Abstract
Protists are less studied for their role and diversity in ecosystems. Notably, protists have played and still play an important role in microbialites. Microbialites, or lithified microbial mats, represent the oldest evidence of fossil biofilms (~3.5 Gyr). Modern microbialites may offer a unique proxy to study the potential role of protists within a geological context. We examined protist diversity in freshwater (Kelly and Pavilion Lake in British Columbia, Canada) and marine (Highborne Cay, Bahamas) to hypersaline (Shark Bay, Australia) microbialites to decipher their geomicrobiological role. The freshwater microbialite communities were clearly distinct from their marine and hypersaline counterparts. Chlorophytes had higher numerical abundance in freshwater microbialites; whereas pennate diatoms dominated numerically in marine microbialites. Despite the differences, protists across ecosystems may have adopted similar roles and functions. We suggest a consistent biogeochemical role of protists across microbialites globally; but that salinity may shape protist composition and evolution in these ecosystems.
Similar content being viewed by others
Introduction
Microbialites are important geobiological signposts as the most abundant evidence of Precambrian life1,2,3. They are fossil biofilms that form through lithification of microbial mats4. The entire microbial mat community, including both prokaryotes and eukaryotes, alter the carbonate saturation index through their metabolic activities, and produce an organic matrix (so called exopolymeric substances, or EPS) that forms the mineral nucleation site5,6,7. Understanding the composition and ecology of microbialites is important to interpret their formation through geologic time1,8,9. Eukaryotes in extant microbialites are low in abundance (<1%), when considering either physical counts10 or sequence abundance11,12,13,14 but are thought to have played and still play an important role in the development of microbialites.
Stromatolites, microbialites with a layered internal fabric, represent the oldest form of microbialites, dating back ~3.5 Gy15. Their fossilized forms potentially provide the earliest evidence of life8,16. Eukaryotes are thought to have evolved in microbial mats or stromatolites3,7,17 and, importantly, played a pivotal role in the first transition from layered stromatolites to clotted thrombolites18, either by using these as a holdfast in the intertidal zone or by grazing pressure8. It has been generally thought that microbialites are rare environments within the contemporary biosphere, even though they have a cosmopolitan distribution19,20,21. The commonly accepted thesis is that due to eukaryotic pressure, mentioned above, present day microbialites are typically only thriving in “extreme” environments7,19. Thus, extant microbialites are found in diverse, extreme environments, with alkaline water (pH > 7.5, carbonate alkalinity > 120 mg CaCO3 L−1) containing high concentrations of free divalent cations (Ca2+ or Mg2+) and limited nutrients (N, P), where photosynthetically active radiation can penetrate to the bottom6. The variety of ecosystems where contemporary microbialites are found include hot springs, lakes, coastal oceans, hypersaline lakes and tropical lagoons5,19,21,22,23.
Despite the hypothesized role in their decline, eukaryotes are vital to healthy, functional modern microbialite communities. Protists have the potential to significantly impact microbialite structures through their photosynthetic activities, by grazing on bacterial communities, and through bioturbation9,11,24,25,26. Furthermore, several eukaryotic taxa have a potential role in the accretion of microbialites2. Photoautotrophic protists aid in the carbonate mineral formation by increasing the alkalinity during photosynthesis, and grazing by heterotrophic eukaryotes may change the prokaryotic community, ultimately altering the overall impact of microbial metabolic processes on the carbonate saturation index5. Metazoans, Stramenopiles, Alveolata, Amoebozoa, Chlorophyta, and Rhodophyta were metabolically active in stromatolites and thrombolites in both Highborne Cay and Shark Bay27. Green algae (predominantly a single Chlorophyte OTU) and diatoms were the predominant protistan oxygenic photosynthesizers in the microbialites of Lake Alchichica, a saline (i.e., 8–13 psu), alkaline crater lake in Mexico11,28. Heterotrophic protists including basal Fungi, Choanoflagellida, Nucleariida, Amoebozoa, Alveolata and Stramenopiles were also present in this lake11.
Negatively charged functional groups in EPS concentrate calcium ions and form the nucleation site for carbonate minerals. EPS are abundant in extant microbialites and support sediment stability that resist erosion, retain critical nutrients within the community, allow for cell-to-cell communication and trap viruses29. Many protists, both photosynthetic and heterotrophic, produce copious amounts of EPS29,30, and thus play a pivotal role in microbialite formation. Diatom-rich subspherical aggregates filled with EPS have been proposed as a major contributor of carbonate precipitation in the hypersaline Laguna Negra (Argentina) microbial mats31. Once lithified structures are formed, protists are capable of altering the fabric (internal structure) of the microbialite: diatoms from the Fragilarophyceae and Bacillariophyceae classes form biofilm networks, which can trap sediment grains and eventually constitute a stromatolite framework enhancing the cyanobacteria and heterotrophic prokaryotes32,33 and reticulopodia from naked or thecate foraminifera and amoebae were found to alter laminated fabrics of microbialites in Shark Bay, Australia, Highborne Cay, Bahamas, and Fayetteville Green Lakes, NY9,34.
Clearly, a wide variety of protists exists in extant freshwater, marine and hypersaline microbialites where they contribute to the ecology and geomicrobiology of these systems. The most critical roles of protists in microbialite formation can be summed up as follows6,18: 1) Changing the carbonate saturation index by contributing to photosynthesis, and through grazing on prokaryotes, by changing the community composition and consequently the sum of the metabolisms; 2) Producing EPS by which the number of sites for binding of cations and nucleation of carbonate minerals increases; 3) Altering the internal fabric through bioturbation and predation using reticulopodia (Fig. 1).
Two key components of microbialite formation are 1) changes in the saturation index (alkalinity) due to changes in microbial metabolisms (see refs. 6,66 for details) and, 2) production exopolymeric substances (EPS) that initially bind Ca2+ and then act as nucleation site for carbonates (see refs. 5,6). Once formed, microbial internal fabrics can be altered by grazing, bioturbation and formation of EPS networks by protists (3). Photosynthetic and heterotrophic (bacterivorous) protists sometimes have different impacts on these processes.
External factors such as location and physicochemical conditions (e.g., salinity, light, nutrients) may drive changes in community composition of microbialites, however the functional capabilities (e.g., to construct a microbialite) of these distinct communities remain mostly consistent33,35,36. Salinity and light availability can be major drivers of microbialite community composition37,38. Notably the former likely impacts lithification rates5 by inhibiting the growth or metabolic activity of certain groups of microbes37,38,39,40,41. Several bacterial diversity studies along spatial or temporal physicochemical gradients concluded that not only is salinity a main determinant of prokaryotic and eukaryotic community composition42,43,44, but it also provokes large-scale changes in the metabolic potential of the entire community7,37,40,45,46. Comparing the eukaryotic community assemblages of saline and freshwater microbialites can potentially uncover the underlying taxonomic similarities and differences in these functionally convergent communities.
