Abstract
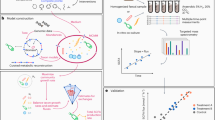
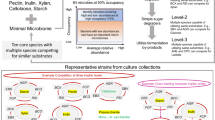
The impact of diet on gut microbial metabolism is essential for advancing microbiome-based health interventions. This study introduces a novel systems biology pipeline that integrates genome-scale metabolic models (GSMMs) with Thai dietary intake data to simulate gut microbiome metabolism and assess prebiotic responses. Utilizing metagenomic data from healthy Thai adults and an average Thai diet derived from national surveys, community-scale metabolic models (CSMMs) were developed and simulated under both typical dietary and prebiotic-supplemented condition. Flux variability analysis was employed to assess metabolic capacities, short-chain fatty acids (SCFAs) production in relation to microbial taxonomy. The results promisingly revealed inter-individual variability in SCFA profiles, with Bacteroides and Phocaeicola notably linked to isobutyrate production and Bifidobacterium emerged as a key responder to prebiotic supplementation. This integrative framework offers biological insights into diet-gut microbiome interactions and provides a foundation for the development of precision nutrition strategies tailored to the Thai population.
Similar content being viewed by others
Data availability
The sequencing data used in the project are openly available from SRA. Raw data supporting the findings (e.g., nutrient-derived metabolites in the average Thai diet and CSMM statistics for Thai adults) are provided as Supplementary Material.
Code availability
Codes and models developed and used in this study are available at https://github.com/nachonase/ThaiCSMMs.
References
Gentile, C. L. & Weir, T. L. The gut microbiota at the intersection of diet and human health. Science 362, 776–780 (2018).
Hindle, V. K., Veasley, N. M. & Holscher, H. D. Microbiota-focused dietary approaches to support health: a systematic review. J. Nutr. 155, 381–401 (2025).
Hitch, T. C. A. et al. Microbiome-based interventions to modulate gut ecology and the immune system. Mucosal Immunol. 15, 1095–1113 (2022).
Chung, Y. W. et al. Functional dynamics of bacterial species in the mouse gut microbiome revealed by metagenomic and metatranscriptomic analyses. PLoS ONE 15, e0227886 (2020).
Patumcharoenpol, P. et al. Exploring longitudinal gut microbiome towards metabolic functional changes associated in atopic dermatitis in early childhood. Biology 12, 1262 (2023).
Kingkaw, A. et al. Analyzing predominant bacterial species and potential short-chain fatty acid-associated metabolic routes in human gut microbiome using integrative metagenomics. Biology 12, 21 (2022).
Patumcharoenpol, P. et al. MetGEMs Toolbox: metagenome-scale models as integrative toolbox for uncovering metabolic functions and routes of human gut microbiome. PLoS Comput. Biol. 17, e1008487 (2021).
Raethong, N. et al. Analysis of human gut microbiome: taxonomy and metabolic functions in Thai adults. Genes 12, 331 (2021).
Yachida, S. et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat. Med. 25, 968–976 (2019).
Zierer, J. et al. The fecal metabolome as a functional readout of the gut microbiome. Nat. Genet. 50, 790–795 (2018).
Noecker, C. et al. Defining and evaluating microbial contributions to metabolite variation in microbiome-metabolome association studies. mSystems 4, e00579–19 (2019).
Shaffer, M. et al. Microbiome and metabolome data integration provides insight into health and disease. Transl. Res. 189, 51–64 (2017).
Magnúsdóttir, S. et al. Generation of genome-scale metabolic reconstructions for 773 members of the human gut microbiota. Nat. Biotechnol. 35, 81–89 (2017).
Baldini, F. et al. The microbiome modeling toolbox: from microbial interactions to personalized microbial communities. Bioinformatics 35, 2332–2334 (2019).
Heinken, A. & Thiele, I. Microbiome modelling toolbox 2.0: efficient, tractable modelling of microbiome communities. Bioinformatics 38, 2367–2368 (2022).
Shaaban, R. et al. Personalized modeling of gut microbiome metabolism throughout the first year of life. Commun. Med. 4, 281 (2024).
Heinken, A. et al. Systematic assessment of secondary bile acid metabolism in gut microbes reveals distinct metabolic capabilities in inflammatory bowel disease. Microbiome 7, 75 (2019).
