Abstract
For neural network potentials (NNPs) to gain widespread use, researchers must be able to trust model outputs. However, the blackbox nature of neural networks and their inherent stochasticity are often deterrents, especially for foundation models trained over broad swaths of chemical space. Uncertainty information provided at the time of prediction can help reduce aversion to NNPs. In this work, we detail two uncertainty quantification (UQ) methods. Readout ensembling, by finetuning the readout layers of an ensemble of foundation models, provides information about model uncertainty, while quantile regression, by replacing point predictions with distributional predictions, provides information about uncertainty within the underlying training data. We demonstrate our approach with the MACE-MP-0 model, applying UQ to the foundation model and a series of finetuned models. The uncertainties produced by the readout ensemble and quantile methods are demonstrated to be distinct measures by which the quality of the NNP output can be judged.
Similar content being viewed by others
Introduction
Neural network potentials (NNPs) are a class of machine learning interatomic potentials (MLIPs) trained to approximate the energy landscape of atomic systems in order to drive atomistic simulations. Specifically, NNPs model the relationship between the atomic configuration and associated system energy and atomic forces. When well-trained and used in-domain, NNPs combine the accuracy of quantum mechanical methods with the efficiency of classical potentials1,2,3,4. In practice, distinguishing in-domain from out-of-domain structures is challenging. Out-of-domain structures can easily be generated during the course of a simulation begun from an in-domain sample. Errors on these new out-of-domain structures can compound over the course of the simulation, leading to inaccurate probability distributions, incorrect observables, or even unphysical results. This effect is especially pronounced in cases where errors lead to the creation of artificial attractive forces5.
Uncertainty quantification (UQ) is used to identify poorly learned or out-of-domain structures for active learning, with model ensembling being a popular technique. In this approach, a set of NNPs are independently trained using a common dataset but different initializations and/or network architectures. A variety of methods for calculating the uncertainty have been demonstrated6,7,8,9, typically involving the standard deviation of ensemble predictions. Because of the computational expense of training NNPs, ensembles are often limited to 5–10 independent models.
Single-model UQ techniques have been explored to reduce the computational expense of ensembling. Wen et al. developed a NNP architecture that incorporated dropout-based uncertainty10. Zhu et al. used a Gaussian mixture model11, while Thaler et al. applied a Bayesian method12 to estimate the uncertainty of a single NNP. Soleimany et al.13 implemented evidential deep learning for molecular property prediction, which has since been extended to NNPs14,15,16. In a similar fashion, Busk et al.17 and Carrete et al.18 coupled a pretrained model head and a nonlinear scaling function to attach a variance to the energy contribution from each atom, which were then summed to produce the sample uncertainty.
Debate is ongoing as to the technique that best measures uncertainty in neural networks19. In an examination of the quality of ensemble-based uncertainty estimates, Kahle et al. observed that ensembles tended to underestimate uncertainty and suggested that the ideal ensembling technique must be optimized for each dataset and network architecture20. Conversely, Tan et al. claimed that ensembling leads to more generalizable and robust NNPs than single-model uncertainty techniques15. For single-model NNP active learning, Thomas-Mitchell et al. found that uncertainties from Gaussian processes are not reliable, even after post-hoc calibration, and advocated the use of a student-t process21. Meanwhile, Dai et al. performed a broad examination of existing UQ methods for atomistic machine learning approaches and found that in many cases predicted uncertainties do not match well with the observed errors22.
Further confounding the issue, the dataset used to train the NNP can also contain inherent uncertainty. In classical molecular dynamics (MD) simulations, stochastic uncertainty arises from the chaotic nature of MD and the extreme sensitivity of Newtonian dynamics to initial conditions23,24. Density functional theory (DFT), which is used to collect the vast majority of NNP training data, introduces energy fluctuations that are dependent on the exchange-correlation functional25. For higher levels of theory, statistical noise results from convergence criteria, among other subtle computational choices26.
In an effort to improve the generalizablility of NNPs and potentially reduce epistemic uncertainties arising from poorly approximated energy landscapes, NNP researchers have begun to produce foundation models. Foundation models are trained over large, structurally diverse datasets, often at significant computational cost, to capture general relationships present in the data. Such models can then be adapted to specific applications through finetuning with less data and at reduced computational cost.
