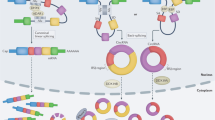
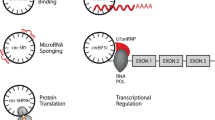
Abstract
Over the past decade, circular RNA (circRNA) research has evolved into a bona fide research field shedding light on the functional consequence of this unique family of RNA molecules in cancer. Although the method of formation and the abundance of circRNAs can differ from their cognate linear mRNA, the spectrum of interacting partners and their resultant cellular functions in oncogenesis are analogous. However, with 10 times more diversity in circRNA variants compared with linear RNA variants, combined with their hyperstability in the cell, circRNAs are equipped to influence every stage of oncogenesis. This is an opportune time to address the breadth of circRNA in cancer focused on their spatiotemporal expression, mutations in biogenesis factors and contemporary functions through each stage of cancer. In this Review, we highlight examples of functional circRNAs in specific cancers, which satisfy critical criteria, including their physical co-association with the target and circRNA abundance at stoichiometrically valid quantities. These considerations are essential to develop strategies for the therapeutic exploitation of circRNAs as biomarkers and targeted anticancer agents.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Yoshida, K. et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 478, 64–69 (2011).
Seiler, M. et al. Somatic mutational landscape of splicing factor genes and their functional consequences across 33 cancer types. Cell Rep. 23, 282–296.e4 (2018).
Suzuki, H. et al. Recurrent noncoding U1 snRNA mutations drive cryptic splicing in SHH medulloblastoma. Nature 574, 707–711 (2019).
Shuai, S. et al. The U1 spliceosomal RNA is recurrently mutated in multiple cancers. Nature 574, 712–716 (2019).
Wedge, E. et al. Impact of U2AF1 mutations on circular RNA expression in myelodysplastic neoplasms. Leukemia 37, 1113–1125 (2023).
Conn, V. M. et al. Circular RNAs drive oncogenic chromosomal translocations within the MLL recombinome in leukemia. Cancer Cell 41, 1309–1326.e10 (2023). This paper presents the first example of an endogenous RNA carcinogen; a circular RNA able to drive oncogenic chromosomal translocations in acute leukaemia.
Trsova, I. et al. Expression of circular RNAs in myelodysplastic neoplasms and their association with mutations in the splicing factor gene SF3B1. Mol. Oncol. 17, 2565–2583 (2023).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013). Together with Memczak et al. (2013), this paper invigorated the circular RNA (circRNA) research field by identifying functional circRNAs acting as microRNA sponges.
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013). Together with Hansen et al. (2013), this paper invigorated the circular RNA (circRNA) research field by identifying functional circRNAs acting as microRNA sponges.
Bradley, R. K. & Anczuków, O. RNA splicing dysregulation and the hallmarks of cancer. Nat. Rev. Cancer 23, 135–155 (2023).
Marasco, L. E. & Kornblihtt, A. R. The physiology of alternative splicing. Nat. Rev. Mol. Cell Biol. 24, 242–254 (2023).
Chen, L.-L. & Yang, L. Regulation of circRNA biogenesis. RNA Biol. 12, 381–388 (2015).
Conn, V. M. et al. SRRM4 expands the repertoire of circular RNAs by regulating microexon inclusion. Cells 9, 2488 (2020).
Chiang, T.-W. et al. FL-circAS: an integrative resource and analysis for full-length sequences and alternative splicing of circular RNAs with nanopore sequencing. Nucleic Acids Res. 52, D115–D123 (2024).
Vo, J. N. et al. The landscape of circular RNA in cancer. Cell 176, 869–881.e13 (2019). Vast profiling of circular RNAs (circRNAs) across more than 2,000 samples from patients with cancer, which produced a searchable public database called miOncoCirc to aid in the identification of circRNA biomarkers.
Liang, D. et al. The output of protein-coding genes shifts to circular RNAs when the pre-mRNA processing machinery is limiting. Mol. Cell 68, 940–954.e3 (2017).
Guarnerio, J. et al. Oncogenic role of fusion-circRNAs derived from cancer-associated chromosomal translocations. Cell 165, 289–302 (2016).
Liu, X. et al. Identification of mecciRNAs and their roles in the mitochondrial entry of proteins. Sci. China Life Sci. 63, 1429–1449 (2020).
Liu, X. et al. Interior circular RNA. RNA Biol. 17, 87–97 (2020).
Li, Z. et al. Exon–intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 22, 256–264 (2015).
Zhang, J. et al. Comprehensive profiling of circular RNAs with nanopore sequencing and CIRI-long. Nat. Biotechnol. 39, 836–845 (2021).
