Abstract
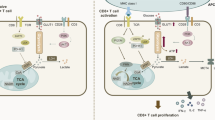
The long-term persistence of naive T lymphocytes is maintained by a state of relative quiescence. Upon antigenic stimulation, these naive T cells undergo rapid activation and proliferation, differentiating into effector cells with specific clonal expansion. Recently, in-depth studies have revealed a fundamental difference in the metabolic requirements of distinct T cell subsets. The fate of CD4 + T cells is influenced by glucose-mediated glycolysis and oxidative phosphorylation (OXPHOS). In this context, key enzymes and various glycolytic intermediates, in conjunction with transcription factors and cytokines, play a crucial role in CD4 + T cell differentiation and function. In our study, we investigated the mechanisms underlying glycolytic reprogramming in CD4 + T cells, with a particular focus on the role of glycolytic enzymes in modulating cytokines and transcription factors that govern T cell differentiation.Our aim is to provide novel insights into the treatment of clinically relevant immune diseases by thoroughly elucidating the characteristics and potential regulatory mechanisms of glucose metabolism in CD4 + T cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data availability is not applicable to this article as no new data were created or analyzed in this study.
References
Taniuchi I. CD4 helper and CD8 cytotoxic T cell differentiation. Annu Rev Immunol. 2018;36:579–601.
Zhu J. T Helper cell differentiation, heterogeneity, and plasticity. Cold Spring Harb Perspect Biol. 2018;10:a030338.
MacIver NJ, Michalek RD, Rathmell JC. Metabolic regulation of T lymphocytes. Annu Rev Immunol. 2013;31:259–83.
Freemerman AJ, Johnson AR, Sacks GN, Milner JJ, Kirk EL, Troester MA, et al. Metabolic reprogramming of macrophages: glucose transporter 1 (GLUT1)-mediated glucose metabolism drives a proinflammatory phenotype. J Biol Chem. 2014;289:7884–96.
Peng M, Yin N, Chhangawala S, Xu K, Leslie CS, Li MO. Aerobic glycolysis promotes T helper 1 cell differentiation through an epigenetic mechanism. Sci (N Y, NY). 2016;354:481–4.
Liu S, Liao S, Liang L, Deng J, Zhou Y. The relationship between CD4+ T cell glycolysis and their functions. Trends Endocrinol Metab. 2023;34:345–60.
Almeida L, Dhillon-LaBrooy A, Carriche G, Berod L, Sparwasser T. CD4(+) T-cell differentiation and function: Unifying glycolysis, fatty acid oxidation, polyamines NAD mitochondria. J Allergy Clin Immunol. 2021;148:16–32.
Palmer CS, Ostrowski M, Balderson B, Christian N, Crowe SM. Glucose metabolism regulates T cell activation, differentiation, and functions. Front Immunol. 2015;6:1.
Song W, Li D, Tao L, Luo Q, Chen L. Solute carrier transporters: the metabolic gatekeepers of immune cells. Acta Pharmaceutica Sin B. 2020;10:61–78.
Chen LY, Phelix CF. Extracellular gating of glucose transport through GLUT 1. Biochemical Biophysical Res Commun. 2019;511:573–8.
Yang L, Miao L, Liang F, Huang H, Teng X, Li S, et al. The mTORC1 effectors S6K1 and 4E-BP play different roles in CNS axon regeneration. Nat Commun. 2014;5:5416.
Zhang Z, Li X, Yang F, Chen C, Liu P, Ren Y, et al. DHHC9-mediated GLUT1 S-palmitoylation promotes glioblastoma glycolysis and tumorigenesis. Nat Commun. 2021;12:5872.
Wu N, Zheng B, Shaywitz A, Dagon Y, Tower C, Bellinger G, et al. AMPK-dependent degradation of TXNIP upon energy stress leads to enhanced glucose uptake via GLUT1. Mol Cell. 2013;49:1167–75.
Tanegashima K, Sato-Miyata Y, Funakoshi M, Nishito Y, Aigaki T, Hara T. Epigenetic regulation of the glucose transporter gene Slc2a1 by β-hydroxybutyrate underlies preferential glucose supply to the brain of fasted mice. Genes Cells. 2016;22:71–83.
Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Sci (N Y, NY). 2009;324:1029–33.