Here we examined the eukaryotic diversity of freshwater microbialites (located in Kelly Lake and Pavilion Lake; Southwestern BC, Canada) in the context of other microbialites from the marine (Highborne Cay, Bahamas) and hypersaline (Shark Bay, Australia) environments. To this end, 18S rRNA gene metabarcoding was performed to characterize the microbial communities of the freshwater sites. Both Kelly and Pavilion Lakes have microbialites that form across depths, to account for this we performed sampling across a depth gradient. These communities were analyzed in a meta-analysis incorporating the publicly available 18S rRNA gene data from Highborne Cay and Shark Bay, which represent saline microbialite communities27. Our results highlight the similarities and differences within the metazoan and protist communities of freshwater vs saltwater microbialites providing further evidence for their importance in these ancient systems.
Results and discussion
The Euk1A forward primer used to amplify the 18S rRNA gene in the Kelly Lake and Pavilion Lake and the TAR-EUK-454F primer used for Shark Bay and Highborne Cay had comparable taxonomic coverage according to an in silico test against the PR2 database (Supplementary Table 1). Only the Opisthokonta supergroup, which includes the metazoans, had slightly lower coverage (~30% less, but still above 50%) with TAR-EUK-454F. An analysis of the protist communities (protists hereby referring to all ASVs not classified as metazoans or embryophytes) from each microbialite location showed that Highborne Cay had the most diverse and significantly different community compared to the other three locations according to Shannon–Weiner Diversity Indices. The other locations did not show a significant difference in Shannon–Weiner alpha diversity between locations, highlighting consistency in Shannon–Weiner alpha diversity across three fairly distant microbialite locations (Fig. 2A). Grouping the locations based on salinity revealed a significantly higher (padj = 0.005) Shannon–Weiner alpha diversity in the saltwater vs freshwater microbialites (Fig. 2B). These results support the hypothesis that salinity plays a major role in structuring microbialite communities across space. Examining the beta diversity across the dataset, we found that both location (ANOSIM; 1000 permutations, R = 0.5278, p = 0.001) and salinity (ANOSIM; 1000 permutations, R = 0.669, p = 0.001) were major drivers of dissimilarity (Fig. 2C). However, primer-bias may also be playing a role in the dissimilarities as it was also shown to be a significant driver of dissimilarity in our dataset (ANOSIM; 1000 permutations, R = 0.669, p = 0.001). As the forward primers used for the analyses had comparable coverage across the major eukaryotic Supergroups (Supplementary Table 1), a compositional analysis of the community can still be performed above the ASV level.
A Shannon–Weiner Alpha Diversity measurements for the four microbialites communities. B Shannon–Weiner Alpha Diversity measurements of the combined freshwater communities (Kelly & Pavilion Lake) and the combined marine communities (Highborne Cay & Shark Bay) showing a significant (P = 0.0053) difference in community diversity. C Aitchison Distance PCA of the microbialite communities included in this study with ellipses showing the 95% confidence interval for each location.
Comparing the compositional makeup of the microbialite communities in this study, we see stark/clear differences between the freshwater and saline locations. The microbialite protist communities at the freshwater locations were consistently dominated by Phaeophyceae, Ulvales-relatives, and Cladophorales, which represented a contrast to the marine and hypersaline microbialite protist communities from Highborne Cay and Shark Bay, respectively, which were dominated primarily by Raphid-pennate and Araphid-pennate diatoms (Fig. 3A). A core microbiome analysis which identified genera which were found in at least 75% of samples and had a relative abundance above 1% showed no overlap between marine and freshwater microbialites (Fig. 3B). Thirty-one percent of genera in the dataset were found as core members of the freshwater microbialite community including Bodanella spp., Epithemia spp., Desmochloris spp., Notosolenus spp., Stentor spp. among others. Core members of the marine microbialite community were Trachelocercidae spp., Rhapid-pennate spp., Labrinthulaceae ANT10 spp., Navicula spp., Amphora spp., Monhystrella spp., and Dinophyceae spp (Fig. 3B). Diatoms are key players in microbialite accretion by increasing alkalinity, producing EPS and potentially altering internal fabric6,9,27,33,47, so their high abundance in these saline environments can potentially contribute to these roles. Stalky diatoms present in Highborne Cay stromatolites trap sediment particles33. The two saline locations also showed a high abundance of Pinguiochrysidaceae and Labyrinthulaceae, which were nearly absent in the freshwater locations27. These two protist taxa are known to exist predominantly in marine environments. Some members of the Pinguiochrysidaceae precipitate CaCO3 and are likely capable of disrupting internal microbialite fabric48. Labyrinthulaceae inhabit multiple environments including seagrasses and produce polymers, but could also contribute to carbohydrate degradation49, including parts of EPS. Likewise, the freshwater locations both had Tokophryidae ciliates, which were completely absent in the saltwater locations. Tokophryidae ciliates have been described as epibionts on lacustrine copepods50, so they may not be microbialite-specific associates, but rather associates of the crustaceans which live on the microbialites. However, free-living Tokopphryideaes are bacterivorous and could graze on the microbialite-associated prokaryotes, potentially changing the biofilm community51.
A Bubble plot depicting the relative abundance (mean ± SD) of eukaryotic communities of microbialites from the sampling locations. The proportion of protist to metazoan reads is also shown. B Core genera with a minimum relative abundance of 1% and prevalence in samples of at least 75% found between the freshwater and marine microbialite communities. C Weighted UniFracs of the eukaryotic communities of the samples used in this study, showing the clear delineation between marine (blue) and freshwater (green) microbialite communities. D Analysis of Compositions of Microbiomes with Bias Correction (ANCOM-BC) controlling for False Discovery Rates depicting the significantly (P < 0.05) differentially abundant eukaryotic taxa between marine vs freshwater microbialite communities.
An ANCOM-BC analysis revealed the significantly differentially abundant taxa and their log-fold differences in abundance between the freshwater and marine microbialite communities. Marine and hypersaline microbialites were significantly dominated by diatoms, specifically Araphid-pennate (~4 log folds more abundant; padj = 0.002) and Raphid-pennates (~2 log folds more abundant; padj = 0.029). (Fig. 3D). Diatoms fulfill a critical role in the production of cohesive sediments, notably in the marine environment52. Previous studies in Highborne Cay and Shark Bay did note a high relative abundance of diatoms present in the microbialite communities, however, the varied abundance of each diatom clade was not clear27. Although a key role for diatoms in accretion of microbialites and microbialite-forming mats has long been accepted2,53,54,55, their contribution to microbialite accretion is controversial both in Shark Bay56,57 and Highborne Cay locations35,57,58. Nevertheless, diatoms are major EPS producers and inevitably play a role in microbialite ecology.
