Abstract
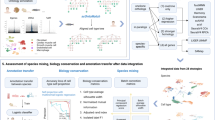
Single-cell RNA-seq (scRNA-seq) data from multiple species present remarkable opportunities to explore cellular origins and evolution. However, integrating and annotating scRNA-seq data across different species remains challenging due to the variations in sequencing techniques, ambiguity of homologous relationships, and limited biological knowledge. To tackle the above challenges, we introduce CAMEX, a heterogeneous Graph Neural Network (GNN) tool that leverages many-to-many homologous relationships for multi-species integration, alignment, and annotation of scRNA-seq data from multiple species. Notably, CAMEX outperforms state-of-the-art methods integration on various cross-species benchmarking datasets (ranging from one to eleven species). Besides, CAMEX facilitates the alignment of diverse species across different developmental stages, significantly enhancing our understanding of organ and organism origins. Furthermore, CAMEX enables the detection of species-specific cell types and marker genes through cell and gene embedding. In short, CAMEX holds the potential to provide invaluable insights into how evolutionary forces operate across different species at single-cell resolution.
Similar content being viewed by others
Data availability
The details of all datasets can be found in Supplementary Data 26. We preprocessed all raw data following the pipeline by Scanpy80 and upload the processed data in h5ad format to Google Driver. This dataset is freely accessible without requiring a password: https://drive.google.com/drive/folders/1rwdjEvWFEFw82a0x2JzMi2jXICbUc5eb?usp=sharing and the dataset can also be available at: https://figshare.com/articles/dataset/Dataset_for_CAMEX/31131808. Source data are provided with this paper.
Code availability
The source codes of CAMEX package, along with code and detailed tutorials for reproducibility, are available at https://github.com/zhanglabtools/CAMEX/ under MIT license, and in Zenodo (https://zenodo.org/records/17991379)81.
References
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Chen, W. et al. A multicenter study benchmarking single-cell RNA sequencing technologies using reference samples. Nat. Biotechnol. 39, 1103–1114 (2021).
Rozenblatt-Rosen, O. et al. Building a high-quality human cell atlas. Nat. Biotechnol. 39, 149–153 (2021).
Han, L. et al. Cell transcriptomic atlas of the non-human primate Macaca fascicularis. Nature 604, 723–731 (2022).
Iram, T. & Consortium, T. M. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Farrell, J. A. et al. Single-cell reconstruction of developmental trajectories during zebrafish embryogenesis. Science 360, eaar3131 (2018).
Tanay, A. & Sebé-Pedrós, A. Evolutionary cell type mapping with single-cell genomics. Trends Genet. 37, 919–932 (2021).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Potter, S. S. Single-cell RNA sequencing for the study of development, physiology and disease. Nat. Rev. Nephrol. 14, 479–492 (2018).
Koonin, E. V. & Galperin, M. Y. Evolutionary Concept in Genetics and Genomics. Sequence–Evolution–Function: Computational Approaches in Comparative Genomics (Kluwer Academic, 2003).
Shafer, M. E. Cross-species analysis of single-cell transcriptomic data. Front. Cell Dev. Biol. 7, 175 (2019).
Wang, J. et al. Tracing cell-type evolution by cross-species comparison of cell atlases. Cell Rep. 34, 108803 (2021).
Wang, R. et al. Construction of a cross-species cell landscape at single-cell level. Nucleic Acids Res. 51, 501–516 (2023).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 1–32 (2020).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with harmony. Nat. Methods 16, 1289–1296 (2019).
Way, G. P. & Greene, C. S. Bayesian deep learning for single-cell analysis. Nat. Methods 15, 1009–1010 (2018).
Tan, Y. & Cahan, P. SingleCellNet: a computational tool to classify single cell RNA-Seq data across platforms and across species. Cell Syst. 9, 207–213. e2 (2019).
Cui, H. et al. scGPT: toward building a foundation model for single-cell multi-omics using generative AI. Nat. Methods 21, 1470–1480 (2024).
Song, Y., Miao, Z., Brazma, A. & Papatheodorou, I. Benchmarking strategies for cross-species integration of single-cell RNA sequencing data. Nat. Commun. 14, 6495 (2023).
Ding, H., Blair, A., Yang, Y. & Stuart, J. M. Biological process activity transformation of single cell gene expression for cross-species alignment. Nat. Commun. 10, 4899 (2019).
Tarashansky, A. J. et al. Mapping single-cell atlases throughout Metazoa unravels cell type evolution. Elife 10, e66747 (2021).
Rosen, Y. et al. Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN. Nat. Methods 21, 1492–1500 (2024).
Liu, X., Shen, Q. & Zhang, S. Cross-species cell-type assignment from single-cell RNA-seq data by a heterogeneous graph neural network. Genome Res. 33, 96–111 (2023).