Buck MD, O’Sullivan D, Klein Geltink RI, Curtis JD, Chang CH, Sanin DE, et al. Mitochondrial Dynamics Controls T Cell Fate through Metabolic Programming. Cell. 2016;166:63–76.
Liberti MV, Locasale JW. The Warburg effect: how does it benefit cancer cells?. Trends Biochem Sci. 2016;41:211–8.
Ganapathy-Kanniappan S. Molecular intricacies of aerobic glycolysis in cancer: current insights into the classic metabolic phenotype. Crit Rev Biochem Mol Biol. 2018;53:667–82.
Pearce EL, Pearce EJ. Metabolic pathways in immune cell activation and quiescence. Immunity. 2013;38:633–43.
Stark JM, Tibbitt CA, Coquet JM. The metabolic requirements of Th2 cell differentiation. Front Immunol. 2019;10:2318.
Koopman R, Ly CH, Ryall JG. A metabolic link to skeletal muscle wasting and regeneration. Front Physiol. 2014;5:32.
Lunt SY, Vander Heiden MG. Aerobic glycolysis: meeting the metabolic requirements of cell proliferation. Annu Rev Cell Dev Biol. 2011;27:441–64.
Bailis W, Shyer JA, Zhao J, Canaveras JCG, Al Khazal FJ, Qu R, et al. Distinct modes of mitochondrial metabolism uncouple T cell differentiation and function. Nature. 2019;571:403–7.
Gerriets VA, Kishton RJ, Nichols AG, Macintyre AN, Inoue M, Ilkayeva O, et al. Metabolic programming and PDHK1 control CD4+ T cell subsets and inflammation. J Clin Invest. 2015;125:194–207.
Angiari S, Runtsch MC, Sutton CE, Palsson-McDermott EM, Kelly B, Rana N, et al. Pharmacological activation of pyruvate kinase M2 inhibits CD4(+) T cell pathogenicity and suppresses autoimmunity. Cell Metab. 2020;31:391–405.e8.
Liberti MV, Dai Z, Wardell SE, Baccile JA, Liu X, Gao X, et al. A predictive model for selective targeting of the Warburg Effect through GAPDH inhibition with a natural product. Cell Metab. 2017;26:648–59.e8.
Kornberg MD, Bhargava P, Kim PM, Putluri V, Snowman AM, Putluri N, et al. Dimethyl fumarate targets GAPDH and aerobic glycolysis to modulate immunity. Sci (N Y, NY). 2018;360:449–53.
Kono M, Maeda K, Stocton-Gavanescu I, Pan W, Umeda M, Katsuyama E, et al. Pyruvate kinase M2 is requisite for Th1 and Th17 differentiation. JCI Insight. 2019;4:e127395.
Almeida L, Lochner M, Berod L, Sparwasser T. Metabolic pathways in T cell activation and lineage differentiation. Semin Immunol. 2016;28:514–24.
Maciolek JA, Pasternak JA, Wilson HL. Metabolism of activated T lymphocytes. Curr Opin Immunol. 2014;27:60–74.
Robinson BH, Williams GR, Halperin ML, Leznoff CC. Factors affecting the kinetics and equilibrium of exchange reactions of the citrate-transporting system of rat liver mitochondria. J Biol Chem. 1971;246:5280–6.
Buck MD, O’Sullivan D, Pearce EL. T cell metabolism drives immunity. J Exp Med. 2015;212:1345–60.
Magaway C, Kim E, Jacinto E. Targeting mTOR and metabolism in cancer: lessons and innovations. Cells. 2019;8:1584.
Liu Q, Zhu F, Liu X, Lu Y, Yao K, Tian N, et al. Non-oxidative pentose phosphate pathway controls regulatory T cell function by integrating metabolism and epigenetics. Nat Metab. 2022;4:559–74.
Wong BW, Wang X, Zecchin A, Thienpont B, Cornelissen I, Kalucka J, et al. The role of fatty acid β-oxidation in lymphangiogenesis. Nature. 2016;542:49–54.
Xiong J, Kawagishi H, Yan Y, Liu J, Wells QS, Edmunds LR, et al. A metabolic basis for endothelial-to-mesenchymal transition. Mol Cell. 2018;69:689–98.e7.