A recent study on the saline Lake Alchichica microbialite community in Mexico showed the high abundance of Chlorophytes and Ochrophyta within the microbialites59. Some taxa that exhibited a similar distribution across all our locations include: Thoracosphaeraceae, Pseudokeronopsidae, Holostichidae, and Oxytrichidae. A weighted UniFrac was constructed to compare the microbialite communities of the locations in a phylogenetic context and showed that freshwater microbialites from Kelly Lake and Pavilion Lake clustered distinctly away from the marine microbialites of Shark Bay and Highborne Cay, no matter whether metazoans were included in the unifrac analysis or not, further supporting the notion of salinity as a driving factor in microbialite microbial diversity (Fig. 3C). As for the metazoans, the freshwater locations showed a strong dominance of arthropods within their microbialite communities with Insecta, Ostracoda, and Maxillopoda making up most of the metazoan community composition and core microbiome (Fig. 3A, B). In contrast, the saltwater metazoan communities were dominated by Nematoda, specifically Chromadorea. The relative abundance of metazoan reads to protists reads did not significantly influence the distribution patterns present in each of the locations (Fig. 3A). The ecological role of these metazoans is likely similar in both the freshwater and saline microbialites, as they rely for part of their food source on microbes60. Overall, the consistency of these protist community composition patterns across distant locations in our dataset emphasizes the core microbial members, which are consistently associated with microbialites belonging to fresh and saltwater, despite other contrasting environmental variables, such temperature, light regime and nutrient availability.
Ulvales-relatives (Chlorophyta) were over ~5 log folds more abundant in the freshwater locations than the marine microbialite locations (Fig. 3D; padj = 3.74 × 10−7). Chlorophytes, specifically the filamentous green algae Gongrosira sp., along with cyanobacteria are associated with calcite precipitation in Pavilion Lake microbialites61 and likely contribute to this by photosynthetic activity, which increases local alkalinity, but also producing organic carbon that fuels aerobic and anaerobic heterotrophic prokaryotes62. Interestingly, a recent phylogenetic analysis concluded that Gongrosira fluminesis which was previously identified as Chaetophorales actually belonged to Ulvales63. Carbonate-precipitating Gongrosira species have been found associated with freshwater microbialites around the world61. We hypothesize that our Ulvales-relatives detected in high abundance in Kelly Lake and Pavilion lake may play a significant role in microbialite carbonate deposition in freshwater microbialite communities along with cyanobacteria. Relatedly, chlorophytes were also recovered in high abundance from Lake Alchichica microbialites where they likely play a similarly important role11. Other ANCOM significant taxa in freshwater microbialites include Ostracoda, Maxillopoda, and Cladophorales, Sessilida, and Chytridiomycetes. Chytridiomycetes can be highly abundant in aquatic environments64 and have also been noted in microbialites communities before11, where they may serve an important role in microbialite structural stability65, potentially by accumulating glycogen, the release of which supports microbial heterotrophs66. This group includes many different fungal parasites of algae which may play an important role in the “mycoloop” which allows for the digestion of planktonic matter which would otherwise sink67. Chytridium polysiphoniae and a Rhyzophidiales spp., which were found associated with the freshwater microbialites, are recognized parasites of protists including red algae and amoebas68,69, so we theorize their presence in the microbialite communities is likely supported by a parasitic relationship with other members of the eukaryome.
Of note, the microbialite system in the saline and alkaline Lake Alchichica found a significant presence of both chlorophytes and diatoms in the microbial communities59. These observations support the integral role these protists likely play in global microbialite communities. An important caveat of our results is that neither primer used for the marine and freshwater microbialites can amplify foraminiferal 18S rRNA according to our in silico analysis (Supplementary Table 1). These protists, some of which can produce calcium carbonate tests, have been observed at a high abundance in the microbialites of Shark Bay and Highborne Cay where they are stabilizing sediment grains and affecting stromatolitic mat lamination9. Notably, these protists and related amoebae have been shown to alter microbialite fabric from laminated (stromatolitic) to clotted (thrombolitic) by their reticulopodia and thus the structure and longevity of these structures9,34.
To better understand freshwater microbialite protist communities assemblages were assessed for both Kelly Lake and Pavilion Lake across a depth gradient revealing differing patterns of community composition between the two lakes. Phaeophytes are abundant at all depths in Kelly Lake, however Raphid-pennate diatoms tend to be more abundant at shallower depths (Fig. 4). Certain Phaeophytes, such as Padina, are capable of calcification70, and thus may also contribute and complement the carbonate deposition role of Ulvales and cyanobacteria in freshwater microbialites. In Pavilion Lake, this pattern is reversed with Phaeophytes abundance at greater depths compared to shallower locations (Fig. 4). As for Chlorophytes, Cladophorales show consistent assemblage patterns in both lakes’ depth distribution being more abundant at greater depths (Fig. 4). Ulvales-relatives tended to be more abundant at shallower depths in both lakes (Fig. 4). Groups of Cercozoans, Rhodophytes, and ciliates showed mostly consistent distribution across depths in both lakes. As for the Metazoan families along the depth gradient, Nematodes from the Haliplectoidea family and Chromadorea are more abundant in shallower microbialites than deeper ones in Pavilion Lake (Fig. 4). Metazoan read abundance is lowest at 20 m depth in Kelly Lake (Fig. 4), while Pavilion Lake shows a gradual decrease in metazoan read relative abundance as microbialites are found deeper (Fig. 4). While the two freshwater lakes share many environmental characteristics, Kelly Lake has much less microbialite microbial mat than Pavilion Lake, than even described by Ferris et al.71
