Abstract
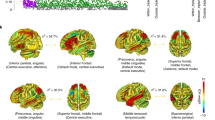
Investigating the genetic underpinnings of functional brain connectivity is essential to understand how genetic variation influences brain health and disease. Here, a mass-univariate approach was adopted to study the genetic architecture of functional brain circuitry (Ntotal = 28,159 subjects) with high spatial resolution (82 brain regions). Common genetic variants explained individual differences in 33% of all 3321 inter-regional functional pathways with 72 significant associations reflecting widespread, pleiotropic effects across the connectome. These associations were mapped to five genes—PAX8, EphA3, SLC39A12, THBS1 and APOE—with known associations with brain phenotypes and which converged in biological processes related to neurodevelopment and cardiovascular and cognitive traits (enrichment minimum p = 3.0 × 10−6 and p = 1.6 × 10−5, respectively). Our findings show that the genetic component of individual differences in functional brain connectivity is largely shared throughout the brain, highlighting the importance of genetic variation in large-scale brain organisation and its relationship with cognitive function and overall health.
Similar content being viewed by others
Data availability
The genome-wide summary statistics data generated in this study can be accessed through https://doi.org/10.5281/zenodo.1842946089. The imaging and genotyping data are protected and are not available due to data privacy. The Source data used to plot each figure are provided with this paper. The following data have been used to perform the analyses on this manuscript: LD reference for LDSC, https://www.internationalgenome.org/category/reference/; RSN Annotation Files, https://surfer.nmr.mgh.harvard.edu/fswiki/CorticalParcellation_Yeo2011; MNI template: https://nist.mni.mcgill.ca/mni-average-brain-305-mri/; FLAMES pathway-naïve feature set: https://zenodo.org/records/10409723; Case-control disease sumstats: https://pgc.unc.edu/for-researchers/download-results/. Source data are provided with this paper.
Code availability
All used software tools and code are publicly available: CATO, http://www.dutchconnectomelab.nl/CATO/; FUMA, http://fuma.ctglab.nl/; MAGMA, https://ctg.cncr.nl/software/magma; LDSC, https://github.com/bulik/ldsc; PLINK, https://www.cog-genomics.org/plink/; FlashPCA2, https://github.com/gabraham/flashpca; FLAMES, https://github.com/Marijn-Schipper/FLAMES; FINEMAP & LDStore2, http://www.christianbenner.com/; coloc R package, https://chr1swallace.github.io/coloc/.
References
Yeo, B. T. et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J. Neurophysiol. 106, 1125–1165 (2011).
Petersen, S. E. & Sporns, O. Brain networks and cognitive architectures. Neuron 88, 207–219 (2015).
van den Heuvel, M. P. & Sporns, O. A cross-disorder connectome landscape of brain dysconnectivity. Nat. Rev. Neurosci. 20, 435–446 (2019).
Fornito, A., Zalesky, A. & Breakspear, M. The connectomics of brain disorders. Nat. Rev. Neurosci. 16, 159–172 (2015).
Gu, Z., Jamison, K. W., Sabuncu, M. R. & Kuceyeski, A. Heritability and interindividual variability of regional structure-function coupling. Nat. Commun. 12, 4894 (2021).
Glahn, D. C. et al. Genetic control over the resting brain. Proc. Natl. Acad. Sci. 107, 1223–1228 (2010).
Reineberg, A. E., Hatoum, A. S., Hewitt, J. K., Banich, M. T. & Friedman, N. P. Genetic and environmental influence on the human functional connectome. Cereb. Cortex 30, 2099–2113 (2020).
Tissink, E. et al. The genetic architectures of functional and structural connectivity properties within cerebral resting-state networks. eNeuro 10, (2023).
Elliott, L. T. et al. Genome-wide association studies of brain imaging phenotypes in UK Biobank. Nature 562, 210–216 (2018).
Zhao, B. et al. Common variants contribute to intrinsic human brain functional networks. Nat. Genet. 54, 508–517 (2022).
Sha, Z., Wager, T. D., Mechelli, A. & He, Y. Common dysfunction of large-scale neurocognitive networks across psychiatric disorders. Biol. Psychiatry 85, 379–388 (2019).
Smith, S. M. et al. An expanded set of genome-wide association studies of brain imaging phenotypes in UK Biobank. Nat. Neurosci. 24, 737–745 (2021).
Brouwer, R. M. et al. Genetic variants associated with longitudinal changes in brain structure across the lifespan. Nat. Neurosci. 25, 421–432 (2022).
Régy, M. et al. The role of dementia in the association between APOE4 and all-cause mortality: pooled analyses of two population-based cohort studies. Lancet Healthy Longev. 5, e422–e430 (2024).
