Abstract
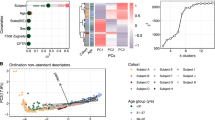
Bacterial infection and inflammation of the airways are the leading causes of morbidity and mortality in persons with cystic fibrosis (CF). The ecology of the bacterial communities inhabiting CF airways is poorly understood, especially with respect to how community structure, dynamics, and microbial metabolic activity relate to clinical outcomes. In this study, the bacterial communities in 818 sputum samples from 109 persons with CF were analyzed by sequencing bacterial 16S rRNA gene amplicons. We identified eight alternative community types (pulmotypes) by using a Dirichlet multinomial mixture model and studied their temporal dynamics in the cohort. Across patients, the pulmotypes displayed chronological patterns in the transition among each other. Furthermore, significant correlations between pulmotypes and patient clinical status were detected by using multinomial mixed effects models, principal components regression, and statistical testing. Constructing pulmotype-specific metabolic activity profiles, we found that pulmotype microbiota drive distinct community functions including mucus degradation or increased acid production. These results indicate that pulmotypes are the result of ordered, underlying drivers such as predominant metabolism, ecological competition, and niche construction and can form the basis for quantitative, predictive models supporting clinical treatment decisions.
Similar content being viewed by others
Log in or create a free account to read this content
Gain free access to this article, as well as selected content from this journal and more on nature.com
or
Data availability
The sequencing data are available in the NCBI Bioproject database (accession numbers: PRJNA423040, PRJNA756039). Detailed SRA accession numbers are additionally listed as Supplementary information.
References
Filkins LM, Hampton TH, Gifford AH, Gross MJ, Hogan DA, Sogin ML, et al. Prevalence of Streptococci and increased polymicrobial diversity associated with cystic fibrosis patient stability. J Bacteriol. 2012;194:4709–17.
Fodor AA, Klem ER, Gilpin DF, Elborn JS, Boucher RC, Tunney MM, et al. The adult cystic fibrosis airway microbiota is stable over time and infection type, and highly resilient to antibiotic treatment of exacerbations. PLoS One. 2012;7:e45001.
Goddard AF, Staudinger BJ, Dowd SE, Joshi-Datar A, Wolcott RD, Aitken ML, et al. Direct sampling of cystic fibrosis lungs indicates that DNA-based analyses of upper-airway specimens can misrepresent lung microbiota. Proc Natl Acad Sci USA. 2012;109:13769–74.
Guss AM, Roeselers G, Newton IL, Young CR, Klepac-Ceraj V, Lory S, et al. Phylogenetic and metabolic diversity of bacteria associated with cystic fibrosis. ISME J. 2011;5:20–9.
Harris JK, De Groote MA, Sagel SD, Zemanick ET, Kapsner R, Penvari C, et al. Molecular identification of bacteria in bronchoalveolar lavage fluid from children with cystic fibrosis. Proc Natl Acad Sci USA. 2007;104:20529–33.
Brown PS, Pope CE, Marsh RL, Qin X, McNamara S, Gibson R, et al. Directly sampling the lung of a young child with cystic fibrosis reveals diverse microbiota. Ann Am Thorac Soc. 2014;11:1049–55.
Jorth P, Staudinger BJ, Wu X, Hisert KB, Hayden H, Garudathri J, et al. Regional isolation drives bacterial diversification within cystic fibrosis lungs. Cell Host Microbe. 2015;18:307–19.
Sibley CD, Parkins MD, Rabin HR, Duan K, Norgaard JC, Surette MG. A polymicrobial perspective of pulmonary infections exposes an enigmatic pathogen in cystic fibrosis patients. Proc Natl Acad Sci USA. 2008;105:15070–5.
van der Gast CJ, Walker AW, Stressmann FA, Rogers GB, Scott P, Daniels TW, et al. Partitioning core and satellite taxa from within cystic fibrosis lung bacterial communities. ISME J. 2011;5:780–91.
