Abstract
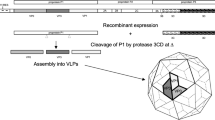
The inherent instability of poliovirus capsids presents a formidable challenge for developing next-generation vaccines suitable for a post-eradication world. Here, we address this by engineering a thermally stabilized virus-like particle (sVLP) derived from the poliovirus serotype 1 (PV1) Mahoney-SC7 mutant and elucidating its atomic-level structure. Produced at remarkably high yields in Pichia pastoris yeast, our engineered sVLP maintains a native, D-antigenic conformation and elicits a potent neutralizing antibody response in mice, in sharp contrast to unstable wild-type VLP (wtVLP) which adopts an expanded, non-immunogenic form. Our 2.43 Å resolution cryo-EM structure reveals precisely how seven stabilizing mutations cooperatively enhance inter-protomer contacts and rigidify surface loops to lock the particle in its immunogenic state. We further define a critical D-antigenic epitope by determining the 2.60 Å structure of the sVLP in complex with a novel D-antigen-specific, neutralizing monoclonal antibody, 3G10, elucidating the structural mechanisms of D-antigen recognition and virus neutralization by 3G10. These findings provide a definitive structural blueprint for engineering stable, immunogenic vaccines for PVs and other enteroviruses and also deliver a vital reagent for ensuring vaccine quality control.
Similar content being viewed by others
Data availability
Cryo-EM maps have been deposited in the Electron Microscopy Data Bank (http://www.emdataresource.org) with the accession numbers of EMD-65817 for wtVLP, EMD-65818 for sVLP, and EMD-65819 for sVLP-3G10. The associated models have been deposited to the Protein Data Bank (https://www.rcsb.org/) with accession codes 9WAG, 9WAH, and 9WAI.
References
Minor, P. D. An introduction to poliovirus: pathogenesis, vaccination, and the endgame for global eradication. Methods Mol. Biol. 1387, 1–10 (2016).
Hogle, J. M. Poliovirus cell entry: common structural themes in viral cell entry pathways. Annu Rev. Microbiol 56, 677–702 (2002).
Kitamura, N. et al. Primary structure, gene organization and polypeptide expression of poliovirus RNA. Nature 291, 547–553 (1981).
Hogle, J. M., Chow, M. & Filman, D. J. Three-dimensional structure of poliovirus at 2.9 A resolution. Science 229, 1358–1365 (1985).
Ypma-Wong, M. F., Dewalt, P. G., Johnson, V. H., Lamb, J. G. & Semler, B. L. Protein 3CD is the major poliovirus proteinase responsible for cleavage of the P1 capsid precursor. Virology 166, 265–270 (1988).
Hindiyeh, M., Li, Q. H., Basavappa, R., Hogle, J. M. & Chow, M. Poliovirus mutants at histidine 195 of VP2 do not cleave VP0 into VP2 and VP4. J. Virol. 73, 9072–9079 (1999).
Basavappa, R. et al. Role and mechanism of the maturation cleavage of VP0 in poliovirus assembly: structure of the empty capsid assembly intermediate at 2.9 A resolution. Protein Sci. 3, 1651–1669 (1994).
Jacobson, M. F. & Baltimore, D. Morphogenesis of poliovirus. I. Association viral RNA coat protein. J. Mol. Biol. 33, 369–378 (1968).
Hummeler, K. & Hamparian, V. V. Studies on the complement fixing antigens of poliomyelitis. I. Demonstration of type and group specific antigens in native and heated viral preparations. J. Immunol. 81, 499–505 (1958).
Mayer, M. M. et al. The purification of poliomyelitis virus as studied by complement fixation. J. Immunol. 78, 435–455 (1957).
Le Bouvier, G. L. The modification of poliovirus antigens by heat and ultraviolet light. Lancet 269, 1013–1016 (1955).
Strauss, M., Schotte, L., Karunatilaka, K. S., Filman, D. J. & Hogle, J. M. Cryo-electron microscopy structures of expanded poliovirus with VHHs sample the conformational repertoire of the expanded state. J. Virol. 91, e10443–16 (2017).
Minor, P. Vaccine-derived poliovirus (VDPV): impact on poliomyelitis eradication. Vaccine 27, 2649–2652 (2009).