Talhouarne, G. J. S. & Gall, J. G. Lariat intronic RNAs in the cytoplasm of vertebrate cells. Proc. Natl Acad. Sci. USA 115, E7970–E7977 (2018).
Zhang, Y. et al. Circular intronic long noncoding RNAs. Mol. Cell 51, 792–806 (2013).
Lu, Z. et al. Metazoan tRNA introns generate stable circular RNAs in vivo. RNA 21, 1554–1565 (2015).
Gao, Y., Wang, J. & Zhao, F. CIRI: an efficient and unbiased algorithm for de novo circular RNA identification. Genome Biol. 16, 4 (2015).
Chen, L.-L. et al. A guide to naming eukaryotic circular RNAs. Nat. Cell Biol. 25, 1–5 (2023).
Liang, D. & Wilusz, J. E. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 28, 2233–2247 (2014).
Ashwal-Fluss, R. et al. CircRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 56, 55–66 (2014).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013). A seminal paper identifying circular RNAs (circRNAs) as an abundant RNA family, with insight into Alu-driven circRNA biogenesis.
Conn, S. J. et al. The RNA binding protein quaking regulates formation of circRNAs. Cell 160, 1125–1134 (2015).
Errichelli, L. et al. FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons. Nat. Commun. 8, 14741 (2017).
Stagsted, L. V. W., O’Leary, E. T., Ebbesen, K. K. & Hansen, T. B. The RNA-binding protein SFPQ preserves long-intron splicing and regulates circRNA biogenesis in mammals. eLife 10, e63088 (2021).
Li, X. et al. Coordinated circRNA biogenesis and function with NF90/NF110 in viral infection. Mol. Cell 67, 214–227.e7 (2017).
Knupp, D., Cooper, D. A., Saito, Y., Darnell, R. B. & Miura, P. NOVA2 regulates neural circRNA biogenesis. Nucleic Acids Res. 49, 6849–6862 (2021).
Aktaş, T. et al. DHX9 suppresses RNA processing defects originating from the Alu invasion of the human genome. Nature 544, 115–119 (2017).
Ivanov, A. et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 10, 170–177 (2015).
Rybak-Wolf, A. et al. Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol. Cell 58, 870–885 (2015).
Yang, L., Wilusz, J. E. & Chen, L.-L. Biogenesis and regulatory roles of circular RNAs. Annu. Rev. Cell Dev. Biol. 38, 263–289 (2022).
Kristensen, L. S. et al. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 20, 675–691 (2019).
Goodall, G. J. & Wickramasinghe, V. O. RNA in cancer. Nat. Rev. Cancer 21, 22–36 (2021).
Wang, Z.-L. et al. Comprehensive genomic characterization of RNA-binding proteins across human cancers. Cell Rep. 22, 286–298 (2018).
Pillman, K. A. et al. miR-200/375 control epithelial plasticity-associated alternative splicing by repressing the RNA-binding protein Quaking. EMBO J. 37, e99016 (2018).
Wang, J.-Z. et al. QKI-5 regulates the alternative splicing of cytoskeletal gene ADD3 in lung cancer. J. Mol. Cell Biol. 13, 347–360 (2021).
Rabbitts, T. H., Forster, A., Larson, R. & Nathan, P. Fusion of the dominant negative transcription regulator CHOP with a novel gene FUS by translocation t(12;16) in malignant liposarcoma. Nat. Genet. 4, 175–180 (1993).
Owen, I. et al. The oncogenic transcription factor FUS-CHOP can undergo nuclear liquid–liquid phase separation. J. Cell Sci. 134, jcs258578 (2021).
Rezvykh, A. P. et al. Cytoplasmic aggregation of mutant FUS causes multistep RNA splicing perturbations in the course of motor neuron pathology. Nucleic Acids Res. 51, 5810–5830 (2023).
Colantoni, A. et al. FUS alters circRNA metabolism in human motor neurons carrying the ALS-linked P525L mutation. Int. J. Mol. Sci. 24, 3181 (2023).
Quesnel-Vallières, M., Irimia, M., Cordes, S. P. & Blencowe, B. J. Essential roles for the splicing regulator nSR100/SRRM4 during nervous system development. Genes Dev. 29, 746 (2015).
Head, S. A. et al. Silencing of SRRM4 suppresses microexon inclusion and promotes tumor growth across cancers. PLoS Biol. 19, e3001138 (2021).
Li, Y. et al. SRRM4 drives neuroendocrine transdifferentiation of prostate adenocarcinoma under androgen receptor pathway inhibition. Eur. Urol. 71, 68–78 (2017).
