Abstract
Small cell lung carcinoma (SCLC) is an aggressive neuroendocrine cancer that rapidly develops resistance to platinum-based chemotherapy. A key feature of SCLC is its ability to switch between neuroendocrine (NE) and non-neuroendocrine (non-NE) states, a process linked to therapeutic failure, yet the underlying mechanisms driving this plasticity remain incompletely understood. Here, we show that the translation initiation factor eIF6 is a critical regulator of non-NE transdifferentiation in SCLC. eIF6 expression is consistently upregulated in non-NE states across cell lines, mouse models, and patient samples, accompanied by global remodelling of the translational landscape. Mechanistically, eIF6 dissociates from ribosomes and interacts with the CD104-FAK complex, leading to MAPK pathway activation. Intervening eIF6 suppresses non-NE transdifferentiation and enhances SCLC chemotherapy sensitivity in vitro and in vivo. These findings position the eIF6-CD104-FAK axis as a prognostic marker and therapeutic target, offering a potential strategy to mitigate SCLC resistance.
Similar content being viewed by others
Data availability
The RNA-seq data generated in this study have been deposited in the Gene Expression Omnibus under accession code GSE262597. The mass spectrometry proteomics data generated in this study have been deposited in the ProteomeXchange Consortium via the iProX partner repository under identifier PXD051035. The RNA-seq and proteomics data are fully available without restrictions. Processed data and all data supporting the findings of this study are provided in the Supplementary Information. Source data are provided with this paper.
References
Megyesfalvi, Z. et al. Clinical insights into small cell lung cancer: Tumor heterogeneity, diagnosis, therapy, and future directions. CA Cancer J. Clin. 73, 620–652 (2023).
Rudin, C. M., Brambilla, E., Faivre-Finn, C. & Sage, J. Small-cell lung cancer. Nat. Rev. Dis. Prim. 7, 3 (2021).
Horn, L. et al. First-Line Atezolizumab plus Chemotherapy in Extensive-Stage Small-Cell Lung Cancer. N. Engl. J. Med. 379, 2220–2229 (2018).
Paz-Ares, L. et al. Durvalumab plus platinum-etoposide versus platinum-etoposide in first-line treatment of extensive-stage small-cell lung cancer (CASPIAN): a randomised, controlled, open-label, phase 3 trial. Lancet 394, 1929–1939 (2019).
Tang, X. et al. Metabolism and mRNA translation: a nexus of cancer plasticity. Trends Cell Biol. 35, 562–575 (2025).
Redin, E., Quintanal-Villalonga, A. & Rudin, C. M. Small cell lung cancer profiling: an updated synthesis of subtypes, vulnerabilities, and plasticity. Trends Cancer 10, 935–946 (2024).
Gardner, E. E. et al. Chemosensitive Relapse In Small Cell Lung Cancer Proceeds Through an EZH2-SLFN11 Axis. Cancer Cell 31, 286–299 (2017).
Drapkin, B. J. et al. Genomic anD Functional Fidelity Of Small Cell Lung Cancer Patient-derived Xenografts. Cancer Discov. 8, 600–615 (2018).
Wagner, A. H. et al. Recurrent WNT pathway alterations are frequent in relapsed small cell lung cancer. Nat. Commun. 9, 3787 (2018).
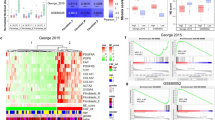
George, J. et al. Evolutionary trajectories of small cell lung cancer under therapy. Nature 627, 880–889 (2024).
Ireland, A. S. et al. MYC Drives Temporal Evolution Of Small Cell Lung Cancer Subtypes By Reprogramming Neuroendocrine Fate. Cancer Cell 38, 60–78 e12 (2020).
Nguyen, E. M. et al. Targeting Lysine-Specific Demethylase 1 Rescues Major Histocompatibility Complex Class I Antigen Presentation and Overcomes Programmed Death-Ligand 1 Blockade Resistance in SCLC. J. Thorac. Oncol. 17, 1014–1031 (2022).