Basile, A. et al. Longitudinal flux balance analyses of a patient with episodic colonic inflammation reveals microbiome metabolic dynamics. Gut Microbes 15, 2226921 (2023).
Martinelli, F. et al. Whole-body metabolic modelling reveals microbiome and genomic interactions on reduced urine formate levels in Alzheimer’s disease. Sci. Rep. 14, 6095 (2024).
Vangay, P. et al. US immigration westernizes the human gut microbiome. Cell 175, 962–972.e10 (2018).
Ministry of Social, D. & T. Human Security, National Food Consumption Survey Report. (2015).
Judprasong, K. et al. Thai Food Composition Database, Online version 2. (Institute of Nutrition, 2015).
Noronha, A. et al. The virtual metabolic human database: integrating human and gut microbiome metabolism with nutrition and disease. Nucleic Acids Res. 47, D614–d624 (2019).
Bureau of Nutrition, Department of Health, Ministry of Public Health. Dietary Reference Intake Tables for Thais (Bangkok, Thailand, 2020).
Hertel, J. et al. Integration of constraint-based modeling with fecal metabolomics reveals large deleterious effects of Fusobacterium spp. on community butyrate production. Gut Microbes 13(1), 1–23 (2021).
Sathitkowitchai, W. et al. A randomized trial to evaluate the impact of copra meal hydrolysate on gastrointestinal symptoms and gut microbiome. PeerJ 9, e12158 (2021).
Rios Garza, D. et al. Metabolic models of human gut microbiota: advances and challenges. Cell Syst. 14, 109–121 (2023).
Ivanovitch, K. et al. The intake of energy and selected nutrients by Thai urban sedentary workers: an evaluation of adherence to dietary recommendations. J. Nutr. Metab. 2014, 145182 (2014).
Tjahyo, A. S. et al. Shifting trend of protein consumption in Southeast Asia: toward health, innovation, and sustainability. Curr. Dev. Nutr. 8, 104443 (2024).
Krongdang, S. et al. Edible insects in Thailand: an overview of status, properties, processing, and utilization in the food industry. Foods 12 https://doi.org/10.3390/foods12112162 (2023).
Chailimpamontree, W. et al. Estimated dietary sodium intake in Thailand: a nationwide population survey with 24-h urine collections. J. Clin. Hypertens. 23, 744–754 (2021).
Yongsmith, B. and W. Malaphan Traditional Fermented Foods in Thailand, in Traditional Foods: General and Consumer Aspects, (eds Kristbergsson K. & Oliveira J.) p. 31–59 (Springer, 2016).
Cook, N. R. et al. Long term effects of dietary sodium reduction on cardiovascular disease outcomes: observational follow-up of the trials of hypertension prevention (TOHP). BMJ 334, 885–8 (2007).
Bromage, S. et al. Development and validation of a novel food-based global diet quality score (GDQS). J. Nutr. 151, 75S–92S (2021).
Bromage, S. et al. Performance of the global diet quality score (GDQS) App in predicting nutrient adequacy and metabolic risk factors among Thai adults. J. Nutr. 153, 3576–3594 (2023).
Huttenhower, C. et al. Structure, function and diversity of the healthy human microbiome. Nature 486(7402), 207–214 (2012).
Almeida, A. et al. A new genomic blueprint of the human gut microbiota. Nature 568, 499–504 (2019).
La-Ongkham, O. et al. Age-related changes in the gut microbiota and the core gut microbiome of healthy Thai humans. Biotechnology 10, 276 (2020).
Lopez-Siles, M. et al. Faecalibacterium prausnitzii: from microbiology to diagnostics and prognostics. ISME J. 11, 841–852 (2017).
Schaus, S. R. et al. Ruminococcus torques is a keystone degrader of intestinal mucin glycoprotein, releasing oligosaccharides used by Bacteroides thetaiotaomicron. mBio 15, e0003924 (2024).
Sung, J. et al. Global metabolic interaction network of the human gut microbiota for context-specific community-scale analysis. Nat. Commun. 8, 15393 (2017).
LeBlanc, J. G. et al. Beneficial effects on host energy metabolism of short-chain fatty acids and vitamins produced by commensal and probiotic bacteria. Microb. Cell Factories 16, 79 (2017).
den Besten, G. et al. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 54, 2325–40 (2013).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–50 (2013).