Romero, C. et al. Exploring the genetic overlap between twelve psychiatric disorders. Nat. Genet. 54, 1795–1802 (2022).
Grotzinger, A. D. et al. Genetic architecture of 11 major psychiatric disorders at biobehavioral, functional genomic and molecular genetic levels of analysis. Nat. Genet. 54, 548–559 (2022).
Roelfs, D. et al. Shared genetic architecture between mental health and the brain functional connectome in the UK Biobank. BMC Psychiatry 23, 461 (2023).
van der Meer, D. et al. Understanding the genetic determinants of the brain with MOSTest. Nat. Commun. 11, 3512 (2020).
Tissink, E. P. et al. Abundant pleiotropy across neuroimaging modalities identified through a multivariate genome-wide association study. Nat. Commun. 15, 2655 (2024).
Mekki, Y. et al. The genetic architecture of language functional connectivity. NeuroImage 249, 118795 (2022).
Foo, H. et al. Novel genetic variants associated with brain functional networks in 18,445 adults from the UK Biobank. Sci. Rep. 11, 14633 (2021).
Bell, S., Tozer, D. J. & Markus, H. S. Genome-wide association study of the human brain functional connectome reveals strong vascular component underlying global network efficiency. Sci. Rep. 12, 14938 (2022).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Oberhuber, M. et al. Four functionally distinct regions in the left supramarginal gyrus support word processing. Cereb. Cortex 26, 4212–4226 (2016).
Davey, J. et al. Exploring the role of the posterior middle temporal gyrus in semantic cognition: Integration of anterior temporal lobe with executive processes. Neuroimage 137, 165–177 (2016).
Seoane, S., Modroño, C., González-Mora, J. L. & Janssen, N. Medial temporal lobe contributions to resting-state networks. Brain Struct. Funct. 227, 995–1012 (2022).
Schipper, M. et al. Prioritizing effector genes at trait-associated loci using multimodal evidence. Nat. Genet. 57, 323–333 (2025).
Fernández-Calle, R. et al. APOE in the bullseye of neurodegenerative diseases: impact of the APOE genotype in Alzheimer’s disease pathology and brain diseases. Mol. Neurodegener. 17, 62 (2022).
Watson, H. J. et al. Genome-wide association study identifies eight risk loci and implicates metabo-psychiatric origins for anorexia nervosa. Nat. Genet. 51, 1207–1214 (2019).
Wightman, D. P. et al. A genome-wide association study with 1,126,563 individuals identifies new risk loci for Alzheimer’s disease. Nat. Genet. 53, 1276–1282 (2021).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Xue, A. et al. Unravelling the complex causal effects of substance use behaviours on common diseases. Commun. Med. 4, 43 (2024).
de Leeuw, C., Savage, J., Bucur, I. G., Heskes, T. & Posthuma, D. Understanding the assumptions underlying Mendelian randomization. Eur. J. Hum. Genet. 30, 653–660 (2022).
Petrovski, S., Wang, Q., Heinzen, E. L., Allen, A. S. & Goldstein, D. B. Genic intolerance to functional variation and the interpretation of personal genomes. PLOS Genet. 9, e1003709 (2013).
Lisabeth, E. M., Falivelli, G. & Pasquale, E. B. Eph receptor signaling and ephrins. Cold Spring Harb. Perspect. Biol. 5, a009159 (2013).
Donoghue, M. J. & Rakic, P. Molecular gradients and compartments in the embryonic primate cerebral cortex. Cereb. Cortex 9, 586–600 (1999).
Jayasena, C. S., Flood, W. D. & Koblar, S. A. High EphA3 expressing ophthalmic trigeminal sensory axons are sensitive to ephrin-A5-Fc: Implications for lobe-specific axon guidance. Neuroscience 135, 97–109 (2005).
Wainberg, M. et al. Genetic architecture of the structural connectome. Nat. Commun. 15, 1962 (2024).
Di Palma, T. et al. Characterization of a novel loss-of-function mutation of PAX8 associated with congenital hypothyroidism. Clin. Endocrinol. 73, 808–814 (2010).
Mansouri, A., Chowdhury, K. & Gruss, P. Follicular cells of the thyroid gland require Pax8 gene function. Nat. Genet. 19, 87–90 (1998).
Son, S. M. et al. Thrombospondin-1 prevents amyloid beta-mediated synaptic pathology in Alzheimer’s disease. Neurobiol. Aging 36, 3214–3227 (2015).
Li, Z., Shue, F., Zhao, N., Shinohara, M. & Bu, G. APOE2: protective mechanism and therapeutic implications for Alzheimer’s disease. Mol. Neurodegener. 15, 63 (2020).