Developers of the ANI-1 architecture27,28 have recently explored its use as a foundation model for condensed phase reactive chemistry of structures containing H, C, N, and/or O29 and for drug-like molecules30. Foundation models for solid-phase materials have been produced for the CHGNet31, MACE32, and M3GNet33 architectures using the Materials Project Trajectory (MPtrj) Dataset, which contains 1.6M materials and spans 89 elements. The Open Catalyst Project has developed foundation models for solid-phase catalysis using an open database of > 10M structures34,35,36.
Despite the success of NNP foundation models to produce accurate energy predictions over a broad range of structures, extension to novel systems remains a challenge. Numerous assessments of current NNP foundation models have noted the need for finetuning when extrapolating to new tasks or out-of-domain atomic environments37,38,39,40,41. The difficulty of distinguishing out-of-domain from in-domain structures necessitates the need for quantifying uncertainty during inference. More generally, if NNPs are to gain widespread practical use, UQ provides a way to establish trust in the output of NNP-driven simulations.
Herein, we demonstrate two UQ methods for NNP foundation models: readout ensembling and quantile regression. Each method has unique advantages. Ensembling is useful for identifying epistemic uncertainties, while quantile regression captures aleatoric uncertainties42. Both approaches are applied to MACE-MP-032 to generate uncertainties for the foundation model. We then demonstrate transfer to novel datasets: a high entropy alloy dataset with high chemical complexity43 and a highly specific zeolite dataset with varying numbers of water molecules inside the pores. We find that quantile regression is useful for capturing variations in chemical complexity, while ensembling is useful for capturing out-of-domain structures. We find that the ensemble is overconfident in its predictions and, though ensemble uncertainty tends to increase with error, the magnitude of uncertainty is lower than the error by orders of magnitude. Conversely, the quantile uncertainty more accurately reflects the model’s prediction ability and tends to increase with system size.
Results
Uncertainty quantification for neural network potentials
Model ensembling helps to reduce model bias and mitigate overfitting, resulting in higher accuracy predictions. For foundation models, training is often highly compute intensive. For instance, the MACE-MP-0 foundation models used 40–80 NVIDIA A100 GPUs when training a single model32. This high computational cost hinders the training of a full ensemble of models. To reduce computational costs and maintain the learned representation of the foundation model, we apply readout ensembling Fig. 1. In readout ensembling, only the weights of the final readout layers are updated during training. Because each model in the ensemble is initialized with the same weights, stochasticity is introduced by finetuning the readout layers on different subsets of the full training set. The lower number of weights to be updated and smaller dataset lead to greatly decreased computational costs, such that each model in the readout ensemble could be trained on a single NVIDIA P100 GPU.
Each model in the ensemble is trained using the Huber loss function, which is a piecewise function that switches between the mean squared error (MSE) and mean absolute error (MAE) depending on a set threshold. The symmetric nature of the Huber loss function – and of MSE and MAE individually – ensures that predictions higher and lower than the target value are penalized equally. Variability in predictions given by an ensemble of models can be standard deviation, while confidence intervals (CIs) can be computed from the Student’s t-distribution.
Quantile regression makes use of an asymmetric function that penalizes above and below the target value differently. In this way, ground truth quantiles are not required for training. For instance, to predict the 95th percentile, a penalty of 0.95 times the prediction error would be given when the prediction is higher than the target value and a penalty of 0.05 would be given when the prediction is lower. The asymmetric penalization causes the prediction, after further training epochs, to move towards the desired quantile. To form uncertainty bounds using quantile regression, we modify the network architecture to have two readout layers with opposite penalization (Fig. 1): one readout is penalized by 0.95 and 0.05 for higher and lower predictions, while the other readout is penalized 0.05 and 0.95, respectively. This architecture produces two predictions targeted at the 95th and 5th percentiles, respectively. The difference between these predictions gives the 90% CI.
Though both ensembling and quantile regression can produce CIs, the methods apply different statistical assumptions. Ensembling approximates the model posterior, while quantile regression approximates the conditional distribution44,45. With infinite data and a perfect model, the model posterior uncertainty would vanish, but the conditional distribution would still exist. These different assumptions lead to different types of uncertainty. Quantile regression captures aleatoric uncertainty in the training data distribution, while ensembling captures both epistemic uncertainty in model parameters and aleatoric uncertainty. Because epistemic uncertainty is captured, CIs derived from the ensemble are wider in regions of parameter uncertainty or sparse data.