Xuekai L, Yan S, Jian C, Yifei S, Xinyue W, Wenyuan Z, et al. Advances in reprogramming of energy metabolism in tumor T cells. Front Immunol. 2024;15:1347181.
Hosios Aaron M, Hecht Vivian C, Danai Laura V, Johnson Marc O, Rathmell Jeffrey C, Steinhauser Matthew L, et al. Amino acids rather than glucose account for the majority of cell mass in proliferating mammalian cells. Developmental Cell. 2016;36:540–9.
Ma S, Ming Y, Wu J, Cui G. Cellular metabolism regulates the differentiation and function of T-cell subsets. Cell Mol Immunol. 2024;21:419–35.
Zhang S, Zhang X, Wang K, Xu X, Li M, Zhang J, et al. Newly generated CD4+ T cells acquire metabolic quiescence after thymic egress. J Immunol. 2018;200:1064–77.
Yang J, Chen Y, Li X, Qin H, Bao J, Wang C, et al. Complex interplay between metabolism and CD4+ T-cell activation, differentiation, and function: a novel perspective for atherosclerosis immunotherapy. Cardiovascular Drugs Ther. 2023;38:1033–46.
Rathmell JC, Vander Heiden MG, Harris MH, Frauwirth KA, Thompson CB. In the Absence of Extrinsic Signals, Nutrient Utilization by Lymphocytes Is Insufficient to Maintain Either Cell Size or Viability. Mol Cell. 2000;6:683–92.
Baixauli F, Acín-Pérez R, Villarroya-Beltrí C, Mazzeo C, Nuñez-Andrade N, Gabandé-Rodriguez E, et al. Mitochondrial respiration controls lysosomal function during inflammatory T cell responses. Cell Metab. 2015;22:485–98.
Tarasenko TN, Pacheco SE, Koenig MK, Gomez-Rodriguez J, Kapnick SM, Diaz F, et al. Cytochrome c oxidase activity is a metabolic checkpoint that regulates cell fate decisions during T cell activation and differentiation. Cell Metab. 2017;25:1254–68.e7.
Chapman NM, Boothby MR, Chi H. Metabolic coordination of T cell quiescence and activation. Nat Rev Immunol. 2020;20:55–70.
Takada K, Jameson SC. Naive T cell homeostasis: from awareness of space to a sense of place. Nat Rev Immunol. 2009;9:823–32.
Sprent J, Surh CD. Normal T cell homeostasis: the conversion of naive cells into memory-phenotype cells. Nat Immunol. 2011;12:478–84.
Mendoza A, Fang V, Chen C, Serasinghe M, Verma A, Muller J, et al. Lymphatic endothelial S1P promotes mitochondrial function and survival in naive T cells. Nature. 2017;546:158–61.
Peng M, Li MO. Metabolism along the life journey of T cells. Life Metab. 2023;2:load002.
Macintyre AN, Gerriets VA, Nichols AG, Michalek RD, Rudolph MC, Deoliveira D, et al. The glucose transporter Glut1 is selectively essential for CD4 T cell activation and effector function. Cell Metab. 2014;20:61–72.
Jacobs SR, Michalek RD, Rathmell JC. IL-7 Is essential for homeostatic control of T cell metabolism in vivo. J Immunol. 2010;184:3461–9.
Yang K, Neale G, Green DR, He W, Chi H. The tumor suppressor Tsc1 enforces quiescence of naive T cells to promote immune homeostasis and function. Nat Immunol. 2011;12:888–97.
Pollizzi KN, Patel CH, Sun I-H, Oh M-H, Waickman AT, Wen J, et al. mTORC1 and mTORC2 selectively regulate CD8+ T cell differentiation. J Clin Investig. 2015;125:2090–108.
Yang K, Shrestha S, Zeng H, Karmaus PeerWF, Neale G, Vogel P, et al. T cell exit from quiescence and differentiation into Th2 cells depend on raptor-mTORC1-mediated metabolic reprogramming. Immunity. 2013;39:1043–56.
Newton RH, Shrestha S, Sullivan JM, Yates KB, Compeer EB, Ron-Harel N, et al. Maintenance of CD4 T cell fitness through regulation of Foxo1. Nat Immunol. 2018;19:838–48.