Kulminski, A. M., Philipp, I., Shu, L. & Culminskaya, I. Definitive roles of TOMM40-APOE-APOC1 variants in the Alzheimer’s risk. Neurobiol. Aging 110, 122–131 (2022).
Ken-Dror, G., Talmud, P. J., Humphries, S. E. & Drenos, F. APOE/C1/C4/C2 gene cluster genotypes, haplotypes and lipid levels in prospective coronary heart disease risk among UK healthy men. Mol. Med. 16, 389–399 (2010).
Blake, S. M. et al. Thrombospondin-1 binds to ApoER2 and VLDL receptor and functions in postnatal neuronal migration. EMBO J. 27, 3069–3080 (2008).
Sensi, S. L., Paoletti, P., Bush, A. I. & Sekler, I. Zinc in the physiology and pathology of the CNS. Nat. Rev. Neurosci. 10, 780–791 (2009).
Chowanadisai, W., Graham, D. M., Keen, C. L., Rucker, R. B. & Messerli, M. A. Neurulation and neurite extension require the zinc transporter ZIP12 (slc39a12). Proc. Natl. Acad. Sci. 110, 9903–9908 (2013).
Scarr, E. et al. Increased cortical expression of the zinc transporter SLC39A12 suggests a breakdown in zinc cellular homeostasis as part of the pathophysiology of schizophrenia. Npj Schizophr. 2, 1–7 (2016).
Karczewski, K. J. et al. Systematic single-variant and gene-based association testing of thousands of phenotypes in 394,841 UK Biobank exomes. Cell Genom. 2, 100168 (2022).
Friston, K. J. Functional and effective connectivity in neuroimaging: a synthesis. Hum. Brain Mapp. 2, 56–78 (1994).
Faskowitz, J., Betzel, R. F. & Sporns, O. Edges in brain networks: contributions to models of structure and function. Netw. Neurosci. 6, 1–28 (2022).
Foo, H. et al. Genetic influence on ageing-related changes in resting-state brain functional networks in healthy adults: a systematic review. Neurosci. Biobehav. Rev. 113, 98–110 (2020).
van der Meer, D. et al. Boosting schizophrenia genetics by utilizing genetic overlap with brain morphology. Biol. Psychiatry 92, 291–298 (2022).
Cao, H., Zhou, H. & Cannon, T. D. Functional connectome-wide associations of schizophrenia polygenic risk. Mol. Psychiatry 26, 2553–2561 (2021).
Sha, Z., Schijven, D., Fisher, S. E. & Francks, C. Genetic architecture of the white matter connectome of the human brain. Sci. Adv. 9, eadd2870 (2023).
Carnevale, L. et al. Brain functional magnetic resonance imaging highlights altered connections and functional networks in patients with hypertension. Hypertension 76, 1480–1490 (2020).
Hussein, A. et al. The association between resting-state functional magnetic resonance imaging and aortic pulse-wave velocity in healthy adults. Hum. Brain Mapp. 41, 2121–2135 (2020).
Chang, C. et al. Association between heart rate variability and fluctuations in resting-state functional connectivity. NeuroImage 68, 93–104 (2013).
Wei, Y. et al. Associated Genetics and Connectomic Circuitry in Schizophrenia and Bipolar Disorder. Biol. Psychiatry 94, 174–183 (2023).
Sjögren, M. et al. Tau and Aβ42 in cerebrospinal fluid from healthy adults 21–93 years of age: establishment of reference values. Clin. Chem. 47, 1776–1781 (2001).
Slot, R. E. R. et al. Apolipoprotein A1 in cerebrospinal fluid and plasma and progression to Alzheimer’s disease in non-demented elderly. J. Alzheimers Dis. 56, 687–697 (2017).
Yu, M., Sporns, O. & Saykin, A. J. The human connectome in Alzheimer disease—relationship to biomarkers and genetics. Nat. Rev. Neurol. 17, 545–563 (2021).
Fry, A. et al. Comparison of sociodemographic and health-related characteristics of UK Biobank participants with those of the general population. Am. J. Epidemiol. 186, 1026–1034 (2017).
Power, J. D. et al. Methods to detect, characterize, and remove motion artifact in resting state fMRI. NeuroImage 84, https://doi.org/10.1016/j.neuroimage.2013.08.048 (2014).
Liu, T. T. & Falahpour, M. Vigilance effects in resting-state fMRI. Front. Neurosci. 14, (2020).
Birn, R. M. et al. The influence of physiological noise correction on test–retest reliability of resting-state functional connectivity. Brain Connect 4, 511–522 (2014).
Botvinik-Nezer, R. et al. Variability in the analysis of a single neuroimaging dataset by many teams. Nature 582, 84–88 (2020).
Jansen, P. R. et al. Genome-wide meta-analysis of brain volume identifies genomic loci and genes shared with intelligence. Nat. Commun. 11, 5606 (2020).