Zhao J, Carmody LA, Kalikin LM, Li J, Petrosino JF, Schloss PD, et al. Impact of enhanced Staphylococcus DNA extraction on microbial community measures in cystic fibrosis sputum. PLoS One. 2012;7:e33127.
Carmody LA, Zhao J, Schloss PD, Petrosino JF, Murray S, Young VB, et al. Changes in cystic fibrosis airway microbiota at pulmonary exacerbation. Ann Am Thorac Soc. 2013;10:179–87.
Cox MJ, Allgaier M, Taylor B, Baek MS, Huang YJ, Daly RA, et al. Airway microbiota and pathogen abundance in age-stratified cystic fibrosis patients. PLoS One. 2010;5:e11044.
Stressmann FA, Rogers GB, van der Gast CJ, Marsh P, Vermeer LS, Carroll MP, et al. Long-term cultivation-independent microbial diversity analysis demonstrates that bacterial communities infecting the adult cystic fibrosis lung show stability and resilience. Thorax. 2012;67:867–73.
Zhao J, Schloss PD, Kalikin LM, Carmody LA, Foster BK, Petrosino JF, et al. Decade-long bacterial community dynamics in cystic fibrosis airways. Proc Natl Acad Sci USA. 2012;109:5809–14.
Rogers GB, Bruce KD, Hoffman LR. How can the cystic fibrosis respiratory microbiome influence our clinical decision-making? Curr Opin Pulm Med. 2017;23:536–43.
Widder S, Knapp S. Microbial metabolites in cystic fibrosis: a target for future therapy? Am J Respir Cell Mol Biol. 2019;61:132–3.
Mahboubi MA, Carmody LA, Foster BK, Kalikin LM, VanDevanter DR, LiPuma JJ. Culture-based and culture-independent bacteriologic analysis of cystic fibrosis respiratory specimens. J Clin Microbiol. 2016;54:613–9.
Carmody LA, Caverly LJ, Foster BK, Rogers MAM, Kalikin LM, Simon RH, et al. Fluctuations in airway bacterial communities associated with clinical states and disease stages in cystic fibrosis. PLoS One. 2018;13:e0194060.
Zhao J, Li J, Schloss PD, Kalikin LM, Raymond TA, Petrosino JF, et al. Effect of sample storage conditions on cultureindependent bacterial community measures in cystic fibrosis sputum specimens. J Clin Microbiol 2011;49:3717–8.
Hnizdo E, Yu L, Freyder L, Attfield M, Lefante J & Glindmeyer HW. The precision of longitudinal lung function measurements: Monitoring and interpretation. Occup Environ Med 2005;62:695–701.
Konstan MW, Wagener JS, VanDevanter DR. Characterizing aggressiveness and predicting future progression of CF lung disease. J Cyst Fibros. 2009;8:S15–19.
Schloss PD, Gevers D, Westcott SL. Reducing the effects of PCR amplification and sequencing artifacts on 16s rRNA-based studies. PLoS One. 2011;6:e27310.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, et al. The ribosomal database project: Improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 2009;37:D141–145.
Sung J, Kim S, Cabatbat JJT, Jang S, Jin YS, Jung GY, et al. Global metabolic interaction network of the human gut microbiota for context-specific community-scale analysis. Nat Commun 2017;8:15393.
Friedman J, Alm EJ. Inferring correlation networks from genomic survey data. PLoS Comput Biol. 2012;8:e1002687.
Holmes I, Harris K, Quince C. Dirichlet multinomial mixtures: generative models for microbial metagenomics. PLoS One. 2012;7:e30126.
Ding T, Schloss PD. Dynamics and associations of microbial community types across the human body. Nature. 2014;509:357–60.
Price KE, Hampton TH, Gifford AH, Dolben EL, Hogan DA, Morrison HG, et al. Unique microbial communities persist in individual cystic fibrosis patients throughout a clinical exacerbation. Microbiome 2013;1:27.