Bandyopadhyay, A. S., Garon, J., Seib, K. & Orenstein, W. A. Polio vaccination: past, present and future. Future Microbiol. 10, 791–808 (2015).
Connor, R. I. et al. Mucosal immunity to poliovirus. Mucosal Immunol. 15, 1–9 (2022).
Duizer, E., Ruijs, W. L., van der Weijden, C. P. & Timen, A. Response to a wild poliovirus type 2 (WPV2)-shedding event following accidental exposure to WPV2, the Netherlands, April 2017. Euro Surveill. 22, 30542. https://doi.org/10.2807/1560-7917.ES.2017.22.21.30542 (2017).
Roldao, A., Mellado, M. C., Castilho, L. R., Carrondo, M. J. & Alves, P. M. Virus-like particles in vaccine development. Expert Rev. Vaccines 9, 1149–1176 (2010).
Kushnir, N., Streatfield, S. J. & Yusibov, V. Virus-like particles as a highly efficient vaccine platform: diversity of targets and production systems and advances in clinical development. Vaccine 31, 58–83 (2012).
Urakawa, T. et al. Synthesis of immunogenic, but non-infectious, poliovirus particles in insect cells by a baculovirus expression vector. J. Gen. Virol. 70, 1453–1463 (1989).
Jore, J. P. et al. Formation of poliomyelitis subviral particles in the yeast Saccharomyces cerevisiae. Yeast 10, 907–922 (1994).
Rombaut, B., Jore, J. & Boeye, A. A competition immunoprecipitation assay of unlabeled poliovirus antigens. J. Virol. Methods 48, 73–79 (1994).
Fox, H., Knowlson, S., Minor, P. D. & Macadam, A. J. Genetically thermo-stabilised, immunogenic poliovirus empty capsids; a strategy for non-replicating vaccines. PLoS Pathog. 13, e1006117 (2017).
Hong, Q. et al. Vaccine potency and structure of yeast-produced polio type 2 stabilized virus-like particles. Vaccines 12, 1077. https://doi.org/10.3390/vaccines12091077 (2024).
Marsian, J. et al. Plant-made polio type 3 stabilized VLPs-a candidate synthetic polio vaccine. Nat. Commun. 8, 245 (2017).
Xu, Y. et al. Virus-like particle vaccines for poliovirus types 1, 2, and 3 with enhanced thermostability expressed in insect cells. Vaccine 37, 2340–2347 (2019).
Bahar, M. W. et al. Mammalian expression of virus-like particles as a proof of principle for next generation polio vaccines. NPJ Vaccines 6, 5 (2021).
Sherry, L. et al. Production and Characterisation of Stabilised PV-3 Virus-like Particles Using Pichia pastoris. Viruses 14, 2159 (2022).
Sherry, L. et al. Recombinant expression systems for production of stabilised virus-like particles as next-generation polio vaccines. Nat. Commun. 16, 831 (2025).
Miller, S. T., Hogle, J. M. & Filman, D. J. phasing of high-symmetry macromolecular complexes:: Successful phasing of authentic poliovirus data to 3.0 Å resolution. J. Mol. Biol. 307, 499–512 (2001).
Shah, P. N. M. et al. Cryo-EM structures reveal two distinct conformational states in a picornavirus cell entry intermediate. PLoS Pathog. 16, e1008920 (2020).
Strauss, M. et al. Nectin-like interactions between poliovirus and its receptor trigger conformational changes associated with cell entry. J. Virol. 89, 4143–4157 (2015).
Sherry, L. et al. Production of an immunogenic trivalent poliovirus virus-like particle vaccine candidate in yeast using controlled fermentation. NPJ Vaccines 10, 64 (2025).
Sherry, L. et al. Protease-independent production of poliovirus virus-like particles in Pichia pastoris: implications for efficient vaccine development and insights into capsid assembly. Microbiol. Spectr. 11, e0430022 (2023).
Minor, P. et al. Principal and subsidiary antigenic sites of VP1 involved in the neutralization of poliovirus type 3. J. Gen. Virol. 66, 1159–1165 (1985).
Charnesky, A. J. et al. A human monoclonal antibody binds within the poliovirus receptor-binding site to neutralize all three serotypes. Nat. Commun. 14, 6335 (2023).
Fiore, L. et al. Poliovirus Sabin type 1 neutralization epitopes recognized by immunoglobulin A monoclonal antibodies. J. Virol. 71, 6905–6912 (1997).
