Abstract
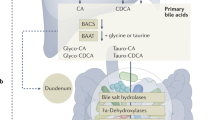
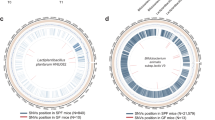
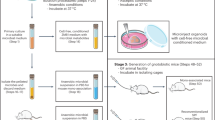
Directed evolution has transformed biomolecular engineering but remains largely untapped for probiotic optimization, despite its immense promise for human health maintenance and disease therapy. Here, we present an in vivo, host-mediated directed-evolution platform that harnesses the gut’s endogenous selective pressures to drive functional enhancement of probiotics. Using Bifidobacterium animalis subsp. lactis as a model, we expose germ-free male mice to stepwise increases in bile-acid stress via a high-fat, high-cholesterol diet. Compared to in vitro evolution, which fails to produce any functional gains, our host-mediated approach yields a variant exhibiting a 77% increase in bile acid metabolism. Multi-omics analysis identifies two critical single-nucleotide variants (SNVs) simultaneously: one in the upstream region of cbh, encoding bile salt hydrolase, and a non-synonymous mutation in mdr, a bile-acid efflux transporter. Functional validation assays confirm that these mutations drive increased corresponding gene expression and enhance substrate binding efficiency. Moreover, to demonstrate its translational potential, we administer the parental and adapted strains daily in a male diet-induced mouse model of non-alcoholic fatty liver disease (NAFLD). We find that the adapted strain significantly improves bile-acid homeostasis, reduces hepatic steatosis, lowers inflammatory and lipid biomarkers, and enhances histological liver health compared to the parental strain. Our work establishes the host gut as a living evolutionary bioreactor for precision engineering of probiotics, enabling targeted phenotypic enhancement in vivo through natural selection. This framework paves the way for personalized, functionally tailored microbiome therapeutics and sets a foundation for next-generation live biotherapeutic agents.
Similar content being viewed by others
Data availability
All the data that support the findings of this study are available within the paper and its Supplementary Data. All third-generation long-read, whole-genome resequencing, transcriptomic (RNA-seq), and metagenomic sequencing data generated in this study have been deposited in the NCBI database under the following BioProject accession numbers: PRJNA1281804, PRJNA1281807, PRJNA1380636, PRJNA1281849, and PRJNA1283298. Metagenomic data for sample H22B656-6 are not obtained due to DNA extraction failure. The predicted structures of the wild-type protein and the N192D mutant have been deposited in Figshare (https://doi.org/10.6084/m9.figshare.31272826). Targeted metabolomics data for bile acids are available in the MetaboLights database under identifier MTBLS13718. Source data are provided with this paper.
References
Beck-Sickinger, A. G. Directed evolution in drug and antibody development: From the Nobel Prize to broad clinical application]. Internist 60, 1014–1020 (2019).
Scott, B. M. et al. Self-tunable engineered yeast probiotics for the treatment of inflammatory bowel disease. Nat. Med. 27, 1212–1222 (2021).
Karpinski, J. et al. Directed evolution of a recombinase that excises the provirus of most HIV-1 primary isolates with high specificity. Nat. Biotechnol. 34, 401–409 (2016).
Chatterjee, R. & Yuan, L. Directed evolution of metabolic pathways. Trends Biotechnol. 24, 28–38 (2006).
Phaneuf, P. V., Gosting, D., Palsson, B. O. & Feist, A. M. ALEdb 1.0: a database of mutations from adaptive laboratory evolution experimentation. Nucleic Acids Res. 47, D1164–D1171 (2019).
Bennett, A. F. & Hughes, B. S. Microbial experimental evolution. Am. J. Physiol. Regul. Integr. Comp. Physiol. 297, R17–R25 (2009).
Sandberg, T. E., Salazar, M. J., Weng, L. L., Palsson, B. O. & Feist, A. M. The emergence of adaptive laboratory evolution as an efficient tool for biological discovery and industrial biotechnology. Metab. Eng. 56, 1–16 (2019).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Gu, H. et al. Physiological mechanism of improved tolerance of Saccharomyces cerevisiae to lignin-derived phenolic acids in lignocellulosic ethanol fermentation by short-term adaptation. Biotechnol. Biofuels 12, 268 (2019).
Li, J. et al. Biofortified tomatoes provide a new route to vitamin D sufficiency. Nat. Plants 8, 611–616 (2022).
Zhang, J. et al. Probiotics maintain the intestinal microbiome homeostasis of the sailors during a long sea voyage. Gut Microbes 11, 930–943 (2020).
Trager, C. et al. A probiotic approach identifies a Treg-centred immunoregulation via modulation of gut microbiota metabolites in people with multiple sclerosis and healthy individuals. EBioMedicine 116, 105743 (2025).