Carmody LA, Zhao J, Kalikin LM, LeBar W, Simon RH, Venkataraman A, et al. The daily dynamics of cystic fibrosis airway microbiota during clinical stability and at exacerbation. Microbiome 2015;3:12.
de Dios Caballero J, Vida R, Cobo M, Maiz L, Suarez L, Galeano J, et al. Individual patterns of complexity in cystic fibrosis lung microbiota, including predator bacteria, over a 1-year period. mBio 2017;8::e00959–17.
Whelan, FJ, Heirali AA, Rossi L, Rabin HR, Parkins MD, & Surette MG. Longitudinal sampling of the lung microbiota in individuals with cystic fibrosis. PLoS One 2017:12:e0172811.
Noecker C, Eng A, Srinivasan S, Theriot CM, Young VB, Jansson JK, et al. Metabolic model-based integration of microbiome taxonomic and metabolomic profiles elucidates mechanistic links between ecological and metabolic variation. mSystems 2016;1.
Douglas, GM, Maffei, VJ, Zaneveld, JR, Yurgel, SN, Brown JR, Taylor CM, et al. PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 2020;38:685–8.
Caspi R, Billington R, Fulcher CA, Keseler IM, Kothari A, Krummenacker M, et al. The MetaCyc database of metabolic pathways and enzymes. Nucleic Acids Res 2018;46:D633–D639.
Quinn RA, Comstock W, Zhang T, Morton JT, da Silva R, Tran A, et al. Niche partitioning of a pathogenic microbiome driven by chemical gradients. Sci Adv 2018;4:eaau1908.
Quinn RA, Whiteson K, Lim YW, Zhao J, Conrad D, LiPuma JJ, et al. Ecological networking of cystic fibrosis lung infections. NPJ Biofilms Microbiomes. 2016;2:4.
Pradeu T, Vivier E. The discontinuity theory of immunity. Sci Immunol. 2016;1:AAG0479.
Flynn JM, Niccum D, Dunitz JM, Hunter RC. Evidence and role for bacterial mucin degradation in cystic fibrosis airway disease. PLoS Pathog. 2016;12:e1005846.
Adamowicz EM, Flynn J, Hunter RC, Harcombe WR. Cross-feeding modulates antibiotic tolerance in bacterial communities. ISME J. 2018;12:2723–35.
Rose MC & Voynow JA. Respiratory tract mucin genes and mucin glycoproteins in health and disease. Physiol Rev 2006;86:245–78.
Tailford LE, Crost EH, Kavanaugh D. & Juge N. Mucin glycan foraging in the human gut microbiome. Front Genet 2015;6:81.
Wheeler KM, Carcamo-Oyarce G, Turner BS, Dellos-Nolan S, Co JY, Lehoux S, et al. Mucin glycans attenuate the virulence of Pseudomonas aeruginosa in infection. Nat Microbiol 2019;4:2146–54.
Twomey KB, O'Connell OJ, McCarthy Y, Dow JM, O'Toole GA, Plant BJ, et al. Bacterial cis-2-unsaturated fatty acids found in the cystic fibrosis airway modulate virulence and persistence of Pseudomonas aeruginosa. ISME J 2012;6:939–50.
Zemanick ET, Wagner BD, Robertson CE, Ahrens RC, Chmiel JF, Clancy JP, et al. Airway microbiota across age and disease spectrum in cystic fibrosis. Eur Respir J 2017;50:1700832.
Lu J, Carmody LA, Opron K, Simon RH, Kalikin LM, Caverly LJ, et al. Parallel analysis of cystic fibrosis sputum and saliva'reveals overlapping communities and an opportunity for sample decontamination. mSystems 2020;5.
Jones KL, Hegab AH, Hillman BC, Simpson KL, Jinkins PA, Grisham MB, et al. Elevation of nitrotyrosine and nitrate concentrations in cystic fibrosis sputum. Pediatr Pulmonol 2000;30:79–85.
Quinn RA, Lim YW, Maughan H, Conrad D, Rohwer F, Whiteson KL. Biogeochemical forces shape the composition and physiology of polymicrobial communities in the cystic fibrosis lung. mBio. 2014;5:e00956–00913.