Eukaryotes have been prescribed an important role in microbialite formation and function through geologic time1,3,8. The oldest microbialites (stromatolites) date back 3.45 Gy, or perhaps even 3.7 Gy72, but the first eukaryotes evolved more than 1.5 Gy later. Early microbial evolution, including eukaryogenesis, was likely facilitated in stromatolites73 owing to the high abundance of organisms, tight cycling of elements that typically occurs at extremely high rates in a stabilized biofilm environment5,6,7. Although these early eukaryotes lacked body parts that fossilized, biomarkers (e.g., sterane) have suggested 1.65 Gya as their earliest occurrence74. However, Neoproterozoic (800 My) steranes are the oldest non-disputed lipid biomarkers, the date of which coincides with that of ample fossil evidence of protists75,76,77. The oldest reliable evidence for cyst-like microeukaryotes is found in 1.7 Gy old Proterozoic rocks78,79 and photosynthetic multicellular eukaryotic fossils date back ~1.6 Gy80. Phylogenomic Bayesian analyses sets the age of the freshwater amoeboid Paulinella lineage at 2.1 Gya81. Another molecular clock approach proposes the crown group of photosynthetic eukaryotes may even be older, perhaps >2.3 Gya82. These dates coincide with the formation of the earliest thrombolites either 1.9 Gya83 or 2.4 Gya84. The first Archaeplastida evolved around this time as well81, an event that likely occurred in a freshwater environment85, and continued to evolve in such an environment well into the Proterozoic81,82. Reports of diatom-like structures in ~800 My old stromatolites86 suggest a Neoproterozoic occurrence of these protists. As mentioned, diatoms are abundant in saline microbialites today, where they fulfill multiple geomicrobiological roles. More convincing fossil and molecular clock evidence, however, suggest an evolution of diatoms ~200 Mya87. Consequently, these eukaryotes could not have contributed to microbialite formation in the Precambrian. However, their role of EPS production could have been fulfilled by other (agglutinated) protist species78. Similar to the diatoms, foraminifera with tests and related amoeboids appeared in the Cambrian fossil record88 or possibly in the late Precambrian89. Unlike their relatives with shells, agglutinated and naked foraminifera do not fossilize, and thus their exact age is not known, although molecular analyses corroborate a Neoproterozoic appearance88. Foraminifera and thecate amoebae are known to ingest biofilms and microbial mats25,26 and as mentioned, their role in the transition from stromatolite to thrombolite fabric in freshwater and saline environments is well established9,34. The 3.48 Gy old stromatolites of the Dresser Formation formed in fresh to brackish hot spring environment90, and diversified later in the Tumbia Fm, which formed as a lacustrine sediment91. It may be coincidental, but it is noteworthy that both microbialites and eukaryotes both appear to have developed in freshwater environments. Modern microbialites house a wide diversity of eukaryotes the diversity of which is impacted by salinity. Chlorophytes (green algae) were significantly more abundant in the freshwater microbialites, while saltwater microbialites communities were primarily composed of pennate diatoms. Thus, salinity appears to be a main driver of the eukaryotic community composition and structure. However, albeit different in diversity, the eukaryotic community in freshwater, marine and hypersaline environments has a similar ecological role across the salinity spectrum. This is particularly important, as lithification of microbial mats, which marks the onset of microbial formation, is predominantly determined by salinity and light conditions5,37. Different salinities result in different dominant microbial metabolisms as well as in different eukaryotic community compositions, yet the geological structures they form are similar in freshwater, marine and hypersaline environments, indicating a convergent geomicrobiological functionality. Clearly, more biogeochemical gradients should be evaluated for structuring composition and role of these eukaryotes in modern microbialites.
While we tried our best to limit or address potential methodological biases, a comparison of our results against those generated using a different sequencing platform and primers will always be limited by these variables. We have kept most comparisons above the ASV level and have also ran in-silco tests of the primers to ensure we were able to make biologically relevant comparisons between the datasets. Uneven sample numbers between locations were also considered, shown by the inclusion of standard deviations in our analyses when possible to show consistency across samples from the same locations.
Methods
Site description and water chemistry
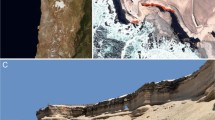
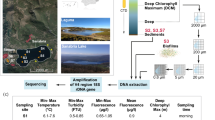
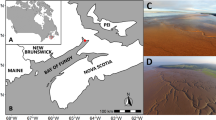
Pavilion Lake (50°51’ N, 121°44’ W) and Kelly Lake (55° 15′ 0″ N, 120° 2′ 0″ W) are mountain lakes in British Columbia, Canada, nestled in a steep-walled valley92,93. Their major limnological characteristics have previously been described by Lim et al., important features include their low phosphate concentrations (~3 µg/L), relatively high CaCO3 concentrations (~182 mg/L), relatively high pH (~8.4), and clear waters with light penetrating to the bottom93. The microbialites consist primarily of calcite thrombolites, whose morphology varies strongly with depth (Fig. 5): Shallow domes resembling “open lettuce-beds” are found at ~10 m depth, intermediate domes resembling “cabbage heads” appear at ~20 m, intermediate-deep domes with hollow conical “asparagus-like” outcroppings at 25 m, and deep domes possessing “artichoke leaflets” appear at and below 25 m92,94. Kelly Lake is shallower with a max depth of ~25 m, and has analogous structures to Pavilion Lake71.
Left) Microbialites of Pavilion and Kelly Lake structure by depth. Right) Map showing the locations of the four microbialite communities analyzed in this study. Fig. 5 was adapted with permission from White III et al.14.
Sampling, DNA extraction, purity, and concentration measurements
Microbialite samples were obtained during the summer of 2010 and 2011. Representative microbialites (~10 kg) were sampled by SCUBA divers along a transect [Pavilion Three Poles (TP) site: 10, 20, 25 m, and Kelly site: 10, 15, 20, 25 m]. A manned DeepWorker submersible sampled the deepest sample in Pavilion Lake (Deep mound site: 45 m; 2010) (Nuytco Research Ltd., North Vancouver, BC, Canada). Divers collected microbialites in autoclaved bags then brought to the surface for dissection for Pavilion and Kelly depths 10–25 m14. The microbiome of the microbialites, sampled using a sterile razor blade, was used to scrape off 3–10 mm (~5 g) across the surface of morphologically similar microbialites collected at each depth. DNA was extracted on-site using a PowerMax® Soil DNA Isolation Kit (Mobio, Carlsbad, CA, USA). DNA concentrations were determined using a Nanodrop-3300 fluorospectrometer (ThermoFisher, Nanodrop Wilmington, DE) with PicoGreen® reagent according to the manufacturer’s instructions (Invitrogen, Carlsbad, CA). Purity of extracted DNA samples was determined by absorbance (260/280 and 260/230 ratios) using a Nanodrop-1000 (ThermoFisher, Nandrop Wilmington, DE). The DNA extraction triplicates were amplified using PCR (with EUK1A [5′-CTGGTTGATCCTGCCAG-3′] and EUK516R [5′-ACCAGACTTGCCCTCC-3′] primers) separately, then pooled for each microbialite depth prior to Illumina library construction95,96,97. OneTaq® Hot Start DNA polymerase was used following standard manufacturers conditions (NEB, Ipswich, MA). PCR thermal cycling conditions were 94 °C—5 min (initial denaturation), 94 °C—1 min, 56 °C—45 s, 72 °C—2 min with 30 cycles, then a final denaturation for 7 min.
Library construction and sequencing
For Illumina library construction, using purified PCR products that were end-paired, A-tailed (Lucigen NxSeq DNA prep kit, Middleton, WI) and ligated to TruSeq adapters (IDT, Coralville, Iowa); small fragments were removed twice using magnetic beads (Beckman Coulter, Danvers, MA). The resulting libraries were pooled, and sequenced using 250 bp paired-end reads on an Illumina MiSeq® by UCLA GenoSeq Core (Los Angeles, CA).