Deshpande, K. et al. SRRM4-mediated REST to REST4 dysregulation promotes tumor growth and neural adaptation in breast cancer leading to brain metastasis. Neuro Oncol. 26, 309–322 (2024).
Ren, L. et al. Mechanisms of circular RNA degradation. Commun. Biol. 5, 1–6 (2022).
Enuka, Y. et al. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 44, 1370–1383 (2016).
Liu, C.-X. et al. Structure and degradation of circular RNAs regulate PKR activation in innate immunity. Cell 177, 865–880.e21 (2019).
Li, X. et al. Linking circular intronic RNA degradation and function in transcription by RNase H1. Sci. China Life Sci. 64, 1795–1809 (2021).
Park, O. H. et al. Endoribonucleolytic cleavage of m6A-containing RNAs by RNase P/MRP complex. Mol. Cell 74, 494–507.e8 (2019).
Casey, G. et al. RNASEL Arg462Gln variant is implicated in up to 13% of prostate cancer cases. Nat. Genet. 32, 581–583 (2002).
Yi, C. et al. A pan-cancer analysis of RNASEH1, a potential regulator of the tumor microenvironment. Clin. Transl. Oncol. 25, 2569–2586 (2023).
Wahba, L., Amon, J. D., Koshland, D. & Vuica-Ross, M. RNase H and multiple RNA biogenesis factors cooperate to prevent RNA–DNA hybrids from generating genome instability. Mol. Cell 44, 978–988 (2011).
Fischer, J. W., Busa, V. F., Shao, Y. & Leung, A. K. L. Structure-mediated RNA decay by UPF1 and G3BP1. Mol. Cell 78, 70–84.e6 (2020).
Schmid, M. & Jensen, T. H. The exosome: a multipurpose RNA-decay machine. Trends Biochem. Sci. 33, 501–510 (2008).
Lasda, E. & Parker, R. Circular RNAs co-precipitate with extracellular vesicles: a possible mechanism for circRNA clearance. PLoS ONE 11, e0148407 (2016).
Zhang, F., Jiang, J., Qian, H., Yan, Y. & Xu, W. Exosomal circRNA: emerging insights into cancer progression and clinical application potential. J. Hematol. Oncol. 16, 67 (2023).
Alhasan, A. A. et al. Circular RNA enrichment in platelets is a signature of transcriptome degradation. Blood 127, e1–e11 (2016).
Preußer, C. et al. Selective release of circRNAs in platelet-derived extracellular vesicles. J. Extracell. Vesicles 7, 1424473 (2018).
Chen, L. et al. Exportin 4 depletion leads to nuclear accumulation of a subset of circular RNAs. Nat. Commun. 13, 5769 (2022).
Ngo, L. H. et al. Nuclear export of circular RNA. Nature 627, 212–220 (2024).
Huang, C., Liang, D., Tatomer, D. C. & Wilusz, J. E. A length-dependent evolutionarily conserved pathway controls nuclear export of circular RNAs. Genes Dev. 32, 639–644 (2018). An elegant seminal report demonstrating that nuclear circular RNA (circRNA) export is dictated by the length of the mature circRNAs.
Hu, Z. et al. Exosome-derived circCCAR1 promotes CD8+ T-cell dysfunction and anti-PD1 resistance in hepatocellular carcinoma. Mol. Cancer 22, 55 (2023).
Pan, Z. et al. EWSR1-induced circNEIL3 promotes glioma progression and exosome-mediated macrophage immunosuppressive polarization via stabilizing IGF2BP3. Mol. Cancer 21, 16 (2022).
Bouwman, B. A. M., Crosetto, N. & Bienko, M. RNA gradients: shapers of 3D genome architecture. Curr. Opin. Cell Biol. 74, 7–12 (2022).
Chen, C.-K. et al. Structured elements drive extensive circular RNA translation. Mol. Cell 81, 4300–4318.e13 (2021).
Fan, X., Yang, W., Chen, C. & Wang, Z. Pervasive translation of circular RNAs driven by short IRES-like elements. Nat. Commun. 13, 3751 (2022).
Dodbele, S., Mutlu, N. & Wilusz, J. E. Best practices to ensure robust investigation of circular RNAs: pitfalls and tips. EMBO Rep. 22, e52072 (2021).
Nielsen, A. F. et al. Best practice standards for circular RNA research. Nat. Methods 19, 1208–1220 (2022).
Koppula, A., Abdelgawad, A., Guarnerio, J., Batish, M. & Parashar, V. CircFISH: a novel method for the simultaneous imaging of linear and circular RNAs. Cancers 14, 428 (2022).