Dutta A, Zhao B, Love PE. New insights into TCR β-selection. Trends Immunol. 2021;42:735–50.
Rangel Rivera GO, Knochelmann HM, Dwyer CJ, Smith AS, Wyatt MM, Rivera-Reyes AM, et al. Fundamentals of T cell metabolism and strategies to enhance cancer immunotherapy. Front Immunol. 2021;12:645242.
Zhang M, Lin X, Yang Z, Li X, Zhou Z, Love PE, et al. Metabolic regulation of T cell development. Front Immunol. 2022;13:946119.
Jones N, Vincent EE, Cronin JG, Panetti S, Chambers M, Holm SR, et al. Akt and STAT5 mediate naive human CD4+ T-cell early metabolic response to TCR stimulation. Nat Commun. 2019;10:2042.
Tan H, Yang K, Li Y, Shaw TI, Wang Y, Blanco DB, et al. Integrative proteomics and phosphoproteomics profiling reveals dynamic signaling networks and bioenergetics pathways underlying T cell activation. Immunity. 2017;46:488–503.
Forgac M. Vacuolar ATPases: rotary proton pumps in physiology and pathophysiology. Nat Rev Mol Cell Biol. 2007;8:917–29.
Li M, Zhang C-S, Feng J-W, Wei X, Zhang C, Xie C, et al. Aldolase is a sensor for both low and high glucose, linking to AMPK and mTORC1. Cell Res. 2020;31:478–81.
He L, Gomes AP, Wang X, Yoon SO, Lee G, Nagiec MJ, et al. mTORC1 Promotes Metabolic Reprogramming by the Suppression of GSK3-Dependent Foxk1 Phosphorylation. Mol Cell. 2018;70:949–60.e4.
Samanta D, Semenza GL. Metabolic adaptation of cancer and immune cells mediated by hypoxia-inducible factors. Biochimica et Biophysica Acta (BBA) - Rev Cancer. 2018;1870:15–22.
Lin J, Fan L, Han Y, Guo J, Hao Z, Cao L, et al. The mTORC1/eIF4E/HIF-1α pathway mediates glycolysis to support brain hypoxia resistance in the Gansu Zokor, Eospalax cansus. Front Physiol. 2021;12:626240.
Choi G, Ju HY, Bok J, Choi J, Shin JW, Oh H, et al. NRF2 is a spatiotemporal metabolic hub essential for the polyfunctionality of Th2 cells. Proc Natl Acad Sci. 2024;121:e2319994121.
DuPage M, Bluestone JA. Harnessing the plasticity of CD4(+) T cells to treat immune-mediated disease. Nat Rev Immunol. 2016;16:149–63.
Hirahara K, Nakayama T. CD4+ T-cell subsets in inflammatory diseases: beyond the Th1/Th2 paradigm. Int Immunol. 2016;28:163–71.
Li P, Spolski R, Liao W, Leonard WJ. Complex interactions of transcription factors in mediating cytokine biology in T cells. Immunol Rev. 2014;261:141–56.
Rogozynski NP, Dixon B. The Th1/Th2 paradigm: a misrepresentation of helper T cell plasticity. Immunol Lett. 2024;268:106870.
Michalek RD, Gerriets VA, Jacobs SR, Macintyre AN, MacIver NJ, Mason EF, et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J Immunol. 2011;186:3299–303.
Ray JP, Staron MM, Shyer JA, Ho PC, Marshall HD, Gray SM, et al. The interleukin-2-mTORc1 kinase axis defines the signaling, differentiation, and metabolism of T Helper 1 and Follicular B Helper T Cells. Immunity. 2015;43:690–702.
Rashida Gnanaprakasam JN, Wu R, Wang R. Metabolic reprogramming in modulating T cell reactive oxygen species generation and antioxidant capacity. Front Immunol. 2018;9:1075.
Tindemans I, Joosse ME, Samsom JN. Dissecting the heterogeneity in T-cell mediated inflammation in IBD. Cells. 2020;9:110.
Krueger PD, Goldberg MF, Hong SW, Osum KC, Langlois RA, Kotov DI, et al. Two sequential activation modules control the differentiation of protective T helper-1 (Th1) cells. Immunity. 2021;54:687–701.e4.