Oser, M. G. et al. The KDM5A/RBP2 histone demethylase represses NOTCH signaling to sustain neuroendocrine differentiation and promote small cell lung cancer tumorigenesis. Genes Dev. 33, 1718–1738 (2019).
Duplaquet, L. et al. KDM6A epigenetically regulates subtype plasticity in small cell lung cancer. Nat. Cell Biol. 25, 1346–1358 (2023).
Chen, H. Y. et al. Regulation of neuroendocrine plasticity by the RNA-binding protein ZFP36L1. Nat. Commun. 13, 4998 (2022).
Fabbri, L., Chakraborty, A., Robert, C. & Vagner, S. The plasticity of mRNA translation during cancer progression and therapy resistance. Nat. Rev. Cancer 21, 558–577 (2021).
Jaako, P. et al. eIF6 rebinding dynamically couples ribosome maturation and translation. Nat. Commun. 13, 1562 (2022).
Gandin, V. et al. Eukaryotic initiation factor 6 is rate-limiting in translation, growth and transformation. Nature 455, 684–688 (2008).
Finch, A. J. et al. Uncoupling of GTP hydrolysis from eIF6 release on the ribosome causes Shwachman-Diamond syndrome. Genes Dev. 25, 917–929 (2011).
George, J. et al. Comprehensive genomic profiles of small cell lung cancer. Nature 524, 47–53 (2015).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009).
Zhang, W. et al. Small cell lung cancer tumors and preclinical models display heterogeneity of neuroendocrine phenotypes. Transl. Lung Cancer Res 7, 32–49 (2018).
Gay, C. M. et al. Patterns of transcription factor programs and immune pathway activation define four major subtypes of SCLC with distinct therapeutic vulnerabilities. Cancer Cell 39, 346–360 e347 (2021).
Canadas, I. et al. Targeting epithelial-to-mesenchymal transition with Met inhibitors reverts chemoresistance in small cell lung cancer. Clin. Cancer Res. 20, 938–950 (2014).
Guo, C. et al. Therapeutic targeting of the mevalonate-geranylgeranyl diphosphate pathway with statins overcomes chemotherapy resistance in small cell lung cancer. Nat. Cancer 3, 614–628 (2022).
Shen, S. et al. An epitranscriptomic mechanism underlies selective mRNA translation remodelling in melanoma persister cells. Nat. Commun. 10, 5713 (2019).
Bohlen, J., Roiuk, M. & Teleman, A. A. Phosphorylation of ribosomal protein S6 differentially affects mRNA translation based on ORF length. Nucleic Acids Res. 49, 13062–13074 (2021).
Leppek, K., Das, R. & Barna, M. Functional 5’ UTR mRNA structures in eukaryotic translation regulation and how to find them. Nat. Rev. Mol. Cell Biol. 19, 158–174 (2018).
Advani, V. M. & Ivanov, P. Translational Control under Stress: Reshaping the Translatome. Bioessays 41, e1900009 (2019).
Jobava, R. et al. Adaptive translational pausing is a hallmark of the cellular response to severe environmental stress. Mol. Cell 81, 4191–4208.e4198 (2021).
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Genuth, N. R. & Barna, M. The discovery of ribosome heterogeneity and its implications for gene regulation and organismal life. Mol. Cell 71, 364–374 (2018).
Ceci, M. et al. Release of eIF6 (p27BBP) from the 60S subunit allows 80S ribosome assembly. Nature 426, 579–584 (2003).
Mollaoglu, G. et al. MYC drives progression of small cell lung cancer to a variant neuroendocrine subtype with vulnerability to aurora kinase inhibition. Cancer Cell 31, 270–285 (2017).
Bankhead, P. et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Nabet, B. Y. et al. Immune heterogeneity in small-cell lung cancer and vulnerability to immune checkpoint blockade. Cancer Cell 42, 429-443.e4 (2024).
Balanis, N. G. et al. Pan-cancer convergence to a small-cell neuroendocrine phenotype that shares susceptibilities with hematological malignancies. Cancer Cell 36, 17–34.e17 (2019).
Canadas, I. et al. Tumor innate immunity primed by specific interferon-stimulated endogenous retroviruses. Nat. Med. 24, 1143–1150 (2018).