Uncertainty in the MACE-MP-0 foundation model
We first examine UQ for the out-of-the-box MACE-MP-0 foundation model by readout ensembling and quantile regression. Each of the 7 models in the readout ensemble was trained on a unique set of 90,000 structures chosen at random from the MPtrj dataset, using 80,000 for training and 10,000 for validation. The quantile model was trained on a separate set of 90,000 MPtrj structures. All testing was performed on a common set of 10,000 MPtrj structures. We report the errors in energy prediction and associated uncertainties as per-electron values (meV/e−) to remove size extensive effects of DFT-calculated energies, which scale with the number of electrons. Such scaling enables comparison among the wide range of structures contained in the MPtrj dataset. Alternative units for the values reported below are given in Table S1 in the Supplementary Information.
The readout ensemble and quantile model give similar values for the mean absolute error (MAE) in energy prediction on the MPtrj test set of 0.721 and 0.890 meV/e−, respectively. It should be noted that these errors are in line with those reported for MACE-MP-0 after finetuning the ‘small’ model for 50 additional epochs with higher weighting of the energy component of the loss function32. The pretrained MACE-MP-0 without finetuning gives a test set error of 0.739 meV/e−, which is equivalent to the MAE of 13 meV/atom reported by Batatia et al. The MAEs of the readout ensemble and quantile model translate to 13 meV/atom and 16 meV/atom, respectively. Therefore, we can claim that our models are well-trained on the MPtrj dataset.
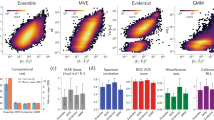
As shown in Table 1, the mean uncertainty of the quantile model (1.391 meV/e−) is over an order of magnitude higher than that of the readout ensemble (0.036 meV/e−). We evaluate the quality of the estimated uncertainties in two ways, shown in Fig. 2. First, we calculate the coverage, defined here as the percent of samples in which the target value falls within the 5th and 95th percentiles derived from the uncertainty. The coverage by the quantile model (87%) greatly exceeds that of the readout ensemble (11%), which results from the larger uncertainties of the quantile model. Next, we examine the correlation between uncertainty and prediction error. In the ideal case, high uncertainty should correspond to high prediction error. Examining the uncertainty distributions of samples within specified MAE ranges, the uncertainty from the quantile model clearly increases with MAE and is closer in magnitude than that from the readout ensemble. It should be noted that as the mean uncertainty increases with MAE, so does the spread of uncertainties. Therefore, a single uncertainty is not directly representative of the prediction error, but in general, larger uncertainties indicate higher errors.
A Regression of uncertainty (U) vs absolute error (AE) of predictions with the readout ensemble and quantile model on the MPtrj test set. The lowess curve shows the 90% CI. B Coverage of the readout ensemble and quantile model. C, D Density maps of configurations contributing to the fitted curve in (A).
The wide breadth of structural space covered by MPtrj and the slight variations in simulation procedures (inconsistent application of Hubbard U correction, varying convergence criteria, etc.) contributes to increased aleatoric uncertainty in the data, which is reflected in the quantile uncertainty. Conversely, the low readout ensemble uncertainty indicates low epistemic uncertainty, which reflects the high quality of MACE-MP-0.
The number of models in an ensemble will affect the resultant uncertainty. More models generally lead to more reliable uncertainty estimates, though with diminishing return. To examine this effect, we recalculated the uncertainties using our readout ensemble models trained on the MPtrj data by leaving 1, 2, or 3 models out of the ensemble. We calculated the uncertainty for each combination of model removal and provide statistics in Table 2. As expected, the uncertainty decreases as the number of models in the ensemble increases.
Transfer learning to a dataset with high chemical complexity
We then explored UQ during transfer learning of the MACE-MP-0 foundation model. Finetuning a foundation model limits the amount of stochasticity when ensembling, specifically in terms of randomized initialization (initial weights are transferred from the foundation model) and dataset splitting (taking unique subsets of a small dataset would lead to very small training sets). This leaves the seed controlling the random number generator, which influences shuffling of the training set between epochs and nondeterministic algorithms used by the PyTorch and cuDNN libraries, as the only source of stochasticity for our finetuned foundation readout ensemble.