Murray, E. et al. Probiotic consumption during puberty mitigates LPS-induced immune responses and protects against stress-induced depression- and anxiety-like behaviors in adulthood in a sex-specific manner. Brain Behav. Immun. 81, 198–212 (2019).
Zhang, J. P. et al. High-throughput screening of probiotics in fermented foods and their potential application in alleviating alcohol-induced damage. Food Funct. 16, 2564–2576 (2025).
Keshri, J. et al. Phenotypic characterization and draft genome sequence analyses of two novel endospore-forming sporosarcina spp. isolated from Canada Goose (Branta canadensis) feces. Microorganisms 12, 70 (2023).
Zhang, C. et al. A key genetic factor governing arabinan utilization in the gut microbiome alleviates constipation. Cell Host Microbe 31, 1989–2006 e1988 (2023).
Fletcher, E. et al. Evolutionary engineering reveals divergent paths when yeast is adapted to different acidic environments. Metab. Eng. 39, 19–28 (2017).
Tilloy, V., Ortiz-Julien, A. & Dequin, S. Reduction of ethanol yield and improvement of glycerol formation by adaptive evolution of the wine yeast Saccharomyces cerevisiae under hyperosmotic conditions. Appl. Environ. Microbiol. 80, 2623–2632 (2014).
Baym, M. et al. Spatiotemporal microbial evolution on antibiotic landscapes. Science 353, 1147–1151 (2016).
Pan, C. et al. Ameba-inspired strategy enhances probiotic efficacy via prebound nutrient supply. Nat. Commun. 16, 1827 (2025).
Chang, Z. et al. Bacterial immunotherapy leveraging IL-10R hysteresis for both phagocytosis evasion and tumor immunity revitalization. Cell 188, 1842–1857.e1820 (2025).
Lee, J. S. et al. E. coli genetically modified for purine nucleobase release promotes butyrate generation and colonic wound healing during DSS insult. Gut Microbes 17, 2490211 (2025).
Charbonneau, M. R., Isabella, V. M., Li, N. & Kurtz, C. B. Developing a new class of engineered live bacterial therapeutics to treat human diseases. Nat. Commun. 11, 1738 (2020).
Ford, S. A. & King, K. C. In vivo microbial coevolution favors host protection and plastic downregulation of immunity. Mol. Biol. Evol. 38, 1330–1338 (2021).
Huang, S. et al. Candidate probiotic Lactiplantibacillus plantarum HNU082 rapidly and convergently evolves within human, mice, and zebrafish gut but differentially influences the resident microbiome. Microbiome 9, 151 (2021).
Zhao, S. et al. Adaptive evolution within gut microbiomes of healthy people. Cell Host Microbe 25, 656–667.e658 (2019).
Li, W. et al. Comparative genomics of in vitro and in vivo evolution of probiotics reveals energy restriction not the main evolution driving force in short term. Genomics 113, 3373–3380 (2021).
Aydin, O. et al. An integrated analysis of bile acid metabolism in humans with severe obesity. Hepatology 81, 19–31 (2025).
Kim, T. Y. et al. A high-fat diet activates the BAs-FXR axis and triggers cancer-associated fibroblast properties in the colon. Cell Mol. Gastroenterol. Hepatol. 13, 1141–1159 (2022).
Olm, M. R. et al. inStrain profiles population microdiversity from metagenomic data and sensitively detects shared microbial strains. Nat. Biotechnol. 39, 727–736 (2021).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Knight, J. C. Functional implications of genetic variation in non-coding DNA for disease susceptibility and gene regulation. Clin. Sci. 104, 493–501 (2003).
Bustos, A. Y., Font de Valdez, G., Fadda, S. & Taranto, M. P. New insights into bacterial bile resistance mechanisms: the role of bile salt hydrolase and its impact on human health. Food Res. Int. 112, 250–262 (2018).
Funabashi, M. et al. A metabolic pathway for bile acid dehydroxylation by the gut microbiome. Nature 582, 566–570 (2020).
Gueimonde, M., Garrigues, C., van Sinderen, D., de los Reyes-Gavilan, C. G. & Margolles, A. Bile-inducible efflux transporter from Bifidobacterium longum NCC2705, conferring bile resistance. Appl. Environ. Microbiol. 75, 3153–3160 (2009).
Smirnova, E. et al. Metabolic reprogramming of the intestinal microbiome with functional bile acid changes underlie the development of NAFLD. Hepatology 76, 1811–1824 (2022).
Pfeifer, E., Gatgens, C., Polen, T. & Frunzke, J. Adaptive laboratory evolution of Corynebacterium glutamicum towards higher growth rates on glucose minimal medium. Sci. Rep. 7, 16780 (2017).
Wannier, T. M. et al. Adaptive evolution of genomically recoded Escherichia coli. Proc. Natl. Acad. Sci. USA 115, 3090–3095 (2018).