Hillary, R. F. & FitzGerald, U. A lifetime of stress: ATF6 in development and homeostasis. J. Biomed. Sci. 25, 48 (2018).
Mello, C. A. et al. Desmoplastic small round cell tumor: a review of main molecular abnormalities and emerging Therapy. Cancers 13, 498 (2021).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Chan, J. M. et al. Signatures of plasticity, metastasis, and immunosuppression in an atlas of human small cell lung cancer. Cancer Cell 39, 1479–1496.e1418 (2021).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Arafeh, R., Shibue, T., Dempster, J. M., Hahn, W. C. & Vazquez, F. The present and future of the Cancer Dependency Map. Nat. Rev. Cancer 25, 59–73 (2025).
Mahadevan, N. R. et al. Intrinsic immunogenicity of small cell lung carcinoma revealed by its cellular plasticity. Cancer Discov. 11, 1952–1969 (2021).
Oughtred, R. et al. The BioGRID database: A comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 30, 187–200 (2021).
Biffo, S. et al. Isolation of a novel beta4 integrin-binding protein (p27(BBP)) highly expressed in epithelial cells. J. Biol. Chem. 272, 30314–30321 (1997).
Sanvito, F. et al. The beta4 integrin interactor p27(BBP/eIF6) is an essential nuclear matrix protein involved in 60S ribosomal subunit assembly. J. Cell Biol. 144, 823–837 (1999).
Shen, S., Girault, I., Malka-Mahieu, H., Robert, C. & Vagner, S. In situ detection of the eIF4F translation initiation complex in mammalian cells and tissues. STAR Protoc. 2, 100621 (2021).
Groft, C. M., Beckmann, R., Sali, A. & Burley, S. K. Crystal structures of ribosome anti-association factor IF6. Nat. Struct. Biol. 7, 1156–1164 (2000).
Jungers, C. F., Elliff, J. M., Masson-Meyers, D. S., Phiel, C. J. & Origanti, S. Regulation of eukaryotic translation initiation factor 6 dynamics through multisite phosphorylation by GSK3. J. Biol. Chem. 295, 12796–12813 (2020).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Tai, Y. L. et al. An EGFR/Src-dependent beta4 integrin/FAK complex contributes to malignancy of breast cancer. Sci. Rep. 5, 16408 (2015).
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M., Arpagaus, S. & Ban, N. Crystal structure of the eukaryotic 60S ribosomal subunit in complex with initiation factor 6. Science 334, 941–948 (2011).
Elliff, J. et al. Dynamic states of eIF6 and SDS variants modulate interactions with uL14 of the 60S ribosomal subunit. Nucleic Acids Res. 51, 1803–1822 (2023).
Keen, A. N. et al. Eukaryotic initiation factor 6 regulates mechanical responses in endothelial cells. J. Cell Biol. 221, e202005213 (2022).
Brina, D., Miluzio, A., Ricciardi, S. & Biffo, S. eIF6 anti-association activity is required for ribosome biogenesis, translational control and tumor progression. Biochim. Biophys. Acta 1849, 830–835 (2015).
Jeffery, C. J. Protein moonlighting: what is it, and why is it important? Philos. Trans. R. Soc. Lond. B. Biol. Sci. 373, 20160523 (2018).
Mazumder, B. et al. Regulated release of L13a from the 60S ribosomal subunit as a mechanism of transcript-specific translational control. Cell 115, 187–198 (2003).
Wan, F. et al. IKKbeta phosphorylation regulates RPS3 nuclear translocation and NF-kappaB function during infection with Escherichia coli strain O157:H7. Nat. Immunol. 12, 335–343 (2011).
Kim, T. S., Kim, H. D. & Kim, J. PKCdelta-dependent functional switch of rpS3 between translation and DNA repair. Biochim Biophys. Acta 1793, 395–405 (2009).
Ray, P. et al. The Saccharomyces cerevisiae 60 S ribosome biogenesis factor Tif6p is regulated by Hrr25p-mediated phosphorylation. J. Biol. Chem. 283, 9681–9691 (2008).
