Ashley EA, et al. Polygenic risk scores for cardiovascular disease: a scientific statement from the American Heart Association. Circulation. 2022;146:e93–e118.
Kullo IJ, Jouni H, Austin EE, Brown SA, Kruisselbrink TM, Isseh IN, et al. Incorporating a genetic risk score into coronary heart disease risk estimates: effect on low-density lipoprotein cholesterol levels (the MI-GENES Clinical Trial). Circulation. 2016;133:1181–8.
Kuchenbaecker KB, McGuffog L, Barrowdale D, Lee A, Soucy P, Dennis J, et al. Evaluation of polygenic risk scores for breast and ovarian cancer risk prediction in BRCA1 and BRCA2 mutation carriers. J Natl Cancer Inst. 2017;109:djw302.
Mars N, Widen E, Kerminen S, Meretoja T, Pirinen M, Della Briotta Parolo P, et al. The role of polygenic risk and susceptibility genes in breast cancer over the course of life. Nat Commun. 2020;11:6383.
Huntley C, Torr B, Sud A, Rowlands CF, Way R, Snape K, et al. Utility of polygenic risk scores in UK cancer screening: a modelling analysis. Lancet Oncol. 2023;24:658–68.
Salazar de Pablo G, Radua J, Pereira J, Bonoldi I, Arienti V, Besana F, et al. Probability of transition to psychosis in individuals at clinical high risk: an updated meta-analysis. JAMA Psychiatry. 2021;78:970–8.
Koutsouleris N, Dwyer DB, Degenhardt F, Maj C, Urquijo-Castro MF, Sanfelici R, et al. Multimodal machine learning workflows for prediction of psychosis in patients with clinical high-risk syndromes and recent-onset depression. JAMA Psychiatry. 2021;78:195–209.
Perkins DO, Olde Loohuis L, Barbee J, Ford J, Jeffries CD, Addington J, et al. Polygenic risk score contribution to psychosis prediction in a target population of persons at clinical high risk. Am J Psychiatry. 2020;177:155–63.
Davies RW, Fiksinski AM, Breetvelt EJ, Williams NM, Hooper SR, Monfeuga T, et al. Using common genetic variation to examine phenotypic expression and risk prediction in 22q11.2 deletion syndrome. Nat Med. 2020;26:1912–8.
Torkamani A, Wineinger NE, Topol EJ. The personal and clinical utility of polygenic risk scores. Nat Rev Genet. 2018;19:581–90.
Pedersen CB, Mors O, Bertelsen A, Waltoft BL, Agerbo E, McGrath JJ, et al. A comprehensive nationwide study of the incidence rate and lifetime risk for treated mental disorders. JAMA Psychiatry. 2014;71:573–81.
Whitehouse AJO, Varcin KJ, Pillar S, Billingham W, Alvares GA, Barbaro J, et al. Effect of preemptive intervention on developmental outcomes among infants showing early signs of autism: a randomized clinical trial of outcomes to diagnosis. JAMA Pediatr. 2021;175:e213298.
Arango C, Diaz-Caneja CM, McGorry PD, Rapoport J, Sommer IE, Vorstman JA, et al. Preventive strategies for mental health. Lancet Psychiatry. 2018;5:591–604.
Riggs ER, Andersen EF, Cherry AM, Kantarci S, Kearney H, Patel A et al. Technical standards for the interpretation and reporting of constitutional copy-number variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics (ACMG) and the Clinical Genome Resource (ClinGen). Genet Med. 2020;22:245–57.
Rammos A, Kirov G, Hubbard L, Walters JTR, Holmans P, Owen MJ, et al. Family-based analysis of the contribution of rare and common genetic variants to school performance in schizophrenia. Mol Psychiatry. 2023;28:2081–7.
van den Broek JJ, Schechter CB, van Ravesteyn NT, Janssens A, Wolfson MC, Trentham-Dietz A, et al. Personalizing breast cancer screening based on polygenic risk and family history. J Natl Cancer Inst. 2021;113:434–42.
Schaaf CP, Betancur C, Yuen RKC, Parr JR, Skuse DH, Gallagher L, et al. A framework for an evidence-based gene list relevant to autism spectrum disorder. Nat Rev Genet. 2020;21:367–76.
Douard E, Zeribi A, Schramm C, Tamer P, Loum MA, Nowak S, et al. Effect sizes of deletions and duplications on autism risk across the genome. Am J Psychiatry. 2021;178:87–98.
Acknowledgements
We thank Dr Robert W Davies at the Department of Statistics, University of Oxford, Oxford, UK, for writing the code underpinning Fig. 2b. This work was supported by the SickKids Psychiatry Associates Chair in Developmental (JV) and by NIH grants U01MH119741-01 (JV) and U01MH119746 (JS). These funding sources did not influence content of this manuscript.
Author information
Authors and Affiliations
Contributions
JV, JS, VRB and SJ contributed to the conception of the topic of this expert review as well as to the writing of the different iterations of this work. JV, JS, VRB and SJ approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
JV has served as a consultant for Nobias Therapeutics Inc. and has received speaker fees for Henry Stewart Talks Ltd.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Vorstman, J., Sebat, J., Bourque, VR. et al. Integrative genetic analysis: cornerstone of precision psychiatry. Mol Psychiatry 30, 229–236 (2025). https://doi.org/10.1038/s41380-024-02706-2
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41380-024-02706-2