Eukaryote sequence analysis for Kelly and Pavilion lakes
An average of 153,467 reads were recovered per sample, with a median read count of 131,618 reads. Reads were quality checked using FastQC and adapters + primers were trimmed off using Cutadapt then processed using the DADA2 pipeline98,99. Only forward reads were used for the analysis, as these were shown to have comparable taxonomic coverage to the primers used in the marine microbialite samples (Supplementary Table 1). Sequences were filtered by length and quality score, then Amplicon Sequence Variants (ASVs) were inferred using DADA2’s sample inference algorithm. ASV tables for each dataset were then merged into a consensus table. Chimeras were removed before assigning taxonomy using the PR2 18S rRNA database v4.14.0100. Due to a high number of low-resolution taxonomically unresolved ASVs, a BLASTN search was performed using the PR2 database with bacteria and archaea sequences added101. Any sequences matching to bacteria or archaea were then removed from the analysis. Any ASV with an e value (<3 × 10−170) and sequence similarity of 97%, 95–97%, and 90–95% were reassigned to the species, genus, and family level, respectively. The final ASV table was then manually inspected for contaminants and possibly misassigned taxa. The final filtered dataset contained 2488 ASVs across the sampling sites. Alpha diversity plots were generated using the microbiome R package (https://microbiome.github.io/). Relative abundance plots were created showing only the taxa with a median relative abundance above 0.01%. Beta-diversity between the communities were assessed using Aitchison distances and ANOSIM of the center-log ratio transformed counts as these methods account for the compositional nature of microbiome data102. Core analysis was performed using AmpVis v2.8.7 with a minimum abundance threshold set at 1% and minimum prevalence set at 75%103. A weighted UniFrac was constructed for further community analysis across locations using a phylogenetic tree of ASVs constructed using FastTree104. Analysis of Compositions of Microbiomes with Bias Correction (ANCOM-BC) was used to find significantly enriched taxa between the freshwater and saltwater microbialites105.
Comparison to Shark Bay and Highborne Cay
18S rRNA gene pyrosequencing from 9 marine microbialites in Highborne Cay (HC), Bahamas and 6 samples from Shark Bay (SB), Australia27 were downloaded from the Sequence Read Archive (SRA; accession numbers: SRA061992 [Highborne Cay, Bahamas] and SRA061825 [Hamelin pool, Shark Bay, Australia]). We completed the same quality filtering and chimera detection as we did with the Pavilion and Kelly lake 18S rRNA gene data. These ASVs were then merged with our other dataset in DADA2 before taxonomic assignment and further processing as described above for the Kelly Lake and Pavilion Lake samples. Primer bias between these two datasets and ours was assessed by running our primers in silico against the PR2 Database (Supplementary Table 1). We kept all community compositional analyses between these two datasets and the freshwater waters ones above the ASV-level to account for differences in the two sequencing platforms (Pyrosequencing vs Illumina). Even though these samples were sequenced using 454 pyrosequencing as compared to ours which were done using an Illumina platform, a previous analysis has shown little to no apparent effect of sequencing method on beta-diversity and only small differences in alpha diversity for high diversity samples106.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All sequence data used generated during and/or analysed in this study is freely available and available for public access Open Science Framework (OSF, https://osf.io/g28ym/, Modern Microbialites Eukaryome).
Code availability
All code, scripts, data, and documentation used for the analysis of this study can be found on github (https://github.com/delCampoLab/microbialite_euks_2023).
References
Knoll, A. H. The early evolution of eukaryotes: a geological perspective. Science 256, 622–627 (1992).
Awramik, S. M. & Riding, R. Role of algal eukaryotes in subtidal columnar stromatolite formation. Proc. Natl. Acad. Sci. 85, 1327–1329 (1988).
Bosak, T., Knoll, A. H. & Petroff, A. P. The meaning of stromatolites. Annu. Rev. Earth Planet. Sci. 41, 21–44 (2013).
Fossil and Recent Biofilms: a Natural History of Life on Earth. https://doi.org/10.1007/978-94-017-0193-8 (Springer Netherlands, Dordrecht, 2003)
Dupraz, C. & Visscher, P. T. Microbial lithification in marine stromatolites and hypersaline mats. Trends Microbiol. 13, 429–438 (2005).
Dupraz, C. et al. Processes of carbonate precipitation in modern microbial mats. Earth.Sci. Rev. 96, 141–162 (2009).
Visscher, P. T. et al. Microbial mats. no. 163: Part B, Volume 1, Chapter 3: Treatise Online (2022).
Grotzinger, J. P. & Knoll, A. H. Stromatolites in Precambrian carbonates: evolutionary mileposts or environmental dipsticks? Annu. Rev. Earth Planet Sci. 27, 313–358 (1999).
Bernhard, J. M. et al. Insights into foraminiferal influences on microfabrics of microbialites at Highborne Cay, Bahamas. Proc. Natl. Acad. Sci. 110, 9830–9834 (2013).
Power, I. M., Wilson, S., Dipple, G. M. & Southam, G. Modern carbonate microbialites from an asbestos open pit pond, Yukon, Canada. Geobiology 9, 180–195 (2011).
Couradeau, E. et al. Prokaryotic and eukaryotic community structure in field and cultured microbialites from the alkaline lake Alchichica (Mexico). PLoS ONE 6, e28767 (2011).
Khodadad, C. L. M. & Foster, J. S. metagenomic and metabolic profiling of nonlithifying and lithifying stromatolitic mats of Highborne Cay, The Bahamas. PLoS ONE 7, e38229 (2012).
White, R. A. III, Power, I. M., Dipple, G. M., Southam, G. & Suttle, C. A. Metagenomic analysis reveals that modern microbialites and polar microbial mats have similar taxonomic and functional potential. Front. Microbiol. 6, 966 (2015).
White, R. A. III et al. Metagenomic analysis suggests modern freshwater microbialites harbor a distinct core microbial community. Front. Microbiol. 6, 1531 (2016).
Allwood, A. C., Walter, M. R., Kamber, B. S., Marshall, C. P. & Burch, I. W. Stromatolite reef from the Early Archaean era of Australia. Nature 441, 714–718 (2006).
Schopf, J. W. Fossil evidence of Archaean life. Philos. Trans. R. Soc. 361, 869–885 (2006).
Margulis, L. Symbiosis and evolution. Sci. Am. 225, 48–57 (1971).
Walter, M. R. & Heys, G. R. Links between the rise of the metazoa and the decline of stromatolites. Precambrian Res. 29, 149–174 (1985).
Myshrall, K. L., Dupraz, C. & Visscher, P. T. Patterns in microbialites throughout geologic time: is the present really the key to the past? in Experimental Approaches to Understanding Fossil Organisms: Lessons from the Living (eds. Hembree, D. I., Platt, B. F. & Smith, J. J.) 111–142. https://doi.org/10.1007/978-94-017-8721-5_6 (Springer Netherlands, Dordrecht, 2014).
Riding, R. Microbialites, Stromatolites, and Thrombolites. in Encyclopedia of Geobiology (eds. Reitner, J. & Thiel, V.) 635–654. https://doi.org/10.1007/978-1-4020-9212-1_196 (Springer Netherlands, Dordrecht, 2011).
