Abstract
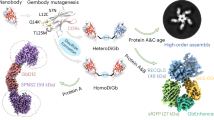
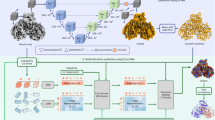
Cryo-electron microscopy (cryo-EM) single-particle analysis faces significant challenges in resolving the structures of small proteins due to low signal-to-noise ratios and insufficient structural features. Here, we present Trimbody, a simple yet robust method that leverages rigid AI-designed scaffolds to overcome these limitations. Trimbody comprises two components: a trimeric scaffold (H3-PrAC-5350A) and a reformatted nanobody fusion (Nb-TAIL). The de novo 3-helix bundle (H3) and TAIL domains, engineered via RFdiffusion and ProteinMPNN, stabilize the interactions between Nbs and the scaffold and enhance the overall rigidity of the system, thereby enabling high-resolution cryo-EM analysis. Using Trimbody, we resolve the atomic structures of four sub-50 kDa test proteins: human Gal10 (2.62 Å), Aequorea coerulescens GFP (2.29 Å), the IgV domain of human Nectin4 (2.43 Å), and membrane protein Escherichia coli LacY (2.50 Å), demonstrating the method’s versatility. Trimbody’s simplicity, cost-effectiveness, and compatibility with standard cryo-EM workflows position it as a universal tool for structural studies of small proteins, further advancing nanobody-based drug development and life science research.
Similar content being viewed by others
Data availability
The cryo-EM density maps generated in this study have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-64015 (hGal10/iTrimbody complex), EMD-63989 (AcGFP/Trimbody complex), EMD-64047 (IgV domain of hNectin4/Trimbody complex), and EMD-67094 (EcLacY10Mu/Trimbody complex). The corresponding atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 9UBR (hGal10/iTrimbody complex), 9UAQ (AcGFP/Trimbody complex), 9UCL (IgV domain of hNectin4/Trimbody complex), and 9XOU (EcLacY10Mu/Trimbody complex). All other published PDB codes cited in this paper are 3OGO, 4JJH, 4FRW, 5GXB, 2V8N, 6I2G, 4W6W, 5C1M, 6P6F, 1WA3, 7RXC, 1HDK and 3LVA. Other data supporting the findings of this study are available within the article and its Supplementary Information. Source data are also provided with this paper. Source data are provided with this paper.
References
Henderson, R. The potential and limitations of neutrons, electrons and X-rays for atomic resolution microscopy of unstained biological molecules. Q Rev. Biophys. 28, 171–193 (1995).
Herzik, M. A. Jr., Wu, M. & Lander, G. C. High-resolution structure determination of sub-100 kDa complexes using conventional cryo-EM. Nat. Commun. 10, 1032 (2019).
Fan, X. et al. Single particle cryo-EM reconstruction of 52 kDa streptavidin at 3.2 Angstrom resolution. Nat. Commun. 10, 2386 (2019).
Yonekura, K., Braunfeld, M. B., Maki-Yonekura, S. & Agard, D. A. Electron energy filtering significantly improves amplitude contrast of frozen-hydrated protein at 300kV. J. Struct. Biol. 156, 524–536 (2006).
Himes, B. & Grigorieff, N. Cryo-TEM simulations of amorphous radiation-sensitive samples using multislice wave propagation. IUCrJ 8, 943–953 (2021).
Lander, G. C. & Glaeser, R. M. Conquer by cryo-EM without physically dividing. Biochem Soc. Trans. 49, 2287–2298 (2021).
Wentinck, K., Gogou, C. & Meijer, D. H. Putting on molecular weight: enabling cryo-EM structure determination of sub-100-kDa proteins. Curr. Res Struct. Biol. 4, 332–337 (2022).
Binz, H. K., Stumpp, M. T., Forrer, P., Amstutz, P. & Pluckthun, A. Designing repeat proteins: well-expressed, soluble and stable proteins from combinatorial libraries of consensus ankyrin repeat proteins. J. Mol. Biol. 332, 489–503 (2003).
Liu, Y., Gonen, S., Gonen, T. & Yeates, T. O. Near-atomic cryo-EM imaging of a small protein displayed on a designed scaffolding system. Proc. Natl. Acad. Sci. USA 115, 3362–3367 (2018).