Ulshöfer, C. J., Pfafenrot, C., Bindereif, A. & Schneider, T. Methods to study circRNA–protein interactions. Methods 196, 36–46 (2021).
Gabryelska, M. M. et al. Native circular RNA pulldown method to simultaneously profile RNA and protein interactions. Methods Mol. Biol. 2765, 299–309 (2024).
Xu, X. et al. CircRNA inhibits DNA damage repair by interacting with host gene. Mol. Cancer 19, 128 (2020).
Yi, S., Singh, S. S., Rozen-Gagnon, K. & Luna, J. M. Mapping RNA–protein interactions with subcellular resolution using colocalization CLIP. RNA 30, 920–937 (2024).
Kocks, C., Boltengagen, A., Piwecka, M., Rybak-Wolf, A. & Rajewsky, N. Single-molecule fluorescence in situ hybridization (FISH) of circular RNA CDR1as. Methods Mol. Biol. 1724, 77–96 (2018).
Zhang, J. et al. Circular RNA profiling provides insights into their subcellular distribution and molecular characteristics in HepG2 cells. RNA Biol. 16, 220–232 (2019).
Zender, L. et al. An oncogenomics-based in vivo RNAi screen identifies tumor suppressors in liver cancer. Cell 135, 852–864 (2008).
Zhang, H. et al. DDX39B contributes to the proliferation of colorectal cancer through direct binding to CDK6/CCND1. Cell Death Discov. 8, 30 (2022).
Xu, Z. et al. Suppression of DDX39B sensitizes ovarian cancer cells to DNA-damaging chemotherapeutic agents via destabilizing BRCA1 mRNA. Oncogene 39, 7051–7062 (2020).
Cai, W. et al. Wanted DEAD/H or alive: helicases winding up in cancers. J. Natl Cancer Inst. 109, djw278 (2017).
Fuller-Pace, F. V. DEAD box RNA helicase functions in cancer. RNA Biol. 10, 121–132 (2013).
Yoo, H. H. & Chung, I. K. Requirement of DDX39 DEAD box RNA helicase for genome integrity and telomere protection. Aging Cell 10, 557–571 (2011).
Bao, Y. et al. DDX39 as a predictor of clinical prognosis and immune checkpoint therapy efficacy in patients with clear cell renal cell carcinoma. Int. J. Biol. Sci. 17, 3158–3172 (2021).
Kuramitsu, Y. et al. Up-regulation of DDX39 in human malignant pleural mesothelioma cell lines compared to normal pleural mesothelial cells. Anticancer Res. 33, 2557–2560 (2013).
Sugiura, T., Sakurai, K. & Nagano, Y. Intracellular characterization of DDX39, a novel growth-associated RNA helicase. Exp. Cell Res. 313, 782–790 (2007).
Xing, C. et al. DDX39 overexpression predicts a poor prognosis and promotes aggressiveness of melanoma by cooperating with SNAIL. Front. Oncol. 10, 1261 (2020).
Bissels, U. et al. Absolute quantification of microRNAs by using a universal reference. RNA 15, 2375–2384 (2009).
Calabrese, J. M., Seila, A. C., Yeo, G. W. & Sharp, P. A. RNA sequence analysis defines Dicer’s role in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 104, 18097–18102 (2007).
Denzler, R., Agarwal, V., Stefano, J., Bartel, D. P. & Stoffel, M. Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Mol. Cell 54, 766–776 (2014).
Mukherji, S. et al. MicroRNAs can generate thresholds in target gene expression. Nat. Genet. 43, 854–859 (2011).
Li, S. et al. Screening for functional circular RNAs using the CRISPR–Cas13 system. Nat. Methods 18, 51–59 (2021).
Ma, X.-K. et al. CIRCexplorer3: a CLEAR pipeline for direct comparison of circular and linear RNA expression. Genomics Proteom. Bioinform. 17, 511–521 (2019).
Zheng, Q. et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat. Commun. 7, 11215 (2016).
Panda, A. C. et al. Identification of senescence-associated circular RNAs (SAC-RNAs) reveals senescence suppressor CircPVT1. Nucleic Acids Res. 45, 4021–4035 (2017).
Militello, G. et al. Screening and validation of lncRNAs and circRNAs as miRNA sponges. Brief. Bioinform. 18, 780–788 (2017).
Ala, U. et al. Integrated transcriptional and competitive endogenous RNA networks are cross-regulated in permissive molecular environments. Proc. Natl Acad. Sci. USA 110, 7154–7159 (2013).
Figliuzzi, M., De Martino, A. & Marinari, E. RNA-based regulation: dynamics and response to perturbations of competing RNAs. Biophys. J. 107, 1011–1022 (2014).