White, R. A. III The Global Distribution of Modern Microbialites: Not So Uncommon After All. in Astrobiology and Cuatro Ciénegas Basin as an Analog of Early Earth (eds. Souza, V., Segura, A. & Foster, J. S.) 107–134. https://doi.org/10.1007/978-3-030-46087-7_5. (Springer International Publishing, Cham, 2020).
Allen, M. A., Goh, F., Burns, B. P. & Neilan, B. A. Bacterial, archaeal and eukaryotic diversity of smooth and pustular microbial mat communities in the hypersaline lagoon of Shark Bay. Geobiology 7, 82–96 (2009).
White, R. A. III, Visscher, P. T. & Burns, B. P. Between a rock and a soft place: the role of viruses in lithification of modern microbial mats. Trends Microbiol. 29, 204–213 (2021).
McNamara, K. J. & Awramik, S. M. Stromatolites: a key to understanding the early evolution of life. Sci. Prog. (1933) 345–364 (1992).
Risse-Buhl, U. et al. Tracking the autochthonous carbon transfer in stream biofilm food webs. FEMS Microbiol. Ecol. 79, 118–131 (2012).
Bernhard, J., Bowser, S., Bernhard, J. M. & Bowser, S. S. Bacterial biofilms as a trophic resource for certain benthic foraminifera. Mar. Ecol. Progr. Ser. 83, 263–272 (1992).
Edgcomb, V. P. et al. Active eukaryotes in microbialites from Highborne Cay, Bahamas, and Hamelin Pool (Shark Bay), Australia. ISME J. 8, 418–429 (2014).
Iniesto, M. et al. Planktonic microbial communities from microbialite-bearing lakes sampled along a salinity-alkalinity gradient. Limnol. Oceanogr. 67, 2718–2733 (2022).
Decho, A. W. & Gutierrez, T. Microbial extracellular polymeric substances (EPSs) in ocean systems. Front. Microbiol. 8, 922 (2017).
Aguilera, A., Souza-Egipsy, V., San Martín-Úriz, P. & Amils, R. Extraction of extracellular polymeric substances from extreme acidic microbial biofilms. Appl. Microbiol. Biotechnol. 78, 1079–1088 (2008).
Gomez, F. J., Mlewski, C., Boidi, F. J., Farías, M. E. & Gérard, E. Calcium carbonate precipitation in diatom-rich microbial mats: the Laguna negra hypersaline lake, Catamarca, Argentina. J. Sediment. Res. 88, 727–742 (2018).
Bowlin, E. M. et al. Environmental controls on microbial community cycling in modern marine stromatolites. Sediment. Geol. 263–264, 45–55 (2012).
Stolz, J. F. et al. The microbial communities of the modern marine stromatolites at Highborne Cay, Bahamas. Atoll Research Bulletin 1–29 (2009).
Bernhard, J. M. et al. Transition from stromatolite to thrombolite fabric: potential role for reticulopodial protists in lake microbialites of a Proterozoic ecosystem analog. Front. Microbiol. 14, 1210781 (2023).
Reid, R. P. et al. The role of microbes in accretion, lamination and early lithification of modern marine stromatolites. Nature 406, 989–992 (2000).
Casaburi, G., Duscher, A. A., Reid, R. P. & Foster, J. S. Characterization of the stromatolite microbiome from Little Darby Island, The Bahamas using predictive and whole shotgun metagenomic analysis. Environ. Microbiol. 18, 1452–1469 (2016).
Visscher, P. T. et al. Biogeochemistry of carbon cycling in hypersaline mats: linking the present to the past through biosignatures. in Microbial Mats: Modern and Ancient Microorganisms in Stratified Systems (eds. Seckbach, J. & Oren, A.) 443–468. https://doi.org/10.1007/978-90-481-3799-2_23 (Springer Netherlands, Dordrecht, 2010).
Dupraz, C. & Strasser, A. Nutritional modes in coral—microbialite reefs (Jurassic, Oxfordian, Switzerland): evolution of trophic structure as a response to environmental change. PALAIOS 17, 449–471 (2002).
Jeffries, T. C. et al. Increases in the abundance of microbial genes encoding halotolerance and photosynthesis along a sediment salinity gradient. Biogeosciences 9, 815–825 (2012).
Wong, H. L., Ahmed-Cox, A. & Burns, B. P. Molecular ecology of hypersaline microbial mats: current insights and new directions. Microorganisms 4, 6 (2016).
Abed, R. M. M., Kohls, K. & De Beer, D. Effect of salinity changes on the bacterial diversity, photosynthesis and oxygen consumption of cyanobacterial mats from an intertidal flat of the Arabian Gulf. Environ. Microbiol. 9, 1384–1392 (2007).
Jacquemot, L. et al. Protist communities along freshwater–marine transition zones in Hudson Bay (Canada). Elementa: Sci. Anthr. 9, 00111 (2021).
Baumgartner, L. K. et al. Microbial species richness and metabolic activities in hypersaline microbial mats: insight into biosignature formation through lithification. Astrobiology 9, 861–874 (2009).
Raes, E. J. et al. Oceanographic boundaries constrain microbial diversity gradients in the South Pacific Ocean. Proc. Natl Acad. Sci. 115, E8266–E8275 (2018).
Dupont, C. L. et al. Functional tradeoffs underpin salinity-driven divergence in microbial community composition. PLOS ONE 9, e89549 (2014).
Lindsay, M. R., Johnston, R. E., Baxter, B. K. & Boyd, E. S. Effects of salinity on microbialite‐associated production in Great Salt Lake, Utah. Bull. Ecol. Soc. Am. 100, e01513 (2019).
Pace, A. et al. Microbial and diagenetic steps leading to the mineralisation of Great Salt Lake microbialites. Sci. Rep. 6, 31495 (2016).
Muthukrishnan, T. et al. Monthly succession of biofouling communities and corresponding inter-taxa associations in the North- and South-West of the Arabian Gulf. Front. Mar. Sci. 8, 787879 (2022).
Tan, M. H. et al. First genome of Labyrinthula sp., an opportunistic seagrass pathogen, reveals novel insight into marine protist phylogeny, ecology and CAZyme cell-wall degradation. Micro. Ecol. 82, 498–511 (2021).
Bickel, S., Tang, K. & Grossart, H.-P. Ciliate epibionts associated with Crustacean zooplankton in German lakes: distribution, motility, and bacterivory. Front. Microbiol. 3, 243 (2012).
Des Marais, D. J. Microbial mats and the early evolution of life. Trends Ecol. Evol. 5, 140–144 (1990).
Underwood, G. J. C. & Paterson, D. M. The importance of extracellular carbohydrate productionby marine epipelic diatoms. Advances in Botanical Research Vol. 40 183–240 (Academic Press, 2003).