Liu, Y., Huynh, D. T. & Yeates, T. O. A 3.8 A resolution cryo-EM structure of a small protein bound to an imaging scaffold. Nat. Commun. 10, 1864 (2019).
Castells-Graells, R. et al. Cryo-EM structure determination of small therapeutic protein targets at 3 A-resolution using a rigid imaging scaffold. Proc. Natl. Acad. Sci. USA 120, e2305494120 (2023).
Wu, S. et al. Fabs enable single-particle cryoEM studies of small proteins. Structure 20, 582–592 (2012).
Lee, Y. et al. Cryo-EM structure of the human L-type amino acid transporter 1 in complex with glycoprotein CD98hc. Nat. Struct. Mol. Biol. 26, 510–517 (2019).
Bloch, J. S. et al. Structure and mechanism of the ER-based glucosyltransferase ALG6. Nature 579, 443–447 (2020).
Kim, J. et al. Structure and drug resistance of the Plasmodium falciparum transporter PfCRT. Nature 576, 315–320 (2019).
Bloch, J.S. et al. Development of a universal nanobody-binding Fab module for fiducial-assisted cryo-EM studies of membrane proteins. Proc. Natl Acad. Sci. USA 118, e2115435118 (2021).
Uchanski, T. et al. Megabodies expand the nanobody toolkit for protein structure determination by single-particle cryo-EM. Nat. Methods 18, 60–68 (2021).
Coupland, C. E. et al. Structure, mechanism, and inhibition of Hedgehog acyltransferase. Mol. Cell 81, 5025–5038.e5010 (2021).
Goutam, K., Ielasi, F. S., Pardon, E., Steyaert, J. & Reyes, N. Structural basis of sodium-dependent bile salt uptake into the liver. Nature 606, 1015–1020 (2022).
Wu, X. & Rapoport, T.A. Cryo-EM structure determination of small proteins by nanobody-binding scaffolds (Legobodies). Proc. Natl. Acad. Sci. USA 118, e2115001118 (2021).
Watson, J. L. et al. De novo design of protein structure and function with RFdiffusion. Nature 620, 1089–1100 (2023).
Dauparas, J. et al. Robust deep learning-based protein sequence design using ProteinMPNN. Science 378, 49–56 (2022).
Frenken, L. G. et al. Isolation of antigen-specific llama VHH antibody fragments and their high-level secretion by Saccharomyces cerevisiae. J. Biotechnol. 78, 11–21 (2000).
Fridy, P. C., Thompson, M. K., Ketaren, N. E. & Rout, M. P. Engineered high-affinity nanobodies recognizing staphylococcal Protein A and suitable for native isolation of protein complexes. Anal. Biochem 477, 92–94 (2015).
Bale, J. B. et al. Accurate design of megadalton-scale two-component icosahedral protein complexes. Science 353, 389–394 (2016).
Brouwer, P. J. M. et al. Enhancing and shaping the immunogenicity of native-like HIV-1 envelope trimers with a two-component protein nanoparticle. Nat. Commun. 10, 4272 (2019).
Brouwer, P. J. M. et al. Lassa virus glycoprotein nanoparticles elicit neutralizing antibody responses and protection. Cell Host Microbe 30, 1759–1772.e1712 (2022).
Fullerton, S. W. et al. Mechanism of the class I KDPG aldolase. Bioorg. Med Chem. 14, 3002–3010 (2006).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pletneva, N. V. et al. Structural evidence for a dehydrated intermediate in green fluorescent protein chromophore biosynthesis. J. Biol. Chem. 285, 15978–15984 (2010).
Jain, R. K., Joyce, P. B., Molinete, M., Halban, P. A. & Gorr, S. U. Oligomerization of green fluorescent protein in the secretory pathway of endocrine cells. Biochem J. 360, 645–649 (2001).
Kubala, M. H., Kovtun, O., Alexandrov, K. & Collins, B. M. Structural and thermodynamic analysis of the GFP:GFP-nanobody complex. Protein Sci. 19, 2389–2401 (2010).