Chang CH, Curtis JD, Maggi LB Jr, Faubert B, Villarino AV, et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell. 2013;153:1239–51.
Jing X, Lyu J, Xiong J, August A. Acetate regulates GAPDH acetylation and T helper 1 cell differentiation. Mol Biol Cell. 2023;34:br10.
Augoff K, Hryniewicz-Jankowska A, Tabola R. Lactate dehydrogenase 5: An old friend and a new hope in the war on cancer. Cancer Lett. 2015;358:1–7.
Wu H, Huang H, Zhao Y. Interplay between metabolic reprogramming and post-translational modifications: from glycolysis to lactylation. Front Immunol. 2023;14:1211221.
Cheng L, Zhao Y, Tang M, Luo Z, Wang X. Knockdown of ISOC1 suppresses cell proliferation in pancreatic cancer in vitro. Oncol Lett. 2019;17:4263–70.
Yamaga R, Ikeda K, Boele J, Horie-Inoue K, Takayama K-i, Urano T, et al. Systemic identification of estrogen-regulated genes in breast cancer cells through cap analysis of gene expression mapping. Biochem Biophys Res Commun. 2014;447:531–6.
Gao B, Zhao L, Wang F, Bai H, Li J, Li M, et al. Knockdown of ISOC1 inhibits the proliferation and migration and induces the apoptosis of colon cancer cells through the AKT/GSK-3β pathway. Carcinogenesis. 2020;41:1123–33.
Teli DM, Gajjar AK. Glycogen synthase kinase-3: A potential target for diabetes. Bioorg Med Chem. 2023;92:117406.
Beurel E, Grieco SF, Jope RS. Glycogen synthase kinase-3 (GSK3): Regulation, actions, and diseases. Pharm Ther. 2015;148:114–31.
Salles, ÉMd, Raeder PL, Angeli CB, Santiago VF, de Souza CN, et al. P2RX7 signaling drives the differentiation of Th1 cells through metabolic reprogramming for aerobic glycolysis. Front Immunol. 2023;14:1140426.
García-Bermúdez J, Cuezva JM. The ATPase Inhibitory Factor 1 (IF1): A master regulator of energy metabolism and of cell survival. Biochimica et Biophysica Acta (BBA) - Bioenerg. 2016;1857:1167–82.
Zhang Y, Zhang Y, Gu W, Sun B. TH1/TH2 cell differentiation and molecular signals. Adv Exp Med Biol. 2014;841:15–44.
Tibbitt CA, Stark JM, Martens L, Ma J, Mold JE, Deswarte K, et al. Single-cell RNA sequencing of the T Helper cell response to house dust mites defines a distinct gene expression signature in airway Th2 cells. Immunity. 2019;51:169–84.e5.
Yang JQ, Kalim KW, Li Y, Zhang S, Hinge A, Filippi MD, et al. RhoA orchestrates glycolysis for TH2 cell differentiation and allergic airway inflammation. J Allergy Clin Immunol. 2016;137:231–45.e4.
Nobs SP, Natali S, Pohlmeier L, Okreglicka K, Schneider C, Kurrer M, et al. PPARγ in dendritic cells and T cells drives pathogenic type-2 effector responses in lung inflammation. J Exp Med. 2017;214:3015–35.
Yamagata T, Skepner J, Yang J. Targeting Th17 effector cytokines for the treatment of autoimmune diseases. Arch Immunol Ther Exp (Warsz). 2015;63:405–14.
Wu L, Hollinshead KER, Hao Y, Au C, Kroehling L, Ng C, et al. Niche-selective inhibition of pathogenic Th17 cells by targeting metabolic redundancy. Cell. 2020;182:641–54.e20.
Xu K, Yin N, Peng M, Stamatiades EG, Chhangawala S, Shyu A, et al. Glycolytic ATP fuels phosphoinositide 3-kinase signaling to support effector T helper 17 cell responses. Immunity. 2021;54:976–87.e7.
Damasceno LEA, Prado DS, Veras FP, Fonseca MM, Toller-Kawahisa JE, Rosa MH, et al. PKM2 promotes Th17 cell differentiation and autoimmune inflammation by fine-tuning STAT3 activation. J Exp Med. 2020;217:e20190613.