The variety of elements present in each sample makes HEA25 a useful dataset for examining effects of chemical complexity on both UQ and model performance. As shown in Table 1, the readout ensemble and quantile model give similar MAEs of 0.971 and 1.013 meV/e−, respectively, while the uncertainty from the readout ensemble (0.132 meV/e−) is much lower than that from the quantile model (1.829 meV/e−). The low uncertainty for the readout ensemble indicates that the model-derived uncertainty is low (i.e., the models have all learned similar sample spaces), while the comparatively higher uncertainty for the quantile model indicates the aleatoric uncertainty is high. Because aleatoric uncertainty reflects the inherent variability in the training data, we can interpret the higher uncertainty in the quantile model to result from the chemical complexity of the HEA25 dataset. Interestingly, the quantile uncertainty roughly correlates with MAE, as shown by fitted loss curve in Fig. 3.
Transfer learning to a dataset of highly ordered configurations
We then examine transfer learning of the MACE-MP-0 model to a dataset with highly ordered configurations: the aluminosilicate zeolite H-ZSM-5 infiltrated with water. Five ab initio molecular dynamics (AIMD) simulations were performed with n = 1, 2, 3, 8, or 16 water molecules in a pore. Simulations with n = 1–3 water molecules were used to finetune MACE-MP-0, and n = 8, 16 were used as holdout sets to examine the effect of larger systems on the uncertainty.
The training dataset is quite small (8543 samples) and has limited chemical complexity (4 types of elements per sample), but each sample has a relatively large number of atoms (295−301). As shown in Table 1, the readout ensemble and quantile model produce similar MAEs on the n = 1–3 test set of 0.035 and 0.031 meV/e−, respectively. The high accuracy is likely a result of 1) the specificity of the H-ZSM-5 dataset and 2) the sufficient representation of ZSM-5 atomic neighborhoods in the MPtrj dataset, as demonstrated by the good zero-shot performance by MACE-MP-032. The finetuned readout ensemble gives comparable uncertainty to the MACE-MP-0 readout ensemble (0.032 vs 0.036 meV/e−, respectively), while the quantile model shows greatly reduced uncertainty (0.056 vs 1.391 meV/e−). The large reduction in uncertainty from the quantile model is likely due to the rigid structure of H-ZSM-5, which limits the configurations that can be sampled during MD simulations.
Table 3 shows the MAEs and uncertainties for each subset, including the holdout sets with n=8, 16 water molecules. The MAE increased almost 6-fold when n = 8 and 10-fold when n = 16 for both the readout ensemble and quantile model. The uncertainty from both models also increases for the n = 8, 16 holdout sets. For the readout ensemble, all training subsets (n = 1–3) have the same mean uncertainty of 0.032 meV/e−, while that of the holdout sets (n = 8, 16) increases slightly with number of water molecules. Meanwhile, the quantile uncertainty increases nearly linearly with number of water molecules across all subsets. These differing uncertainty behaviors further support the notion that ensemble uncertainty reflects configurations that are poorly learned by the model (i.e., out-of-domain), while the quantile uncertainty indicates possible variability in a configuration.
Figure 4 compares the computed versus predicted total energies for each subset and gives uncertainty bands corresponding to the average upper and lower bounds. While the training subsets are centered around the ideal fit, the extrapolation subsets are clearly offset from ideal with the offset increasing with n (see Table 3). Table S2 and Figure S1 show the calibrated MAE and coverage. Note that the uncertainty does not change when correcting for this offset. Correcting for the offset by subtracting the mean difference between the predicted and computed energies for each subset decreases the MAE to 0.041 meV/e− for n = 8 for both the readout ensemble and quantile model and to 0.045 and 0.044 meV/e− for n = 16 for the readout ensemble and quantile model, respectively. As a result, the coverage of the readout ensemble increases to 51 and 53% for n = 8, 16, and that of the quantile model increases to 85 and 91%, respectively.
Comparison of total energies (eV) from DFT and predicted by the readout ensemble and quantile model for H-ZSM-5 infiltrated with n = 1, 2, 3, 8, or 16 water molecules. Markers represent individual predictions, solid lines represent average upper and lower bounds based on the uncertainty, and black dotted lines show the ideal fit. Energies are centered around the mean value from DFT for each subset.
Discussion
In this work, we demonstrated two methods for UQ in NNP foundation models: readout ensembling and quantile regression. The uncertainty derived from quantile regression reflects the chemical complexity in the training set (aleatoric uncertainty), while the uncertainty derived from ensembling predominantly results from model training (epistemic uncertainty). Altering only the readout layers maintains the bulk of the foundation model weights, and therefore, only a small amount of training is necessary to incorporate uncertainty estimation into the foundation model. Notably, both UQ methods can be applied to any NNP foundation model with minimal alteration.