Miluzio, A., Beugnet, A., Volta, V. & Biffo, S. Eukaryotic initiation factor 6 mediates a continuum between 60S ribosome biogenesis and translation. EMBO Rep. 10, 459–465 (2009).
Megyesfalvi, Z. et al. Expression patterns and prognostic relevance of subtype-specific transcription factors in surgically resected small-cell lung cancer: an international multicenter study. J. Pathol. 257, 674–686 (2022).
Sulzmaier, F. J., Jean, C. & Schlaepfer, D. D. FAK in cancer: mechanistic findings and clinical applications. Nat. Rev. Cancer 14, 598–610 (2014).
Aboubakar Nana, F. et al. Increased Expression and activation of FAK in small-cell lung cancer compared to non-small-cell lung cancer. Cancers 11, 1526 (2019).
Aboubakar Nana, F. et al. Therapeutic potential of focal adhesion kinase inhibition in small cell lung cancer. Mol. Cancer Ther. 18, 17–27 (2019).
Carrano, A. C., Eytan, E., Hershko, A. & Pagano, M. SKP2 is required for ubiquitin-mediated degradation of the CDK inhibitor p27. Nat. Cell Biol. 1, 193–199 (1999).
Soria, J. C. et al. A phase I, pharmacokinetic and pharmacodynamic study of GSK2256098, a focal adhesion kinase inhibitor, in patients with advanced solid tumors. Ann. Oncol. 27, 2268–2274 (2016).
Zhang, B. et al. Focal Adhesion Kinase (FAK) inhibition synergizes with KRAS G12C inhibitors in treating cancer through the regulation of the FAK-YAP signaling. Adv. Sci. 8, e2100250 (2021).
Dawson, J. C., Serrels, A., Stupack, D. G., Schlaepfer, D. D. & Frame, M. C. Targeting FAK in anticancer combination therapies. Nat. Rev. Cancer 21, 313–324 (2021).
Calbo, J. et al. A functional role for tumor cell heterogeneity in a mouse model of small cell lung cancer. Cancer Cell 19, 244–256 (2011).
Inoue, Y. et al. Extracellular signal-regulated kinase mediates chromatin rewiring and lineage transformation in lung cancer. Elife 10, e66524 (2021).
Kinbara, K., Goldfinger, L. E., Hansen, M., Chou, F. L. & Ginsberg, M. H. Ras GTPases: integrins’ friends or foes? Nat. Rev. Mol. Cell Biol. 4, 767–776 (2003).
Arang, N. et al. High-throughput chemogenetic drug screening reveals PKC-RhoA/PKN as a targetable signaling vulnerability in GNAQ-driven uveal melanoma. Cell Rep. Med. 4, 101244 (2023).
Sabari, J. K., Lok, B. H., Laird, J. H., Poirier, J. T. & Rudin, C. M. Unravelling the biology of SCLC: implications for therapy. Nat. Rev. Clin. Oncol. 14, 549–561 (2017).
Hu, C. et al. ASCL1 and DLL3 expressions and their clinicopathological implications in surgically resected pure small cell lung cancer: A study of 247 cases from the National Cancer Center of China. Thorac. Cancer 13, 338–345 (2022).
Wei, J. et al. Clinicopathological features and prognostic implications of ASCL1 expression in surgically resected small cell lung cancer. Thorac. Cancer 12, 40–47 (2021).
Augustyn, A. et al. ASCL1 is a lineage oncogene providing therapeutic targets for high-grade neuroendocrine lung cancers. Proc. Natl. Acad. Sci. USA 111, 14788–14793 (2014).
Olsen, R. R. et al. ASCL1 represses a SOX9(+) neural crest stem-like state in small cell lung cancer. Genes Dev. 35, 847–869 (2021).
Woods, L. M. et al. Elevated ASCL1 activity creates de novo regulatory elements associated with neuronal differentiation. BMC Genomics 23, 255 (2022).
Skoulidis, F. et al. Co-occurring genomic alterations define major subsets of KRAS-mutant lung adenocarcinoma with distinct biology, immune profiles, and therapeutic vulnerabilities. Cancer Discov. 5, 860–877 (2015).