Wu, W., Zhang, J., Cao, X., Cai, Z. & Zhao, F. Exploring the cellular landscape of circular RNAs using full-length single-cell RNA sequencing. Nat. Commun. 13, 3242 (2022). The first report detailing the circular RNA (circRNA) landscape using single-cell sequencing, illuminating intertumour and intratumour heterogeneity of circRNAs across 20 patients with breast cancer.
Zhou, Z. et al. CIRI-deep enables single-cell and spatial transcriptomic analysis of circular RNAs with deep learning. Adv. Sci. 11, e2308115 (2024).
Zhong, C., Yu, S., Han, M., Chen, J. & Ning, K. Heterogeneous circRNA expression profiles and regulatory functions among HEK293T single cells. Sci. Rep. 7, 14393 (2017).
Fan, X. et al. Single-cell RNA-seq transcriptome analysis of linear and circular RNAs in mouse preimplantation embryos. Genome Biol. 16, 148 (2015).
Liu, B. et al. Cytoskeleton remodeling mediated by circRNA-YBX1 phase separation suppresses the metastasis of liver cancer. Proc. Natl Acad. Sci. USA 120, e2220296120 (2023).
Chen, S. et al. circVAMP3 drives CAPRIN1 phase separation and inhibits hepatocellular carcinoma by suppressing c‐Myc translation. Adv. Sci. 9, 2103817 (2022).
Piwecka, M. et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 357, eaam8526 (2017).
Kristensen, L. S. et al. Spatial expression analyses of the putative oncogene ciRS-7 in cancer reshape the microRNA sponge theory. Nat. Commun. 11, 4551 (2020).
García-Rodríguez, J. L. et al. Spatial profiling of circular RNAs in cancer reveals high expression in muscle and stromal cells. Cancer Res. 83, 3340–3353 (2023).
Legnini, I. et al. Circ-ZNF609 is a circular RNA that can be translated and functions in myogenesis. Mol. Cell 66, 22–37.e9 (2017). This paper reports for the first time the translation of a circular RNA, circZNF609, to a functional protein that is detectable without transgenic overexpression.
Yang, Y. et al. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res. 27, 626–641 (2017).
Pamudurti, N. R. et al. Translation of circRNAs. Mol. Cell 66, 9–21.e7 (2017).
Ho-Xuan, H. et al. Comprehensive analysis of translation from overexpressed circular RNAs reveals pervasive translation from linear transcripts. Nucleic Acids Res. 48, 10368–10382 (2020).
Hansen, T. B. Signal and noise in circRNA translation. Methods 196, 68–73 (2021).
Nicolet, B. P. et al. Circular RNAs exhibit limited evidence for translation, or translation regulation of the mRNA counterpart in terminal hematopoiesis. RNA 28, 194–209 (2022).
Stagsted, L. V., Nielsen, K. M., Daugaard, I. & Hansen, T. B. Noncoding AUG circRNAs constitute an abundant and conserved subclass of circles. Life Sci. Alliance 2, e201900398 (2019).
Califano, A. & Alvarez, M. J. The recurrent architecture of tumour initiation, progression and drug sensitivity. Nat. Rev. Cancer 17, 116–130 (2017).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the cancer genome atlas. Cell 173, 321–337.e10 (2018).
Martínez-Jiménez, F. et al. A compendium of mutational cancer driver genes. Nat. Rev. Cancer 20, 555–572 (2020).
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489 (2015).
Meyer, C. et al. The KMT2A recombinome of acute leukemias in 2023. Leukemia 37, 988–1005 (2023).
Dal Molin, A. et al. CircRNAs are here to stay: a perspective on the MLL recombinome. Front. Genet. 10, 88 (2019).
Borden, K. L. B. The nuclear pore complex and mRNA export in cancer. Cancers 13, 42 (2020).
Guan, H., Luo, W., Liu, Y. & Li, M. Novel circular RNA circSLIT2 facilitates the aerobic glycolysis of pancreatic ductal adenocarcinoma via miR-510-5p/c-Myc/LDHA axis. Cell Death Dis. 12, 1–9 (2021).
Miller, D. M., Thomas, S. D., Islam, A., Muench, D. & Sedoris, K. c-Myc and cancer metabolism. Clin. Cancer Res. 18, 5546–5553 (2012).
Zhan, W. et al. circCTIC1 promotes the self-renewal of colon TICs through BPTF-dependent c-Myc expression. Carcinogenesis 40, 560–568 (2019).