The two methods provide distinct information about the quality of the predicted output. Ensemble uncertainty identifies when a structure is poorly described by the model (i.e., out-of-domain). In general, such uncertainties can be improved by extending model training, expanding the training dataset, or moving to a new model architecture. In contrast, quantile uncertainty is a reflection of the variability in the underlying training data, which is propagated to the model’s predictions. In principle, this type of uncertainty cannot be improved with additional training or model architecture choices. Altering the training dataset will have an effect, but not always as expected. In fact, more complex samples (i.e., high entropy alloys) were shown to increase the quantile uncertainty, even though the number of unique materials in the training set decreased and a common set of simulation parameters was used to generate the HEA25 dataset.
Though in this work we examined the two methods separately, they could be integrated into a unified framework to adapt to both the amount of available data and the inherent variability in different regions of the input data. For example, to obtain a conservative estimate of the uncertainty, the set of CIs could be combined by taking the maximum of the lower bounds and minimum of the upper bounds. In data-dense regions where the ensemble is confident but the data distribution varies, the intervals from quantile regression would dominate, while the ensemble intervals would dominate in data-poor regions where the ensemble gives high uncertainty. This is just one example of how the two methods could be combined, and more research would be needed to determine the most effective way of combining methods. UQ was applied only to energy predictions in this work. Future work will examine applications of UQ to atomic forces, as well as propagation of uncertainty during NNP-driven simulations. The codebase used in this work is available at github.com/pnnl/SNAP.
Methods
MACE readout ensembling
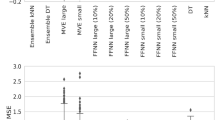
Three different-sized models are provided under the MACE-MP-0 umbrella, labeled ‘small’, ‘medium’, and ‘large’ depending on the model size32. To demonstrate proof of concept, we use the ‘small’ MACE-MP-0 model throughout this work (see Code Block S1 in the Supplementary Information for a full accounting of the model architecture). It should be noted that the ‘small’ model contains approximately 3.85 million learnable parameters. In this instance, ‘small’ is in reference to the other MACE-MP-0 models and should not be considered small in comparison to other NNPs. Each ensemble was made up of 7 models that were trained by freezing the weights of the MACE-MP-0 interaction head and only allowing the final readout module to update. For the foundation ensemble, each model was trained on a unique randomly sampled 80,000 subset of the full MPtrj dataset using a unique seed. When transferring to new datasets, the same training set was used for each model in the ensemble, but unique seeds were set. The ensemble uncertainty U is defined as half the length of the 90% CI of the model predictions determined from the Student’s t-distribution. The 90% CI was chosen to provide a direct comparison to the CI output from the quantile model.
A learning rate of 0.001 was used with MPtrj, which was increased to 0.01 for transfer learning. The learning rate was adaptively lowered after 10 consecutive epochs with no improvement in validation loss. After a minimum of 50 epochs, training was stopped after no improvement in validation loss was obtained over 15 consecutive epochs. All models were trained on a single NVIDIA P100 GPU using the PyTorch Lightning framework46.
Quantile regression
We updated the MACE-MP-0 architecture such that instead of predicting a single target output, our model predicts the 5th and 95th quantiles of the target output. This architecture change is achieved by simply duplicating the final readout module so that each quantile is predicted by a unique readout module. Quantile regression is then applied to simultaneously optimize the readout modules. The quantile model was trained similarly to the readout ensemble models by freezing the MACE-MP-0 interaction head and using the same learning rate and early stopping procedures. The main difference in training is the use of a loss function specific to quantile regression.
Quantile regression typically applies the pinball loss function to train a model to produce a probabilistic prediction of each sample in the form of upper and lower quantiles. See the following works for a thorough discussion of quantile regression47,48. Hatalis et al. showed that a smooth approximation of the pinball loss function, originally described by Songfeng Zheng49, improved the performance of quantile regression on neural networks50. Though their target application was wind power forecasting, we find their smooth pinball loss to be useful for training NNPs. The smooth pinball loss function \({\mathcal{L}}\) is the sum of loss contributions \({{\mathcal{L}}}_{q}\) from the selected quantiles q:
where q is the optimization quantile, E is the prediction target, Eq is the prediction at quantile q, and α is a smoothing parameter. In this work we use q = 0.05, 0.95 as the lower and upper bounds to the energy prediction, which gives \({\mathcal{L}}={{\mathcal{L}}}_{0.05}+{{\mathcal{L}}}_{0.95}\). During inference, the model outputs both the upper and lower bounds. The uncertainty U is calculated as follows: U = (E0.95 − E0.05)/2. Though quantile regression does not make assumptions about the symmetry of the distribution, for simplicity, we take the mean of the quantile predictions as the predicted target value.