Xiao, Z., Zou, Q., Liu, Y. & Yang, X. Genome-wide assessment of differential translations with ribosome profiling data. Nat. Commun. 7, 11194 (2016).
Lin, Y. et al. eIF3 Associates with 80S Ribosomes to Promote Translation Elongation, Mitochondrial Homeostasis, and Muscle Health. Mol. Cell 79, 575–587.e577 (2020).
Li, F. et al. Reanalysis of ribosome profiling datasets reveals a function of rocaglamide A in perturbing the dynamics of translation elongation via eIF4A. Nat. Commun. 14, 553 (2023).
Xiao, Z. et al. De novo annotation and characterization of the translatome with ribosome profiling data. Nucleic Acids Res. 46, e61 (2018).
Zhu, Y., Li, F., Yang, X. & Xiao, Z. De novo identification of actively translated open reading frames with ribosome profiling data. J. Vis. Exp. https://doi.org/10.3791/63366 (2022).
Li, F., Xing, X., Xiao, Z., Xu, G. & Yang, X. RiboMiner: a toolset for mining multi-dimensional features of the translatome with ribosome profiling data. BMC Bioinforma. 21, 340 (2020).
Khajuria, R. K. et al. Ribosome levels selectively regulate translation and lineage commitment in human hematopoiesis. Cell 173, 90–103.e119 (2018).
Wang, D. et al. Schwann cell-derived EVs facilitate dental pulp regeneration through endogenous stem cell recruitment via SDF-1/CXCR4 axis. Acta Biomater. 140, 610–624 (2022).
Killarney, S. T. et al. PKN2 is a dependency of the mesenchymal-like cancer cell state. Cancer Discov. 15, 595–615 (2025).
Acknowledgements
S.S. acknowledges grant support from the National Natural Science Foundation of China (grant no. 82473387) and from the National Clinical Research Center for Geriatrics, West China Hospital, Sichuan University (grant no. Y2022JC002). L. Liu acknowledges financial support ZYGD18021 and ZYJC21002 from the 1.3.5 Project for Disciplines of Excellence, West China Hospital, Sichuan University. M.C. acknowledges grant support from Agence nationale de la recherche of France (grant no. ANR-23-CE14-0023). We thank Li Li, Fei Chen, Chunjuan Bao and Yang Deng from the Institute of Clinical Pathology, West China Hospital of Sichuan University for their technical support with histological staining. We thank Prof. Trudy G Oliver (Duke University, USA) for her generous suggestions for the RPM mouse model.
Author information
Authors and Affiliations
Contributions
H.P., Z.W. and M.W. are co-first authors and contributed equally. S.S. and H.P. designed the study. S.S., H.P., L.L., Z.W., M.W. and M.C. interpreted the data and wrote the original draft of the manuscript. H.P., Z.W., M.W., Z. D., K.L., performed RNA-seq and polysome profiling. H.P., Z.W., and M.W. performed ribosome mass spectrometry. H.P., Z.W. and M.W. performed the in vivo experiment. H.P., Z.D., Y.W., M.C., and Y.D. performed bioinformatics analysis. Y.L. performed protein structure analyses. S.S., H.P. and Z.W. performed patient sample multiplex staining and developed an image analysis pipeline. S.Song., X.Y., L.L. and W.W. performed histopathology analysis. Q.P. and L.Liu. provided administrative support and data resources, collected patient samples. S.S. supervised the overall study, provided strategic oversight and conceived the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interests: S. Shen reports personal fees from Agence nationale de la recherche (France), Krebsliga Schweiz (Switzerland), KWF Kankerbestrijding (Netherlands), Austrian Research Funding, Belgian Foundation against Cancer, Shenzhen Medical Academy of Research and Translation (China), and serving as an Associate Editor for Oncogenesis (Springer Nature, London, UK); M. Cerezo is a CSO for BiPer Therapeutics (Strasbourg, France) and reports personal fees from European Commission. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Pierre Close, Oskar Marin-Bejar and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Peng, H., Wang, Z., Wang, M. et al. Eukaryote initiation factor 6 modulates small-cell lung carcinoma plasticity via the integrin-FAK signaling axis. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69899-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69899-8