Winsborough, B. M. Diatoms and benthic microbial carbonates. in Microbial Sediments (eds. Riding, R. E. & Awramik, S. M.) 76–83. https://doi.org/10.1007/978-3-662-04036-2_10 (Springer, Berlin, Heidelberg, 2000)
Farías, M. E. et al. The discovery of stromatolites developing at 3570 m above sea level in a high-altitude volcanic Lake Socompa, Argentinean Andes. PLoS ONE 8, e53497 (2013).
Brendan, B. P., Walter, M. R. & Neilan, B. A. Microbial communities of stromatolites. in From Fossils to Astrobiology: Records of Life on Earth and Search for Extraterrestrial Biosignatures (eds. Seckbach, J. & Walsh, M.) 143–158. https://doi.org/10.1007/978-1-4020-8837-7_7 (Springer Netherlands, Dordrecht, 2008).
Jahnert, R. J. & Collins, L. B. Characteristics, distribution and morphogenesis of subtidal microbial systems in Shark Bay, Australia. Mar. Geol. 303–306, 115–136 (2012).
Suosaari, E. P. et al. New multi-scale perspectives on the stromatolites of Shark Bay, Western Australia. Sci. Rep. 6, 20557 (2016).
Perkins, R. G., Mouget, J.-L., Kromkamp, J. C., Stolz, J. & Pamela Reid, R. Modern stromatolite phototrophic communities: a comparative study of procaryote and eucaryote phototrophs using variable chlorophyll fluorescence. FEMS Microbiol. Ecol. 82, 584–596 (2012).
Iniesto, M. et al. Core microbial communities of lacustrine microbialites sampled along an alkalinity gradient. Environ. Microbiol 23, 51–68 (2021).
Sherr, E. B. & Sherr, B. F. Significance of predation by protists in aquatic microbial food webs. Antonie Leeuwenhoek 81, 293–308 (2002).
Theisen, C. H. et al. Carbonate fabrics in the modern microbialites of Pavilion Lake: two suites of microfabrics that reflect variation in microbial community morphology, growth habit, and lithification. Geobiology 13, 357–372 (2015).
Dupraz, C. et al. The role of EPS properties and turnover in carbonate mineral nucleation. In Proc. Joint Meeting of the Geological Society of America (Soil Science Society of America, American Society of Agronomy, Crop Science Society of America, Gulf Coast Association of Geological Societies with the Gulf Coast Section of SEPM, 2008).
Liu, B. et al. Taxonomic transfer of Gongrosira fluminensis Fritsch (Chaetophorales, Chlorophyceae) to Lithotrichon Darienko et Pröschold (Ulvales, Ulvophyceae) based on morphological observation and phylogenetic analyses. Fottea 19, 25–32 (2019).
Comeau, A. M., Vincent, W. F., Bernier, L. & Lovejoy, C. Novel chytrid lineages dominate fungal sequences in diverse marine and freshwater habitats. Sci. Rep. 6, 30120 (2016).
Golubic, S., Radtke, G. & Le Campion-Alsumard, T. Endolithic fungi in marine ecosystems. Trends Microbiol. 13, 229–235 (2005).
Visscher, P. T. & Stolz, J. F. Microbial mats as bioreactors: populations, processes, and products. in Geobiology: Objectives, Concepts, Perspectives (ed. Noffke, N.) 87–100. https://doi.org/10.1016/B978-0-444-52019-7.50009-7. (Elsevier, Amsterdam, 2005).
Carreira, C. et al. Fungi and viruses as important players in microbial mats. FEMS Microbiol. Ecol. 96, 1–13 (2020).
Sparrow, F. K. Aquatic Phycomycetes. https://doi.org/10.5962/bhl.title.5685 (University of Michigan Press, 1960).
Karling, J. S. Brazilian Chytrids. IX. Species of rhizophydium. Am. J. Bot. 33, 328–334 (1946).
Yang, F., Wei, Z. & Long, L. Macroalgal calcification and the effects of ocean acidification and global warming. Mar. Freshw. Res. 72, 1697–1710 (2021).
Ferris, F. G., Thompson, J. B. & Beveridge, T. J. Modern freshwater microbialites from Kelly Lake, British Columbia, Canada. PALAIOS 12, 213–219 (1997).
Nutman, A. P., Bennett, V. C., Friend, C. R. L., Van Kranendonk, M. J. & Chivas, A. R. Rapid emergence of life shown by discovery of 3,700-million-year-old microbial structures. Nature 537, 535–538 (2016).
Gudhka, R. K., Neilan, B. A. & Burns, B. P. Adaptation, ecology, and evolution of the halophilic stromatolite archaeon Halococcus hamelinensis inferred through genome analyses. Archaea 2015, 241608 (2015).
Summons, R. E., Welander, P. V. & Gold, D. A. Lipid biomarkers: molecular tools for illuminating the history of microbial life. Nat. Rev. Microbiol 20, 174–185 (2022).
Butterfield, N. J. Bangiomorpha pubescens n. gen., n. sp.: implications for the evolution of sex, multicellularity, and the Mesoproterozoic/Neoproterozoic radiation of eukaryotes. Paleobiology 26, 386–404 (2000).
Porter, S. M. & Knoll, A. H. Testate amoebae in the Neoproterozoic Era: evidence from vase-shaped microfossils in the Chuar Group, Grand Canyon. Paleobiology 26, 360–385 (2000).
Cohen, P. A. & Macdonald, F. A. The proterozoic record of eukaryotes. Paleobiology 41, 610–632 (2015).
Knoll, A. H. Paleobiological perspectives on early eukaryotic evolution. Cold Spring Harb. Perspect. Biol. 6, a016121 (2014).
Knoll, A. H., Javaux, E. J., Hewitt, D. & Cohen, P. Eukaryotic organisms in Proterozoic oceans. Philos. Trans. R. Soc. Lond. B Biol. Sci. 361, 1023–1038 (2006).
Bengtson, S., Sallstedt, T., Belivanova, V. & Whitehouse, M. Three-dimensional preservation of cellular and subcellular structures suggests 1.6 billion-year-old crown-group red algae. PLOS Biol. 15, e2000735 (2017).
Sánchez-Baracaldo, P., Raven, J. A., Pisani, D. & Knoll, A. H. Early photosynthetic eukaryotes inhabited low-salinity habitats. Proc. Natl Acad. Sci. 114, E7737–E7745 (2017).
Blank, C. E. Origin and early evolution of photosynthetic eukaryotes in freshwater environments: reinterpreting proterozoic paleobiology and biogeochemical processes in light of trait evolution. J. Phycol. 49, 1040–1055 (2013).
Kah, L. C. & Grotzinger, J. P. Early proterozoic (1.9 Ga) thrombolites of the Rocknest Formation, Northwest Territories, Canada. PALAIOS 7, 305–315 (1992).