Datasets
MPtrj
The MPtrj dataset31 contains nearly 1.6 million configurations for 150,000 unique inorganic crystals derived from DFT static and relaxation trajectories collected over a decade. MPtrj was used to train MACE-MP-0, and a thorough description of the MPtrj dataset can be found in the supporting information of the report introducing MACE-MP-032. In total, 89 elements are represented, though individual configurations are composed of ≤ 8 elements with the majority containing 3 elements. Configurations range in size from 1–444 atoms, with the vast majority (~ 97%) having < 100 atoms.
HEA25
The HEA25 dataset from Lopanitsyna et al.43 includes 25,628 configurations of high entropy alloys (HEAs) consisting of 36 atoms for body-center cubic (bcc) lattices and 48 atoms for face-centered cubic (fcc) lattices with mixtures of up to 25 d-block transition metals. The disordered multi-element configurations present in HEA25 give rise to a large variety of atomic neighborhoods.
H-ZSM-5 ⋅ nH2O
We prepared a dataset composed of the H form of Zeolite Socony Mobil-5 (H-ZSM-5), in which H+ ions occupy the zeolite ion exchange sites. H-ZSM-5 is composed of 4 elements (H, O, Al, and Si) and belongs to the pentasil family of zeolites. Channels run parallel and perpendicular through the structure, allowing infiltration of solvents and small molecules within the pores. We examined the infiltration of 1, 2, 3, 8, or 16 water molecules. Simulation details are provided in the Supplementary Information.
Code availability
The codebase developed for this study is available on GitHub at https://github.com/pnnl/SNAP.
References
Gokcan, H. & Isayev, O. Learning molecular potentials with neural networks. Wiley Interdiscip. Rev.: Computational Mol. Sci. 12, e1564 (2022).
Kocer, E., Ko, T. W. & Behler, J. Neural network potentials: A concise overview of methods. Annu. Rev. Phys. Chem. 73, 163–186 (2022).
Duval, A. et al. A hitchhiker’s guide to geometric gnns for 3d atomic systems. arXiv preprint arXiv:2312.07511 (2023).
Käser, S., Vazquez-Salazar, L. I., Meuwly, M. & Töpfer, K. Neural network potentials for chemistry: concepts, applications and prospects. Digital Discov. 2, 28–58 (2023).
Behler, J. & Csányi, G. Machine learning potentials for extended systems: a perspective. Eur. Phys. J.B 94, 1–11 (2021).
Smith, J. S., Nebgen, B., Lubbers, N., Isayev, O. & Roitberg, A. E. Less is more: Sampling chemical space with active learning. J. Chem. Phys. 148, 241733 (2018).
Jeong, W., Yoo, D., Lee, K., Jung, J. & Han, S. Efficient atomic-resolution uncertainty estimation for neural network potentials using a replica ensemble. J. Phys. Chem. Lett. 11, 6090–6096 (2020).
Schran, C., Brezina, K. & Marsalek, O. Committee neural network potentials control generalization errors and enable active learning. J. Chem. Phys 153, 104105 (2020).
Kulichenko, M. et al. Uncertainty-driven dynamics for active learning of interatomic potentials. Nat. Compu. Sci. 3, 230–239 (2023).
Wen, M. & Tadmor, E. B. Uncertainty quantification in molecular simulations with dropout neural network potentials. npj Comput. Mater. 6, 124 (2020).
Zhu, A., Batzner, S., Musaelian, A. & Kozinsky, B. Fast uncertainty estimates in deep learning interatomic potentials. J Chem. Phys. 158, 164111 (2023).
Thaler, S., Doehner, G. & Zavadlav, J. Scalable Bayesian uncertainty quantification for neural network potentials: promise and pitfalls. J. Chem. Theory Comput. 19, 4520–4532 (2023).
Soleimany, A. P. et al. Evidential deep learning for guided molecular property prediction and discovery. ACS Cent. Sci. 7, 1356–1367 (2021).