Lu, L. & Welch, J. D. PyLiger: scalable single-cell multi-omic data integration in Python. Bioinformatics 38, 2946–2948 (2022).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887. e17 (2019).
Hie, B., Bryson, B. & Berger, B. Efficient integration of heterogeneous single-cell transcriptomes using Scanorama. Nat. Biotechnol. 37, 685–691 (2019).
Lotfollahi, M., Wolf, F. A. & Theis, F. J. scGen predicts single-cell perturbation responses. Nat. Methods 16, 715–721 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587. e29 (2021).
Rosen, Y. et al. Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN,. Nat. Methods 21, 1492–1500 (2024).
Büttner, M., Miao, Z., Wolf, F. A., Teichmann, S. A. & Theis, F. J. A test metric for assessing single-cell RNA-seq batch correction. Nat. Methods 16, 43–49 (2019).
Hubert, L. & Arabie, P. Comparing partitions. J. Classification 2, 193–218 (1985).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Xu, C. et al. Probabilistic harmonization and annotation of single-cell transcriptomics data with deep generative models. Mol. Syst. Biol. 17, e9620 (2021).
Cao, Z.-J., Wei, L., Lu, S., Yang, D.-C. & Gao, G. Searching large-scale scRNA-seq databases via unbiased cell embedding with Cell BLAST. Nat. Commun. 11, 3458 (2020).
Guilliams, M. et al. Spatial proteogenomics reveals distinct and evolutionarily conserved hepatic macrophage niches. Cell 185, 379–396. e38 (2022).
Fan, X. et al. Single-cell reconstruction of follicular remodeling in the human adult ovary. Nat. Commun. 10, 3164 (2019).
Muraro, M. J. et al. A single-cell transcriptome atlas of the human pancreas. Cell Syst. 3, 385–394. e3 (2016).
Baron, M. et al. A single-cell transcriptomic map of the human and mouse pancreas reveals inter-and intra-cell population structure. Cell Syst. 3, 346–360. e4 (2016).
Xin, Y. et al. RNA sequencing of single human islet cells reveals type 2 diabetes genes. Cell Metab. 24, 608–615 (2016).
Segerstolpe, Å et al. Single-cell transcriptome profiling of human pancreatic islets in health and type 2 diabetes. Cell Metab. 24, 593–607 (2016).
Lawlor, N. et al. Single-cell transcriptomes identify human islet cell signatures and reveal cell-type–specific expression changes in type 2 diabetes. Genome Res. 27, 208–222 (2017).
Wang, S. et al. Single-cell transcriptomic atlas of primate ovarian aging. Cell 180, 585–600. e19 (2020).
Han, X. et al. Mapping the mouse cell atlas by microwell-seq. Cell 172, 1091–1107. e17 (2018).
Li, J. et al. A single-cell transcriptomic atlas of primate pancreatic islet aging. Natl. Sci. Rev. 8, nwaa127 (2021).
Soumillon, M. et al. Cellular source and mechanisms of high transcriptome complexity in the mammalian testis. Cell Rep. 3, 2179–2190 (2013).
Murat, F. et al. The molecular evolution of spermatogenesis across mammals. Nature 613, 308–316 (2023).
Zhong, H. et al. Benchmarking cross-species single-cell RNA-seq data integration methods: towards a cell type tree of life. Nucleic Acids Res. 53, gkae1316 (2025).
Acién, P. & Acién, M. Disorders of sex development: classification, review, and impact on fertility. J. Clin. Med. 9, 3555 (2020).
Fagerberg, L. et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell. Proteom. 13, 397–406 (2014).
MacLean II, J. A. & Wilkinson, M. F. Gene regulation in spermatogenesis. Curr. Top. Dev. Biol. 71, 131–197 (2005).
O’Donnell, L., Stanton, P. & de Kretser, D. M. Endocrinology of the Male Reproductive System and Spermatogenesis. Endotext [Internet] https://www.ncbi.nlm.nih.gov/books/NBK279031/ (updated 11 January 2017).
Griswold, M. D. Spermatogenesis: the commitment to meiosis. Physiol. Rev. 96, 1–17 (2016).
Cardoso-Moreira, M. et al. Gene expression across mammalian organ development. Nature 571, 505–509 (2019).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Garcia-Alonso, L. et al. Single-cell roadmap of human gonadal development. Nature 607, 540–547 (2022).
Smith, K. K. Early development of the neural plate, neural crest and facial region of marsupials. J. Anat. 199, 121–131 (2001).
Tasic, B. et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78 (2018).
Tosches, M. A. et al. Evolution of pallium, hippocampus, and cortical cell types revealed by single-cell transcriptomics in reptiles. Science 360, 881–888 (2018).