Nomchong, B. J. & Van Kranendonk, M. J. Diverse thrombolites from the c. 2.4 Ga Turee Creek Group, Western Australia. Precambrian Res. 338, 105593 (2020).
Lewis, L. A. Hold the salt: freshwater origin of primary plastids. Proc. Natl Acad. Sci. 114, 9759–9760 (2017).
Harwood, C. L. & Sumner, D. Y. Microbialites of the Neoproterozoic Beck Spring Dolomite, Southern California. Sedimentology 58, 1648–1673 (2011).
Bryłka, K., Alverson, A. J., Pickering, R. A., Richoz, S. & Conley, D. J. Uncertainties surrounding the oldest fossil record of diatoms. Sci. Rep. 13, 8047 (2023).
Pawlowski, J. et al. The evolution of early Foraminifera. Proc. Natl Acad. Sci. USA 100, 11494–11498 (2003).
Bosak, T. et al. Possible early foraminiferans in post-Sturtian (716 − 635 Ma) cap carbonates. Geology 40, 67–70 (2012).
Djokic, T., Van Kranendonk, M. J., Campbell, K. A., Walter, M. R. & Ward, C. R. Earliest signs of life on land preserved in ca. 3.5 Ga hot spring deposits. Nat. Commun. 8, 15263 (2017).
Coffey, J. M., Flannery, D. T., Walter, M. R. & George, S. C. Sedimentology, stratigraphy and geochemistry of a stromatolite biofacies in the 2.72 Ga Tumbiana Formation, Fortescue Group, Western Australia. Precambrian Res. 236, 282–296 (2013).
Laval, B. et al. Modern freshwater microbialite analogues for ancient dendritic reef structures. Nature 407, 626–629 (2000).
Lim, D. S. S. L. Limnology of Pavilion Lake, B. C., Canada characterization of a microbialite forming environment. Fundam. Appl. Limnol. 329–351. https://doi.org/10.1127/1863-9135/2009/0173-0329 (2009).
Omelon, C. R. et al. Microstructure variability in freshwater microbialites, Pavilion Lake, Canada. Palaeogeogr., Palaeoclimatol., Palaeoecol. 392, 62–70 (2013).
Díez, B., Pedrós-Alió, C., Marsh, T. L. & Massana, R. Application of denaturing gradient gel electrophoresis (DGGE) to study the diversity of marine picoeukaryotic assemblages and comparison of DGGE with other molecular techniques. Appl. Environ. Microbiol 67, 2942–2951 (2001).
Eland, L. E., Davenport, R. & Mota, C. R. Evaluation of DNA extraction methods for freshwater eukaryotic microalgae. Water Res 46, 5355–5364 (2012).
Mäki, A., Salmi, P., Mikkonen, A., Kremp, A. & Tiirola, M. Sample preservation, DNA or RNA extraction and data analysis for high-throughput phytoplankton community sequencing. Front. Microbiol. 8, 1848 (2017).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10 (2011).
Callahan, B. J., Mcmurdie, P. J., Rosen, M. J., Han, A. W. & A, A. J. DADA2: High resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Guillou, L. et al. The Protist Ribosomal Reference database (PR2): a catalog of unicellular eukaryote small sub-unit rRNA sequences with curated taxonomy. Nucleic Acids Res. 41, D597–D604 (2013).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 421 (2009).
Gloor, G. B., Macklaim, J. M., Pawlowsky-Glahn, V. & Egozcue, J. J. Microbiome datasets are compositional: and this is not optional. Front. Microbiol. 8, 2224 (2017).
Andersen, K., Kirkegaard, R., Karst, S. & Albertsen, M. ampvis2: an R package to analyse and visualise 16S rRNA amplicon data. bioRxiv 299537 https://doi.org/10.1101/299537 (2018).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2 – approximately maximum-likelihood trees for large alignments. PLOS ONE 5, e9490 (2010).
Lin, H. & Peddada, S. D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 3514 (2020).
Nelson, M. C., Morrison, H. G., Benjamino, J., Grim, S. L. & Graf, J. Analysis, optimization and verification of illumina-generated 16S rRNA gene amplicon surveys. PLoS ONE 9, e94249 (2014).
Acknowledgements
We want to acknowledge the Pavilion Lake Research Project (PLRP) research team, including Donnie Reid, Greg Slater, Curtis Suttle, Amy Chan, Kynan Suttle, Danielle Winget, Kynan Suttle, Jan Finke, Marli Vlok, Tyler Nelson, PLRP science divers, and the DeepWorker pilots, for Pavilion and Kelly Lake field work and sampling. Further thanks to Ts’Kw’aylaxw First Nation, British Columbia Parks, Linda and Mickey Macri for their support of the PLRP, MARSLIFE, and this project. R.A. White III was supported by the UNC Charlotte Department Bioinformatics and Genomics start-up package from the North Carolina Research Campus in Kannapolis, NC, and by the NASA Exobiology project NNH22ZDA001N-EXO. National Science Foundation NSF supports Pieter Visscher grant OCE 1561173 (USA), NASA Exobiology project NNH22ZDA001N-EXO, and ISITE project UB18016-BGS-IS (France). Sampling Financial support was provided by the MARSLIFE Project (9F052-10-0176) funded by the Canadian Space Agency, the NASA MMAMA program. A.M.B. and J.d.C. were supported by project PID2020-118836GA-I00 financed by MCIN/AEI/10.13039/501100011033, by project 2021 SCR 00420 financed by Departament de Recerca i Universitats de la Generalitat de Catalunya, and startup funds from the University of Miami, Rosenstiel School of Marine, Atmospheric and Earth Sciences.
Author information
Authors and Affiliations
Contributions
R.A.W. III collected samples, extracted DNA, designed primers, performed PCRs, made libraries, and other molecular work. A.M.B. and R.A.W. III preformed computational analysis and provided code/plots. A.M.B., J.A.C., P.T.V., R.A.W. III analyzed resulting data for publication. J.A.C., A.M.B., P.T.V. wrote, edited, and reviewed manuscript for publication. All authors approve this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflicts of interest. R.A.W. III is the CEO of RAW Molecular Systems (RAW), LLC, but no financial, IP, or others from RAW LLC were used or contributed to the study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bonacolta, A.M., Visscher, P.T., del Campo, J. et al. The eukaryome of modern microbialites reveals distinct colonization across aquatic ecosystems. npj Biofilms Microbiomes 10, 78 (2024). https://doi.org/10.1038/s41522-024-00547-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41522-024-00547-z
This article is cited by
-
Modern microbialites harbor an undescribed diversity of chromerid algae
Environmental Microbiome (2026)
-
Seasonal biogeochemical variations in a modern microbialite reef under early Earth-like conditions
Communications Earth & Environment (2025)
-
Tropical reefs in the aftermath of climate change
Discover Conservation (2025)