Wollschläger, T., Gao, N., Charpentier, B., Ketata, M. A. & Günnemann, S. Uncertainty estimation for molecules: Desiderata and methods. In International conference on machine learning, 37133–37156 (PMLR, 2023).
Tan, A. R., Urata, S., Goldman, S., Dietschreit, J. C. & Gómez-Bombarelli, R. Single-model uncertainty quantification in neural network potentials does not consistently outperform model ensembles. npj Computational Mater. 9, 225 (2023).
Xu, H. et al. Evidential deep learning for interatomic potentials. arXiv preprint arXiv:2407.13994 (2024).
Busk, J., Schmidt, M. N., Winther, O., Vegge, T. & Jørgensen, P. B. Graph neural network interatomic potential ensembles with calibrated aleatoric and epistemic uncertainty on energy and forces. Phys. Chem. Chem. Phys. 25, 25828–25837 (2023).
Carrete, J., Montes-Campos, H., Wanzenböck, R., Heid, E. & Madsen, G. K. Deep ensembles vs committees for uncertainty estimation in neural-network force fields: Comparison and application to active learning. J. Chem. Phys. 158, 204801 (2023).
Gawlikowski, J. et al. A survey of uncertainty in deep neural networks. Artif. Intell. Rev. 56, 1513–1589 (2023).
Kahle, L. & Zipoli, F. Quality of uncertainty estimates from neural network potential ensembles. Phys. Rev. E 105, 015311 (2022).
Thomas-Mitchell, A., Hawe, G. & Popelier, P. L. Calibration of uncertainty in the active learning of machine learning force fields. Mach. Learn.: Sci. Technol. 4, 045034 (2023).
Dai, J., Adhikari, S. & Wen, M. Uncertainty quantification and propagation in atomistic machine learning. Rev. Chem. Eng. https://doi.org/10.1515/revce-2024-0028 (2024).
Wan, S., Sinclair, R. C. & Coveney, P. V. Uncertainty quantification in classical molecular dynamics. Philos. Trans. R. Soc. A 379, 20200082 (2021).
Vassaux, M., Wan, S., Edeling, W. & Coveney, P. V. Ensembles are required to handle aleatoric and parametric uncertainty in molecular dynamics simulation. J. Chem. Theory Comput. 17, 5187–5197 (2021).
Henkel, P. & Mollenhauer, D. Uncertainty of exchange-correlation functionals in density functional theory calculations for lithium-based solid electrolytes on the case study of lithium phosphorus oxynitride. J. Computational Chem. 42, 1283–1295 (2021).
Goswami, S., Käser, S., Bemish, R. J. & Meuwly, M. Effects of aleatoric and epistemic errors in reference data on the learnability and quality of nn-based potential energy surfaces. Artif. Intell. Chem. 2, 100033 (2024).
Smith, J. S., Isayev, O. & Roitberg, A. E. Ani-1: an extensible neural network potential with dft accuracy at force field computational cost. Chem. Sci. 8, 3192–3203 (2017).
Gao, X., Ramezanghorbani, F., Isayev, O., Smith, J. S. & Roitberg, A. E. Torchani: a free and open source pytorch-based deep learning implementation of the ani neural network potentials. J. Chem. Inf. modeling 60, 3408–3415 (2020).
Zhang, S. et al. Exploring the frontiers of condensed-phase chemistry with a general reactive machine learning potential. Nat. Chem. 16, 727–734 (2024).
Allen, A. E. et al. Learning together: Towards foundation models for machine learning interatomic potentials with meta-learning. npj Computational Mater. 10, 154 (2024).
Deng, B. et al. Chgnet: Pretrained universal neural network potential for charge-informed atomistic modeling (2023). 2302.14231.
Batatia, I. et al. A foundation model for atomistic materials chemistry (2023). 2401.00096.
Chen, C. & Ong, S. P. A universal graph deep learning interatomic potential for the periodic table. Nat. Computational Sci. 2, 718–728 (2022).
Chanussot, L. et al. Open catalyst 2020 (oc20) dataset and community challenges. Acs Catal. 11, 6059–6072 (2021).
Kolluru, A. et al. Open challenges in developing generalizable large-scale machine-learning models for catalyst discovery. ACS Catal. 12, 8572–8581 (2022).