Ren, X., Yu, J., Guo, L. & Ma, H. Circular RNA circRHOT1 contributes to pathogenesis of non-small cell lung cancer by epigenetically enhancing C-MYC expression through recruiting KAT5. Aging 13, 20372–20382 (2021).
Wang, X. et al. The circACTN4 interacts with FUBP1 to promote tumorigenesis and progression of breast cancer by regulating the expression of proto-oncogene MYC. Mol. Cancer 20, 91 (2021).
Xie, F. et al. circNR3C1 suppresses bladder cancer progression through acting as an endogenous blocker of BRD4/C-myc complex. Mol. Ther. Nucleic Acids 22, 510–519 (2020).
Yang, Q. et al. A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ. 24, 1609–1620 (2017).
Shen, S. et al. CircECE1 activates energy metabolism in osteosarcoma by stabilizing c-Myc. Mol. Cancer 19, 151 (2020).
Warburg, O. Ober den stoffwechsel der carcmomzelle j. Naturwissenschaften 12, 1131–1137 (1924).
Mo, Y. et al. Circular RNA circRNF13 inhibits proliferation and metastasis of nasopharyngeal carcinoma via SUMO2. Mol. Cancer 20, 112 (2021).
Shangguan, H. et al. Circular RNA circSLC25A16 contributes to the glycolysis of non-small-cell lung cancer through epigenetic modification. Cell Death Dis. 11, 1–10 (2020).
Li, Q. et al. CircACC1 regulates assembly and activation of AMPK complex under metabolic stress. Cell Metab. 30, 157–173.e7 (2019).
Li, J. et al. CIRCRPN2 inhibits aerobic glycolysis and metastasis in hepatocellular carcinoma. Cancer Res. 82, 1055–1069 (2022).
Cai, J. et al. CircRHBDD1 augments metabolic rewiring and restricts immunotherapy efficacy via m6A modification in hepatocellular carcinoma. Mol. Ther. Oncol. 24, 755–771 (2022).
Du, W. W. et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 24, 357–370 (2017).
Ding, L. et al. Circular RNA circ-DONSON facilitates gastric cancer growth and invasion via NURF complex dependent activation of transcription factor SOX4. Mol. Cancer 18, 45 (2019).
Yang, G. et al. circ-BIRC6, a circular RNA, promotes hepatocellular carcinoma progression by targeting the miR-3918/Bcl2 axis. Cell Cycle 18, 976–989 (2019).
Kontos, C. K. et al. Novel circular RNAs of the apoptosis‐related BAX and BCL2L12 genes identified in a chronic lymphocytic leukemia cell line using nanopore sequencing. FEBS Open Bio 13, 1953–1966 (2023).
Forster, J. C., Harriss-Phillips, W. M., Douglass, M. J. & Bezak, E. A review of the development of tumor vasculature and its effects on the tumor microenvironment. Hypoxia 5, 21–32 (2017).
Garcia, J. et al. Bevacizumab (Avastin®) in cancer treatment: a review of 15 years of clinical experience and future outlook. Cancer Treat. Rev. 86, 102017 (2020).
Hendrix, M. J. C., Seftor, E. A., Hess, A. R. & Seftor, R. E. B. Vasculogenic mimicry and tumour-cell plasticity: lessons from melanoma. Nat. Rev. Cancer 3, 411–421 (2003).
Jiang, S. et al. CircRNA-mediated regulation of angiogenesis: a new chapter in cancer biology. Front. Oncol. 11, 553706 (2021).
Shao, Y. & Lu, B. The emerging roles of circular RNAs in vessel co-option and vasculogenic mimicry: clinical insights for anti-angiogenic therapy in cancers. Cancer Metastasis Rev. 41, 173–191 (2022).
He, Q. et al. circ-SHKBP1 regulates the angiogenesis of U87 glioma-exposed endothelial cells through miR-544a/FOXP1 and miR-379/FOXP2 pathways. Mol. Ther. Nucleic Acids 10, 331–348 (2017).
Barbagallo, D. et al. The GAUGAA motif is responsible for the binding between circSMARCA5 and SRSF1 and related downstream effects on glioblastoma multiforme cell migration and angiogenic potential. Int. J. Mol. Sci. 22, 1678 (2021).
Barbagallo, D. et al. CircSMARCA5 regulates VEGFA mRNA splicing and angiogenesis in glioblastoma multiforme through the binding of SRSF1. Cancers 11, 194 (2019).
Geng, Q. et al. CircSMARCA5 silencing impairs cell proliferation and invasion via the miR-17-3p-EGFR signaling in lung adenocarcinoma. Life Sci. 320, 121560 (2023).