Ma, S. et al. Molecular and cellular evolution of the primate dorsolateral prefrontal cortex. Science 377, eabo7257 (2022).
Verkhratsky, A., Margaret, S, Ho. & Parpura, V. Evolution of neuroglia. Adv. Exp. Med. Biol. 1175, 15–44 (2019).
Li, Z. et al. CD83: activation marker for antigen presenting cells and its therapeutic potential. Front. Immunol. 10, 460131 (2019).
Grosche, L. et al. The CD83 molecule–an important immune checkpoint. Front. Immunol. 11, 721 (2020).
Zhao, K. & Ma, Z. “Comprehensive analysis to identify SPP1 as a prognostic biomarker in cervical cancer. Front. Genet. 12, 732822 (2022).
Yi, X. et al. SPP1 facilitates cell migration and invasion by targeting COL11A1 in lung adenocarcinoma. Cancer Cell Int. 22, 324 (2022).
Klein, C. et al. Neuron navigator 3a regulates liver organogenesis during zebrafish embryogenesis. Development 138, 1935–1945 (2011).
Satani, M. et al. Expression and characterization of human bifunctional peptidylglycine α-amidating monooxygenase. Protein Expr. Purif. 28, 293–302 (2003).
Innan, H. & Kondrashov, F. The evolution of gene duplications: classifying and distinguishing between models. Nat. Rev. Genet. 11, 97–108 (2010).
Passalacqua, M. J. & Gillis, J. Coexpression enhances cross-species integration of single-cell RNA sequencing across diverse plant species. Nat. Plants 10, 1075–1080 (2024).
Zhong, H. et al. Unify: learning cellular evolution with universal multimodal embeddings. https://doi.org/10.1101/2025.09.07.674681 (2025).
Wang, P. et al. scCompass: An integrated cross-species scRNA-seq database for AI-ready. https://doi.org/10.1101/2024.11.12.623138 (2024).
Schlichtkrull, M. et al. Modeling relational data with graph convolutional networks. in The Semantic Web: 15th International Conference, ESWC 2018, Heraklion, Crete, Greece, June 3–7, 2018, Proceedings. 593–607 (Springer, 2018).
Rumelhart, D. E., Hinton, G. E. & Williams, R. J. Learning representations by back-propagating errors. Nature 323, 533–536 (1986).
Diederik, K. Adam: A method for stochastic optimization. https://arxiv.org/abs/1412.6980 (2014).
Paszke, A. et al. Automatic differentiation in PyTorch. in 31st Conference on Neural Information Processing Systems. (NIPS, Long Beach, CA, USA, 2017).
Wang, M. et al. Deep graph library: a graph-centric, highly-performant package for graph neural networks. arXiv preprint arXiv:1909.01315 (2019).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Cao, Z.-J., Wei, L., Lu, S., Yang, D.-C. & Gao, G. Searching large-scale scRNA-seq databases via unbiased cell embedding with Cell BLAST. Nat. Commun. 11, 1–13 (2020).
Howe, K. L. et al. Ensembl 2021. Nucl. Acids Res. 49, D884–D891 (2021).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 1–5 (2018).
Guo, Z.-H. zhanglabtools/CAMEX: Multi-species integration, alignment and annotation of single-cell RNA-seq data with CAMEX (v0.0.1). Zenodo. https://doi.org/10.5281/zenodo.17991379 (2015).
Acknowledgements
This work has been supported by the National Key Research and Development Program of China [No. 2021YFA1302500 to S.Z.], the National Natural Science Foundation of China [Nos. 32341013, 12326614 to S.Z., Nos. 62333018, 62372255, 62432013, W2412087, 62402250, 62433001, U22A2039 to D.S.H.], the CAS Project for Young Scientists in Basic Research [No. YSBR-034 to S.Z.], the Zhejiang Province Vanguard Goose-Leading Initiative (No. 2025C01114 to S.Z.), the Natural Science Foundation of Zhejiang Province [No. LMS25F020001 to D.S.H.], the Key Research and Development Program of Ningbo City [Nos. 2024Z112, 2023Z219, 2023Z226 to D.S.H.], the Yongjiang Talent Project of Ningbo, Yongrencaifa [No. 2024-4 to D.S.H.], and the Basic Research Program Project of the Department of Science and Technology of Guizhou Province [No. ZK2024ZD035 to D.S.H)].
Author information
Authors and Affiliations
Contributions
S.Z. conceived and supervised the project. Z. G. collected the datasets and developed the algorithm. Z. G., D. H. and S.Z. performed the analyses and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Doron Betel and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guo, ZH., Huang, DS. & Zhang, S. Multi-species integration, alignment and annotation of single-cell RNA-seq data with CAMEX. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69696-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69696-3