Tran, R. et al. The open catalyst 2022 (oc22) dataset and challenges for oxide electrocatalysts. ACS Catal. 13, 3066–3084 (2023).
Focassio, B., M. Freitas, L. P. & Schleder, G. R. Performance assessment of universal machine learning interatomic potentials: challenges and directions for materials’ surfaces. ACS Appl. Mater. Interfaces 17, 13111–13121 (2024).
Yu, H., Giantomassi, M., Materzanini, G., Wang, J. & Rignanese, G.-M. Systematic assessment of various universal machine-learning interatomic potentials. Mater. Genome Eng. Adv. 2, e58 (2024).
Deng, B. et al. Overcoming systematic softening in universal machine learning interatomic potentials by fine-tuning. arXiv preprint arXiv:2405.07105 (2024).
Niblett, S. P., Kourtis, P., Magdău, I.-B., Grey, C. P. & Csányi, G. Transferability of datasets between machine-learning interaction potentials. arXiv preprint arXiv:2409.05590 (2024).
Clausen, C. M., Rossmeisl, J. & Ulissi, Z. W. Adapting oc20-trained equiformerv2 models for high-entropy materials. J. Phys. Chem. C. 128, 11190–11195 (2024).
Abdar, M. et al. A review of uncertainty quantification in deep learning: Techniques, applications and challenges. Inf. fusion 76, 243–297 (2021).
Lopanitsyna, N., Fraux, G., Springer, M. A., De, S. & Ceriotti, M. Modeling high-entropy transition metal alloys with alchemical compression. Phys. Rev. Mater. 7, 045802 (2023).
Koenker, R. & Hallock, K. F. Quantile regression. J. Econ. Perspect. 15, 143–156 (2001).
Koenker, R. Fundamentals of Quantile Regression, 26–67. Econometric Society Monographs (Cambridge University Press, 2005).
Falcon, W. & The PyTorch Lightning team. PyTorch Lightning (Version 1.4) [Computer software]. (2019).
Koenker, R. Quantile regression. Cambridge Univ Pr (2005).
Takeuchi, I., Le, Q. V., Sears, T. D., Smola, A. J. & Williams, C. Nonparametric quantile estimation. J. Machine Learn. Res. 7, 1231–1264 (2006).
Zheng, S. Gradient descent algorithms for quantile regression with smooth approximation. Int. J. Mach. Learn. Cybern. 2, 191–207 (2011).
Hatalis, K., Lamadrid, A. J., Scheinberg, K. & Kishore, S. Smooth pinball neural network for probabilistic forecasting of wind power. arXiv preprint arXiv:1710.01720 (2017).
Acknowledgements
This work was supported by the "Transferring exascale computational chemistry to cloud computing environment and emerging hardware technologies (TEC4)" project, which is funded by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences, the Division of Chemical Sciences, Geosciences, and Biosciences (under FWP 82037). The simulation data used in this study was supported by the U.S. Department of Energy (DOE), Office of Science, Office of Basic Energy Sciences, Division of Chemical Sciences, Geosciences & Biosciences (under FWP 47319). Pacific Northwest National Laboratory (PNNL) is a multiprogram national laboratory operated for the U.S. Department of Energy (DOE) by Battelle Memorial Institute under Contract No. DE-AC05-76RL0-1830.
Author information
Authors and Affiliations
Contributions
J.A.B. and S.C. conceived of the idea. J.A.B. developed and implemented the method and carried out the experiments. J.S.F. curated and processed the datasets. M-S.L. created the H-ZSM-5 dataset. All authors contributed to the interpretation of the results and writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bilbrey, J.A., Firoz, J.S., Lee, MS. et al. Uncertainty quantification for neural network potential foundation models. npj Comput Mater 11, 109 (2025). https://doi.org/10.1038/s41524-025-01572-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41524-025-01572-y
This article is cited by
-
Emerging Technologies and Integrated Interdisciplinary Strategies for Mitigating Protein Aggregation in Therapeutic Formulations
Pharmaceutical Research (2026)
-
Neural network-driven molecular insights into alkaline wet etching of GaN: toward atomistic precision in nanostructure fabrication
npj Computational Materials (2025)
-
Heterogeneous ensemble enables a universal uncertainty metric for atomistic foundation models
npj Computational Materials (2025)
-
Residual bayesian attention networks for uncertainty quantification in regression tasks
Scientific Reports (2025)
-
Evidential deep learning for interatomic potentials
Nature Communications (2025)