Mirkovic B, Murray MA, Lavelle GM, Molloy K, Azim AA, Gunaratnam C, et al. The role of short-chain fatty acids, produced by anaerobic bacteria, in the cystic fibrosis airway. Am J Respir Crit Care Med. 2015;192:1314–24.
Trompette A, Gollwitzer ES, Pattaroni C, Lopez-Mejia IC, Riva E, Pernot J, et al. Dietary fiber confers protection against flu by shaping Ly6c(-) patrolling monocyte hematopoiesis and CD8(+) t cell metabolism. Immunity. 2018;48:992–1005.e1008.
Flynn JM, Phan C, Hunter RC. Genome-wide survey of Pseudomonas aeruginosa PA14 reveals a role for the glyoxylate pathway and extracellular proteases in the utilization of mucin. Infect Immun. 2017;85:e00182–17.
Jorth P, Ehsan Z, Rezayat A, Caldwell E, Pope C, Brewington JJ, et al. Direct lung sampling indicates that established pathogens dominate early infections in children with cystic fibrosis. Cell Rep. 2019;27:1190–204.e1193.
Charalampous T, Kay GL, Richardson H, Aydin A, Baldan R, Jeanes C, et al. Nanopore metagenomics enables rapid clinical diagnosis of bacterial lower respiratory infection. Nat Biotechnol. 2019;37:783–92.
Cowley ES, Kopf SH, LaRiviere A, Ziebis W, Newman DK. Pediatric cystic fibrosis sputum can be chemically dynamic, anoxic, and extremely reduced due to hydrogen sulfide formation. mBio. 2015;6:e00767.
Cuthbertson L, Walker AW, Oliver AE, Rogers GB, Rivett DW, Hampton TH, et al. Lung function and microbiota diversity in cystic fibrosis. Microbiome 2020;8:45.
Acknowledgements
We gratefully acknowledge the many individuals who provided the sputum samples used in this study. We thank the Michigan Medicine clinical microbiology laboratory for the generous assistance in obtaining sputum samples for this study. We thank Prof. Barbara Bailey for fruitful discussions about random forests. This study was funded by NHLBI grants 1RC1HL100809 and 1R01HL136647 to JJL. Additional support was provided by NIH CTSA grant UL1RR024986 and the Charles Woodson Pediatric Research Fund. SW was supported by the Austrian Science Fund (FWF) Elise Richter V585-B31.
Author information
Authors and Affiliations
Contributions
SW provided conceptualization, investigation, methodology, formal analysis, visualization, writing—original draft, and writing—review and editing. JZ conducted methodology, formal analysis, visualization, writing—original draft, and writing—review and editing. LAC contributed investigation, data curation, writing—original draft, and writing—review and editing. QZ and PDS provided software and supported data curation. LMK contributed to data curation, project administration, and writing—review and editing. JJL contributed to conceptualization, funding acquisition, resources, supervision, writing—original draft, and writing—review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Widder, S., Zhao, J., Carmody, L.A. et al. Association of bacterial community types, functional microbial processes and lung disease in cystic fibrosis airways. ISME J 16, 905–914 (2022). https://doi.org/10.1038/s41396-021-01129-z
Received:
Revised:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/s41396-021-01129-z
This article is cited by
-
Crosstalk between ROS-inflammatory gene expression axis in the progression of lung disorders
Naunyn-Schmiedeberg's Archives of Pharmacology (2025)
-
Microbial community organization designates distinct pulmonary exacerbation types and predicts treatment outcome in cystic fibrosis
Nature Communications (2024)
-
Limited effects of azithromycin on the oropharyngeal microbiome in children with CF and early pseudomonas infection
BMC Microbiology (2023)
-
CDSS for Early Recognition of Respiratory Diseases based on AI Techniques: A Systematic Review
Wireless Personal Communications (2023)
-
CF-Seq, an accessible web application for rapid re-analysis of cystic fibrosis pathogen RNA sequencing studies
Scientific Data (2022)