De Luca, A. et al. The role of the EGFR signaling in tumor microenvironment. J. Cell Physiol. 214, 559–567 (2008).
Klein, C. A. Cancer progression and the invisible phase of metastatic colonization. Nat. Rev. Cancer 20, 681–694 (2020).
Fares, J., Fares, M. Y., Khachfe, H. H., Salhab, H. A. & Fares, Y. Molecular principles of metastasis: a hallmark of cancer revisited. Signal. Transduct. Target. Ther. 5, 28 (2020).
Deng, J. et al. Specific intracellular retention of circSKA3 promotes colorectal cancer metastasis by attenuating ubiquitination and degradation of SLUG. Cell Death Dis. 14, 750 (2023).
Li, K. et al. A circular RNA activated by TGFβ promotes tumor metastasis through enhancing IGF2BP3-mediated PDPN mRNA stability. Nat. Commun. 14, 6876 (2023).
Hanniford, D. et al. Epigenetic silencing of CDR1as drives IGF2BP3-mediated melanoma invasion and metastasis. Cancer Cell 37, 55–70.e15 (2020).
Li, J. et al. Circular RNA IARS (circ-IARS) secreted by pancreatic cancer cells and located within exosomes regulates endothelial monolayer permeability to promote tumor metastasis. J. Exp. Clin. Cancer Res. 37, 177 (2018).
Wang, X. et al. CircURI1 interacts with hnRNPM to inhibit metastasis by modulating alternative splicing in gastric cancer. Proc. Natl Acad. Sci. USA 118, e2012881118 (2021).
Sun, L. et al. Association between duration of immunotherapy and overall survival in advanced non-small cell lung cancer. JAMA Oncol. 9, 1075–1082 (2023).
Liu, X.-Y. et al. The role of circular RNAs in the drug resistance of cancers. Front. Oncol. 11, 790586 (2022).
Chao, H.-M. et al. Y-box binding protein-1 promotes hepatocellular carcinoma-initiating cell progression and tumorigenesis via Wnt/β–catenin pathway. Oncotarget 8, 2604–2616 (2017).
Wilhelm, S. et al. Discovery and development of sorafenib: a multikinase inhibitor for treating cancer. Nat. Rev. Drug Discov. 5, 835–844 (2006).
Xu, J. et al. CircRNA-SORE mediates sorafenib resistance in hepatocellular carcinoma by stabilizing YBX1. Signal. Transduct. Target. Ther. 5, 298 (2020).
Wang, X. et al. CircRNA-CREIT inhibits stress granule assembly and overcomes doxorubicin resistance in TNBC by destabilizing PKR. J. Hematol. Oncol. 15, 122 (2022).
Zheng, S. et al. Extracellular vesicle-packaged circBIRC6 from cancer-associated fibroblasts induce platinum resistance via SUMOylation modulation in pancreatic cancer. J. Exp. Clin. Cancer Res. 42, 324 (2023).
Gao, X. et al. Circular RNA-encoded oncogenic E-cadherin variant promotes glioblastoma tumorigenicity through activation of EGFR-STAT3 signalling. Nat. Cell Biol. 23, 278–291 (2021).
Ma, Y., Wang, T., Zhang, X., Wang, P. & Long, F. The role of circular RNAs in regulating resistance to cancer immunotherapy: mechanisms and implications. Cell Death Dis. 15, 312 (2024).
Liu, T., Zhang, L., Joo, D. & Sun, S.-C. NF-κB signaling in inflammation. Sig. Transduct. Target. Ther. 2, 1–9 (2017).
Zhou, C. et al. Mutant KRAS-activated circATXN7 fosters tumor immunoescape by sensitizing tumor-specific T cells to activation-induced cell death. Nat. Commun. 15, 499 (2024). This article details an abundant circular RNA, which mediates crosstalk among mutant KRAS, NF-κB inactivation and immunotherapy resistance in multiple cancers.
Tang, Y., Zhou, L. & Liu, L. Circ_0085616 contributes to the radio-resistance and progression of cervical cancer by targeting miR-541-3p/ARL2 signaling. Histol. Histopathol. 38, 571–584 (2023).
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N. & Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 7, e30733 (2012).
Meng, S. et al. CircRNA: functions and properties of a novel potential biomarker for cancer. Mol. Cancer 16, 94 (2017).
Memczak, S., Papavasileiou, P., Peters, O. & Rajewsky, N. Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLoS ONE 10, e0141214 (2015).
Wu, Z. et al. Circ-RPL15: a plasma circular RNA as novel oncogenic driver to promote progression of chronic lymphocytic leukemia. Leukemia 34, 919–923 (2020).
Li, Y. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 25, 981–984 (2015).