Zhao Z, Wang Y, Gao Y, Ju Y, Zhao Y, Wu Z, et al. The PRAK-NRF2 axis promotes the differentiation of Th17 cells by mediating the redox homeostasis and glycolysis. Proc Natl Acad Sci. 2023;120:e2212613120.
Kono M, Yoshida N, Maeda K, Skinner NE, Pan W, Kyttaris VC, et al. Pyruvate dehydrogenase phosphatase catalytic subunit 2 limits Th17 differentiation. Proc Natl Acad Sci USA. 2018;115:9288–93.
Newton R, Priyadharshini B, Turka LA. Immunometabolism of regulatory T cells. Nat Immunol. 2016;17:618–25.
Zeng H, Chi H. Metabolic control of regulatory T cell development and function. Trends Immunol. 2015;36:3–12.
De Rosa V, Galgani M, Porcellini A, Colamatteo A, Santopaolo M, Zuchegna C, et al. Glycolysis controls the induction of human regulatory T cells by modulating the expression of FOXP3 exon 2 splicing variants. Nat Immunol. 2015;16:1174–84.
Gerriets VA, Kishton RJ, Johnson MO, Cohen S, Siska PJ, Nichols AG, et al. Foxp3 and Toll-like receptor signaling balance T(reg) cell anabolic metabolism for suppression. Nat Immunol. 2016;17:1459–66.
Böttcher M, Renner K, Berger R, Mentz K, Thomas S, Cardenas-Conejo ZE, et al. D-2-hydroxyglutarate interferes with HIF-1α stability skewing T-cell metabolism towards oxidative phosphorylation and impairing Th17 polarization. OncoImmunology. 2018;7:e1445454.
Miao Y, Zheng Y, Geng Y, Yang L, Cao N, Dai Y, et al. The role of GLS1-mediated glutaminolysis/2-HG/H3K4me3 and GSH/ROS signals in Th17 responses counteracted by PPARγ agonists. Theranostics. 2021;11:4531–48.
Zhang Y-T, Xing M-L, Fang H-H, Li W-D, Wu L, Chen Z-P. Effects of lactate on metabolism and differentiation of CD4+T cells. Mol Immunol. 2023;154:96–107.
Gu J, Zhou J, Chen Q, Xu X, Gao J, Li X, et al. Tumor metabolite lactate promotes tumorigenesis by modulating MOESIN lactylation and enhancing TGF-β signaling in regulatory T cells. Cell Rep. 2022;39:110986.
Angelin A, Gil-de-Gomez L, Dahiya S, Jiao J, Guo L, Levine MH, et al. Foxp3 reprograms T cell metabolism to function in low-glucose, high-lactate environments. Cell Metab. 2017;25:1282–93.e7.
Watson MJ, Vignali PDA, Mullett SJ, Overacre-Delgoffe AE, Peralta RM, Grebinoski S, et al. Metabolic support of tumour-infiltrating regulatory T cells by lactic acid. Nature. 2021;591:645–51.
London CA, Lodge MP, Abbas AK. Functional responses and costimulator dependence of memory CD4+ T cells. J Immunol. 2000;164:265–72.
Rogers PR, Dubey C, Swain SL. Qualitative changes accompany memory t cell generation: faster, more effective responses at lower doses of antigen. J Immunol. 2000;164:2338–46.
Martín-Leal A, Blanco R, Casas J, Sáez ME, Rodríguez-Bovolenta E, de Rojas I, et al. CCR5 deficiency impairs CD4 T-cell memory responses and antigenic sensitivity through increased ceramide synthesis. EMBO J. 2020;39:e104749.
Kumar R, Ferez M, Swamy M, Arechaga I, Rejas María T, Valpuesta Jose M, et al. Increased sensitivity of antigen-experienced T cells through the enrichment of oligomeric T cell receptor complexes. Immunity. 2011;35:375–87.
Blanco R, Gómez de Cedrón M, Gámez-Reche L, Martín-Leal A, González-Martín A, Lacalle RA, et al. The chemokine receptor CCR5 links memory CD4+ T cell metabolism to T cell antigen receptor nanoclustering. Front Immunol. 2021;12:722320.
O’Sullivan D. The metabolic spectrum of memory T cells. Immunol Cell Biol. 2019;97:636–46.