He, J. et al. Short-chain fatty acids and their association with signalling pathways in inflammation, glucose and lipid metabolism. Int. J. Mol. Sci. 21, 6356 (2020).
Aguirre, M. et al. Diet drives quick changes in the metabolic activity and composition of human gut microbiota in a validated in vitro gut model. Res. Microbiol. 167, 114–25 (2016).
Zarling, E. J. & Ruchim, M. A. Protein origin of the volatile fatty acids isobutyrate and isovalerate in human stool. J. Lab Clin. Med. 109, 566–70 (1987).
Smith, E. A. & Macfarlane, G. T. Enumeration of amino acid fermenting bacteria in the human large intestine: effects of pH and starch on peptide metabolism and dissimilation of amino acids. FEMS Microbiol. Ecol. 25, 355–368 (1998).
Procházková, N. et al. Advancing human gut microbiota research by considering gut transit time. Gut 72, 180 (2023).
Rodríguez-Romero, J. J. et al. What we know about protein gut metabolites: Implications and insights for human health and diseases. Food Chem. X 13, 100195 (2022).
Duizer, C. & de Zoete, M. R. The role of microbiota-derived metabolites in colorectal cancer. Int. J. Mol. Sci. 24, 8024 (2023).
Prescott, S. L. et al. Biodiversity, the human microbiome and mental health: moving toward a new clinical ecology for the 21st century? Int. J. Biodivers. 2016, 2718275 (2016).
Gibson, G. R. et al. Expert consensus document: the International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat. Rev. Gastroenterol. Hepatol. 14, 491–502 (2017).
Nguyen, N. K. et al. Gut microbiota modulation with long-chain corn bran arabinoxylan in adults with overweight and obesity is linked to an individualized temporal increase in fecal propionate. Microbiome 8, 118 (2020).
Rodriguez, J. et al. Discovery of the gut microbial signature driving the efficacy of prebiotic intervention in obese patients. Gut 69, 1975–1987 (2020).
Prayoonthien, P. et al. In vitro fermentation of copra meal hydrolysate by human fecal microbiota. Biotechnology 9, 93 (2019).
Kraikaew, J., Morakul, S. & Keawsompong, S. Nutritional improvement of copra meal using mannanase and Saccharomyces cerevisiae. Biotechnology 10, 274 (2020).
Pomyen, Y. et al. Gut dysbiosis in Thai intrahepatic cholangiocarcinoma and hepatocellular carcinoma. Sci. Rep. 13, 11406 (2023).
Aroney, S. et al. CoverM: read alignment statistics for metagenomics. Bioinformatics 41, btaf147 (2025).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Heirendt, L. et al. Creation and analysis of biochemical constraint-based models using the COBRA Toolbox v.3.0. Nat. Protoc. 14, 639–702 (2019).
Acknowledgements
This project is funded by National Research Council of Thailand (NRCT): Contract number N42A660907 and N42A650235. The authors gratefully acknowledge the faculty members and research staffs from the Food and Nutrition Database Unit, Institute of Nutrition, for their insightful feedback and problem-solving suggestions, which significantly contributed to the research implementation. Their support also included access to software tools, food composition databases, and national FCS reports. Lastly, appreciation is extended to the Galaxy server (https://usegalaxy.eu), maintained by the Freiburg Galaxy Team, for offering storage space and high-performance computing resources that enabled the efficient analysis of large-scale microbiome datasets. We also would like to thank Department of Zoology, Faculty of Science, Kasetsart University, SciKU Biodata Server for computing facilities, Postdoctoral Fellowship Program at Kasetsart University, and Kasetsart University Research and Development Institute (KURDI).
Author information
Authors and Affiliations
Contributions
N.R. performed the computational experiments, visualized the data, interpreted the results, and prepared the manuscript. P.P. assisted with data curation and contributed to data interpretation. N.R. and W.V. conceived the study, acquired funding, and critically reviewed the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Raethong, N., Patumcharoenpol, P. & Vongsangnak, W. Modeling diet-gut microbiome interactions and prebiotic responses in Thai adults. npj Biofilms Microbiomes (2026). https://doi.org/10.1038/s41522-026-00921-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41522-026-00921-z