Nielsen, R., Paul, J. S., Albrechtsen, A. & Song, Y. S. Genotype and SNP calling from next-generation sequencing data. Nat. Rev. Genet. 12, 443–451 (2011).
Smith, S., Alfaro-Almagro, F. & Miller, K. L. UK Biobank Brain Imaging Documentation Version 1.9. (2022).
de Lange, S. C., Helwegen, K. & van den Heuvel, M. P. Structural and functional connectivity reconstruction with CATO—a connectivity analysis TOolbox. NeuroImage 273, 120108 (2023).
Jenkinson, M., Beckmann, C. F., Behrens, T. E. J., Woolrich, M. W. & Smith, S. M. FSL. NeuroImage 62, 782–790 (2012).
Fischl, B. et al. Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain. Neuron 33, 341–355 (2002).
Desikan, R. S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral-based regions of interest. NeuroImage 31, 968–980 (2006).
Power, J. D., Barnes, K. A., Snyder, A. Z., Schlaggar, B. L. & Petersen, S. E. Spurious but systematic correlations in functional connectivity MRI networks arise from subject motion. NeuroImage 59, 2142–2154 (2012).
Alfaro-Almagro, F. et al. Confound modelling in UK Biobank brain imaging. NeuroImage 224, 117002 (2021).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Abraham, G., Qiu, Y. & Inouye, M. FlashPCA2: principal component analysis of Biobank-scale genotype datasets. Bioinformatics 33, 2776–2778 (2017).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLOS Genet. 10, e1004383 (2014).
Benner, C. et al. Prospects of fine-mapping trait-associated genomic regions by using summary statistics from genome-wide association studies. Am. J. Hum. Genet. 101, 539–551 (2017).
Benner, C. et al. FINEMAP: efficient variable selection using summary data from genome-wide association studies. Bioinformatics 32, 1493–1501 (2016).
Weeks, E. M. et al. Leveraging polygenic enrichments of gene features to predict genes underlying complex traits and diseases. Nat. Genet. 55, 1267–1276 (2023).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinforma 27, 1739–1740 (2011).
MacArthur, J. et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 45, D896–D901 (2017).
de Azevedo Pinto Castro Maciel, B. et al. Summary statistics data for ‘The genetic landscape of human functional brain connectivity’ by Maciel et al. (2026). Zenodo https://doi.org/10.5281/zenodo.18429460 (2026).
Acknowledgements
B.A.P.C.M., D.P., C.R., M.P.vd.H., and M.S. were funded by a Dutch Research Council (NWO) Gravitation grant: BRAINSCAPES: A Roadmap from Neurogenetics to Neurobiology (grant no. 024.004.012 [to D.P.]). D.P. and C.d.L. were funded by a European Research Council advanced grant (grant no. ERC-2018-AdG GWAS2FUNC 834057 [to D.P.]). M.P.vd.H. and K.H. were funded by an European Research Council Consolidator grant: CONNECT (grant no. 101001062 [to M.P.vd.H]). M.P.vd.H. was funded by a Dutch Research Council (NWO) VICI grant: BrainDiversity (grant no. VI.C.241.074). J.E.S. is supported by the Dutch Research Council (NWO) VENI Grant 201G-064. This manuscript was submitted as a preprint to medRxiv. This research has been conducted using the UK Biobank resource under application 16406. We thank the numerous participants, researchers, and staff who collected and contributed to the data. We thank SURF for the support in using the National Supercomputers Snellius and LISA. Part of the work in this manuscript was carried out with the support of the SURF Cooperative using grant EINF-14888. We thank Dr Rachel M. Brouwer, Dr Sophie van der Sluis and Ilan Libedinsky for all the feedback. Finally, we would also like to acknowledge Dr Elleke P. Tissink and Dr Siemon C. de Lange, who preprocessed the MRI data.
Author information
Authors and Affiliations
Contributions
B.d.A.P.C.M.: Conceptualisation, Methodology, Formal Analysis, Writing, Visualisation. M.S.: Formal Analysis, Feedback. C.R.: Visualisation, Feedback. C.d.L.: Methodology, Feedback. K.H.: Feedback. D.P.: Funding acquisition, Feedback. J.E.S.: Methodology, Supervision, Feedback. M.P.v.d.H: Conceptualisation, Methodology, Resources, Supervision, Project Administration.
Corresponding author
Ethics declarations
Competing interests
M.P.v.d.H. works as a consultant for Hoffman-La Roche and is part of the editorial team of Wiley Human Brain Mapping; they had no role in this study. The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Maciel, B.d.A., Schipper, M., Romero, C. et al. The genetic landscape of human functional brain connectivity. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69442-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69442-9