He J, Shangguan X, Zhou W, Cao Y, Zheng Q, Tu J, et al. Glucose limitation activates AMPK coupled SENP1-Sirt3 signalling in mitochondria for T cell memory development. Nat Commun. 2021;12:4371.
Tan S, Li S, Min Y, Gisterå A, Moruzzi N, Zhang J, et al. Platelet factor 4 enhances CD4+ T effector memory cell responses via Akt-PGC1α-TFAM signaling-mediated mitochondrial biogenesis. J Thrombosis Haemost. 2020;18:2685–700.
Ma EH, Verway MJ, Johnson RM, Roy DG, Steadman M, Hayes S, et al. Metabolic profiling using stable isotope tracing reveals distinct patterns of glucose utilization by physiologically activated CD8(+) T cells. Immunity. 2019;51:856–70.e5.
Sukumar M, Liu J, Ji Y, Subramanian M, Crompton JG, Yu Z, et al. Inhibiting glycolytic metabolism enhances CD8+ T cell memory and antitumor function. J Clin Invest. 2013;123:4479–88.
Wu B, Goronzy JJ, Weyand CM. Metabolic fitness of T cells in autoimmune disease. Immunometabolism. 2020;2:e200017.
Makowski L, Chaib M, Rathmell JC. Immunometabolism: from basic mechanisms to translation. Immunol Rev. 2020;295:5–14.
Guo C, He J, Song X, Tan L, Wang M, Jiang P, et al. Pharmacological properties and derivatives of shikonin—A review in recent years. Pharm Res. 2019;149:104463.
Koga T, Sato T, Furukawa K, Morimoto S, Endo Y, Umeda M, et al. Promotion of calcium/calmodulin-dependent protein kinase 4 by GLUT1-dependent glycolysis in systemic lupus erythematosus. Arthritis Rheumatol. 2019;71:766–72.
Bishop EL, Gudgeon NH, Mackie GM, Chauss D, Roberts J, Tennant DA, et al. 1,25-Dihydroxyvitamin D3 suppresses CD4+ T-cell effector functionality by inhibition of glycolysis. Immunology. 2022;166:299–309.
Tan SY, Kelkar Y, Hadjipanayis A, Shipstone A, Wynn TA, Hall JP. Metformin and 2-deoxyglucose collaboratively suppress human CD4+ T cell effector functions and activation-induced metabolic reprogramming. J Immunol. 2020;205:957–67.
Martins CP, New LA, O’Connor EC, Previte DM, Cargill KR, Tse IL, et al. Glycolysis inhibition induces functional and metabolic exhaustion of CD4+ T cells in type 1 diabetes. Front Immunol. 2021;12:669456.
Lin Y, Xue K, Li Q, Liu Z, Zhu Z, Chen J. et al. Cyclin-Dependent Kinase 7 Promotes Th17/Th1 Cell Differentiation in Psoriasis by Modulating Glycolytic Metabolism. J Invest Dermatol. 2021;141:2656–2667.e11.
Guo Q, Chu Y, Li H, Shi D, Lin L, Lan W. et al. Dickkopf-related protein 3 alters aerobic glycolysis in pancreatic cancer BxPC-3cells, promoting CD4+ T-cell activation and function. Eur J Med Res. 2021;14:93.
Acknowledgements
We would like to thank the other members of the lab for fruitful discussion and feedback during the writing of this manuscript.
Funding
This work was funded by support from the National Natural Science Foundation of China (NSFC 82171787 to LZ, 82371820 to LJ, 32270987 to JZ), from the Natural Science Foundation of Chongqing (CSTB2024NSCQ-KJFZZDX0001to LJ, CSTB2024NSCQ-MSX0622 to JZ) and from Chongqing Natural Science Foundation Innovation Development Joint Fund Project (CSTB2022NSCQ-LZX0066 to LJ).
Author information
Authors and Affiliations
Contributions
LZ and YL performed study concept and design; YL, YZ, JZ wrote the manuscript and created the figures; LZ,JL revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study did not involve human or animal subjects, and thus, no ethical approval was required.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Y., Zhou, Y., Zhang, J. et al. Regulation of CD4 + T cell differentiation and function by glucose metabolism. Genes Immun 26, 287–296 (2025). https://doi.org/10.1038/s41435-025-00340-8
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41435-025-00340-8