Kubach, J. et al. Human CD4+CD25+ regulatory T cells: proteome analysis identifies galectin-10 as a novel marker essential for their anergy and suppressive function. Blood 110, 1550–1558 (2007).
Chatterjee, S., Sinha, S. & Kundu, C. N. Nectin cell adhesion molecule-4 (NECTIN-4): a potential target for cancer therapy. Eur. J. Pharm. 911, 174516 (2021).
Tomiyama, E. et al. Expression of nectin-4 and PD-L1 in upper tract urothelial carcinoma. Int. J. Mol. Sci. 21, 5390 (2020).
Guan, L. & Kaback, H. R. Lessons from lactose permease. Annu Rev. Biophys. Biomol. Struct. 35, 67–91 (2006).
Guan, L., Mirza, O., Verner, G., Iwata, S. & Kaback, H. R. Structural determination of wild-type lactose permease. Proc. Natl. Acad. Sci. USA 104, 15294–15298 (2007).
Kumar, H. et al. Structure of sugar-bound LacY. Proc. Natl. Acad. Sci. USA 111, 1784–1788 (2014).
Smirnova, I. et al. Outward-facing conformers of LacY stabilized by nanobodies. Proc. Natl. Acad. Sci. USA 111, 18548–18553 (2014).
Chaptal, V. et al. Crystal structure of lactose permease in complex with an affinity inactivator yields unique insight into sugar recognition. Proc. Natl. Acad. Sci. USA 108, 9361–9366 (2011).
Gotzke, H. et al. The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications. Nat. Commun. 10, 4403 (2019).
Clift, D., So, C., McEwan, W. A., James, L. C. & Schuh, M. Acute and rapid degradation of endogenous proteins by trim-away. Nat. Protoc. 13, 2149–2175 (2018).
Chen, G. et al. A promising intracellular protein-degradation strategy: TRIMbody-away technique based on nanobody fragment. Biomolecules 11, 1512 (2021).
Wu, X. et al. Structural basis of ER-associated protein degradation mediated by the Hrd1 ubiquitin ligase complex. Science 368, eaaz2449 (2020).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgments
We thank the staff at the Biomedical Research Center for Structural Analysis, Shandong University, for technical assistance during cryo-EM data collection. We also thank all staff members of the Translational Medicine Core Facility of the Advanced Medical Research Institute (AMRI), Shandong University. This work was supported by the National Key Research and Development Program of China (No. 7100, Z.L.), and the National Natural Science Foundation of China (32571436 and 32171207, W.W.; 22377068, X.H.). Additional support was provided by the following: Shandong Provincial Natural Foundation (ZR2024MC199, W.W.), Taishan Scholars Program for Young Experts of Shandong Province (tsqn202408004, W.W.), the Joint Research Grant of Shandong Provincial Third Hospital and Shandong University (GYY202405, W.W.), the Instrument Improvement Fund of Shandong University Public Technology Platform (ts20230203, W.W.), the Innovation Research and Development Special Funds of the Municipality-province-ministry Co-constructed (GJZX-HYSW-2024-02, Z.L.), and the Science and Technology Planning Project of Fujian Province (2022L3022, Z.L.).
Author information
Authors and Affiliations
Contributions
J.S. prepared vectors and protein samples for scaffolds, nanobodies, and target proteins with assistance from Y.L., X.Z., Y.H., Z.Z., and Y.G.; L.Q. and J.S. collected cryo-EM datasets and performed structure determination with assistance from D.H. and M.G.; Z.L. conducted nanobody screening; J.S., X.H., H.F., and W.W. designed nanobody-binding scaffolds and performed computational optimization; J.S. and W.W. built structural models and conducted subsequent analysis; Figures were prepared, and the manuscript was written by W.W. and J.S., with W.W. supervising the entire project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interests: Shandong University has filed patent application No. 202512010140.8 relating to the AI-designed scaffold protein constructs (Trimbody) described in this study. W.W., Z.L., and J.S. are listed as inventors on this application. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Alex De Marco, Jan Steyaert, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Song, J., Qi, L., Li, Y. et al. Trimbody with rigid AI-designed scaffolds enables atomic-resolution cryo-EM structure determination of small proteins. Nat Commun (2026). https://doi.org/10.1038/s41467-026-69941-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-69941-9