Yu, T. et al. Reprogramming Yeast Metabolism from Alcoholic Fermentation to Lipogenesis. Cell 174, 1549–1558.e1514 (2018).
Li, X., Pei, G., Liu, L., Chen, L. & Zhang, W. Metabolomic analysis and lipid accumulation in a glucose tolerant Crypthecodinium cohnii strain obtained by adaptive laboratory evolution. Bioresour. Technol. 235, 87–95 (2017).
Li, J. et al. Advances in adaptive laboratory evolutionary engineering to microbial breeding. Sheng Wu Gong. Cheng Xue Bao 37, 130–141 (2021).
Jian, X. et al. Microbial microdroplet culture system (MMC): An integrated platform for automated, high-throughput microbial cultivation and adaptive evolution. Biotechnol. Bioeng. 117, 1724–1737 (2020).
Tenaillon, O. et al. The molecular diversity of adaptive convergence. Science 335, 457–461 (2012).
Vento, J. M., Crook, N. & Beisel, C. L. Barriers to genome editing with CRISPR in bacteria. J. Ind. Microbiol. Biotechnol. 46, 1327–1341 (2019).
Nethery, M. A., Hidalgo-Cantabrana, C., Roberts, A. & Barrangou, R. CRISPR-based engineering of phages for in situ bacterial base editing. Proc. Natl. Acad. Sci. USA 119, e2206744119 (2022).
Yan, Q. & Fong, S. S. Challenges and advances for genetic engineering of non-model bacteria and uses in consolidated bioprocessing. Front. Microbiol. 8, 2060 (2017).
Hong, W. et al. An inexpensive anaerobic chamber for the genetic manipulation of strictly anaerobic bacteria. Anaerobe 69, 102349 (2021).
Thompson, D. B. et al. The future of multiplexed eukaryotic genome engineering. ACS Chem. Biol. 13, 313–325 (2018).
Abdelrahman, M., Wei, Z., Rohila, J. S. & Zhao, K. Multiplex genome-editing technologies for revolutionizing plant biology and crop improvement. Front. Plant Sci. 12, 721203 (2021).
Jiang, S. et al. Gut microbiota drives structural variation of exogenous probiotics to enhance colonization. Gut Microbes 17, 2503371 (2025).
Barbaro, M. R. et al. Lactiplantibacillus plantarum (CECT7484 and CECT7485) and Pedioccoccus acidilactici (CECT7483) enhance actin cytoskeleton and CYP1A1 expression restoring epithelial permeability alterations induced by irritable bowel syndrome mediators. Gut Microbes 17, 2452235 (2025).
Ke, X. et al. Synbiotic-driven improvement of metabolic disturbances is associated with changes in the gut microbiome in diet-induced obese mice. Mol. Metab. 22, 96–109 (2019).
Haghikia, A. et al. Dietary fatty acids directly impact central nervous system autoimmunity via the small intestine. Immunity 43, 817–829 (2015).
Shen, S. et al. Understanding the “individual drug reaction” from the perspective of the interaction between probiotics and lovastatin in vitro and in vivo. Microbiome 11, 209 (2023).
Tanaka, H., Hashiba, H., Kok, J. & Mierau, I. Bile salt hydrolase of Bifidobacterium longum-biochemical and genetic characterization. Appl. Environ. Microbiol. 66, 2502–2512 (2000).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Vaser, R., Sovic, I., Nagarajan, N. & Sikic, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, W182–W185 (2007).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Eberhardt, J., Santos-Martins, D., Tillack, A. F. & Forli, S. AutoDock Vina 1.2.0: new docking methods, expanded force field, and Python bindings. J. Chem. Inf. Model 61, 3891–3898 (2021).
Arab, J. P., Karpen, S. J., Dawson, P. A., Arrese, M. & Trauner, M. Bile acids and nonalcoholic fatty liver disease: Molecular insights and therapeutic perspectives. Hepatology 65, 350–362 (2017).
Blanco-Miguez, A. et al. Extending and improving metagenomic taxonomic profiling with uncharacterized species using MetaPhlAn 4. Nat. Biotechnol. 41, 1633–1644 (2023).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 32525049) awarded to J.Z.
Author information
Authors and Affiliations
Contributions
J.Z. and Z.S. conceptualized the research framework and formulated the experimental design. The experimental work and sample preparation were conducted by Z.H., X.L., D.Z., Q.G., L.Z., and S.S. Data analysis was performed by Z.H. in collaboration with Z.S. Z.H., Z.S., S.J., and Z.Z. contributed to figure preparation. The manuscript was drafted by Z.S. and Z.H., and J.Z. and Z.S. provided critical review and substantially refined the final version. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Qi Wang, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, Z., Sun, Z., Liu, X. et al. Harnessing a germ‑free mouse gut bioreactor for directed evolution of probiotics to combat non-alcoholic fatty liver disease. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69823-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69823-0