Diamond, D. C. et al. Antigenic variation and resistance to neutralization in poliovirus type 1. Science 229, 1090–1093 (1985).
Minor, P. D., Ferguson, M., Evans, D. M., Almond, J. W. & Icenogle, J. P. Antigenic structure of polioviruses of serotypes 1, 2 and 3. J. Gen. Virol. 67, 1283–1291 (1986).
Page, G. et al. Three-dimensional structure of poliovirus serotype 1 neutralizing determinants. J. Virol. 62, 1781–1794 (1988).
Puligedda, R. D. et al. Characterization of human monoclonal antibodies that neutralize multiple poliovirus serotypes. Vaccine 35, 5455–5462 (2017).
Liu, Q. et al. Detection, characterization and quantitation of coxsackievirus A16 using polyclonal antibodies against recombinant capsid subunit proteins. J. Virol. Methods 173, 115–120 (2011).
Zhang, C. et al. High-yield production of recombinant virus-like particles of enterovirus 71 in Pichia pastoris and their protective efficacy against oral viral challenge in mice. Vaccine 33, 2335–2341 (2015).
Ye, J., Ma, N., Madden, T. L. & Ostell, J. M. IgBLAST: an immunoglobulin variable domain sequence analysis tool. Nucleic Acids Res. 41, W34–W40 (2013).
Yi, Y. et al. Identification of human norovirus GII.3 blockade antibody epitopes. Viruses 13, 2058 (2021).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, W252–W258 (2014).
Miller, S. T., Hogle, J. M. & Filman, D. J. Ab initio phasing of high-symmetry macromolecular complexes: successful phasing of authentic poliovirus data to 3.0 A resolution. J. Mol. Biol. 307, 499–512 (2001).
DiMaio, F. et al. Atomic-accuracy models from 4.5-A cryo-electron microscopy data with density-guided iterative local refinement. Nat. Methods 12, 361–365 (2015).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Schlee, S. et al. Prediction of quaternary structure by analysis of hot spot residues in protein-protein interfaces: the case of anthranilate phosphoribosyltransferases. Proteins 87, 815–825 (2019).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Acknowledgements
We are grateful to the staff of the NCPSS Electron Microscopy Facility, Database and Computing Facility, and Protein Expression and Purification Facility for instrument support and technical assistance. This work was supported by grants from the National Natural Science Foundation of China (32370991 to Z.H.), National Key R&D Program of China (2024YFA1803102 to Y.C.), the Shanghai Municipal Science and Technology Major Project (ZD2021CY001 to Z.H.), the Strategic Priority Research Program of CAS (XDB0570000 to Y.C.), the NSFC (32130056, 32570924 to Y.C.), and the Shanghai Pilot Program for Basic Research from CAS (JCYJ-SHFY-2022-008 to Y.C.). Q.H. was supported by the China National Postdoctoral Program for Innovative Talents (BX20250141).
Author information
Authors and Affiliations
Contributions
Z.H., Y.C., X.W., and Q.L. designed the experiments. T.C., W.H., S.W., C.Y.L. (Chaoyang Lian), Y.Z., L.R., T.W., and C.S. performed the biochemical and animal experiments. Q.H. collected the cryo-EM data and performed cryo-EM reconstructions and model building. C.L. is involved in the initial cryo-EM sample screen. Q.H., T.C., and W.H. analyzed the data. Z.H., Y.C., X.W., Q.L., Q.H., and T.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
W.H., C.L., L.R., Q.L., and X.W. are current employees of Huasong (Shanghai) Biomedical Technology Co., LTD. Q.L., X.W., L.R., T.C., and W.H. are inventors of a patent (CN 202410797193.1) on the 5C6 mAb owned by Huasong (Shanghai) Biomedical Technology Co., LTD. Q.L., X.W., L.R., C.Y.L., Z.H., and T.C. are listed as inventors in a pending patent application (CN 202511196350.4) on the 3G10 mAb filed by Huasong (Shanghai) Biomedical Technology Co., LTD. The other authors declare that they have no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hong, Q., Chen, T., Han, W. et al. Structural insight into the assembly and D antigenicity of polio type 1 stabilized virus-like particles. npj Vaccines (2026). https://doi.org/10.1038/s41541-026-01404-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-026-01404-0