Xu, C. et al. A circulating panel of circRNA biomarkers for the noninvasive and early detection of pancreatic ductal adenocarcinoma. Gastroenterology 166, 178–190.e16 (2024).
Tirpe, A. et al. The glioblastoma circularRNAome. Int. J. Mol. Sci. 24, 14545 (2023).
Xia, D. & Gu, X. Plasmatic exosome-derived circRNAs panel act as fingerprint for glioblastoma. Aging 13, 19575–19586 (2021).
Wen, G., Zhou, T. & Gu, W. The potential of using blood circular RNA as liquid biopsy biomarker for human diseases. Protein Cell 12, 911–946 (2021).
Roy, S. et al. Diagnostic efficacy of circular RNAs as noninvasive, liquid biopsy biomarkers for early detection of gastric cancer. Mol. Cancer 21, 42 (2022).
Kwong, G. A. et al. Synthetic biomarkers: a twenty-first century path to early cancer detection. Nat. Rev. Cancer 21, 655–668 (2021).
Song, H. et al. Novel exosomal circEGFR facilitates triple negative breast cancer autophagy via promoting TFEB nuclear trafficking and modulating miR-224-5p/ATG13/ULK1 feedback loop. Oncogene 43, 821–836 (2024).
Weng, W. et al. Circular RNA ciRS-7-A promising prognostic biomarker and a potential therapeutic target in colorectal cancer. Clin. Cancer Res. 23, 3918–3928 (2017).
Zhang, Y. et al. Optimized RNA-targeting CRISPR/Cas13d technology outperforms shRNA in identifying functional circRNAs. Genome Biol. 22, 41 (2021).
Shi, P. et al. Collateral activity of the CRISPR/RfxCas13d system in human cells. Commun. Biol. 6, 334 (2023).
Han, Y. et al. Synthetic RNA therapeutics in cancer. J. Pharmacol. Exp. Ther. 386, 212–223 (2023).
Amaya, L. et al. Circular RNA vaccine induces potent T cell responses. Proc. Natl Acad. Sci. USA 120, e2302191120 (2023).
Li, H. et al. Circular RNA cancer vaccines drive immunity in hard-to-treat malignancies. Theranostics 12, 6422–6436 (2022).
Qu, L. et al. Circular RNA vaccines against SARS-CoV-2 and emerging variants. Cell 185, 1728–1744.e16 (2022).
Chen, Y. G. et al. N6-methyladenosine modification controls circular RNA immunity. Mol. Cell 76, 96–109.e9 (2019).
Wesselhoeft, R. A. et al. RNA circularization diminishes immunogenicity and can extend translation duration in vivo. Mol. Cell 74, 508–520.e4 (2019).
Huang, D. et al. Tumour circular RNAs elicit anti-tumour immunity by encoding cryptic peptides. Nature 625, 593–602 (2024). This manuscript highlights the potential clinical translation of circFAM53B as an immunotherapeutic strategy against cancer.
Xu, Y. et al. Exosomal transfer of circular RNA FBXW7 ameliorates the chemoresistance to oxaliplatin in colorectal cancer by sponging miR-18b-5p. Neoplasma 68, 108–118 (2021).
Yang, Y. et al. Novel role of FBXW7 circular RNA in repressing glioma tumorigenesis. J. Natl Cancer Inst. 110, 304–315 (2018).
Ye, F. et al. circFBXW7 inhibits malignant progression by sponging miR-197-3p and encoding a 185-aa protein in triple-negative breast cancer. Mol. Ther. Nucleic Acids 18, 88–98 (2019).
Zhou, Q. et al. Exosome-mediated delivery of artificial circular RNAs for gene therapy of bladder cancer. J. Cancer 15, 1770–1778 (2024).
Han, M. et al. Regulatory mechanism and promising clinical application of exosomal circular RNA in gastric cancer. Front. Oncol. 13, 1236679 (2023).
Singh, S., Shyamal, S., Das, A. & Panda, A. C. Global identification of mRNA-interacting circular RNAs by CLiPPR-Seq. Nucleic Acids Res. 52, e29 (2024).
Dolgin, E. Why rings of RNA could be the next blockbuster drug. Nature 622, 22–24 (2023).
Erasmus, J. H. et al. An Alphavirus-derived replicon RNA vaccine induces SARS-CoV-2 neutralizing antibody and T cell responses in mice and nonhuman primates. Sci. Transl. Med. 12, eabc9396 (2020).
de Alwis, R. et al. A single dose of self-transcribing and replicating RNA-based SARS-CoV-2 vaccine produces protective adaptive immunity in mice. Mol. Ther. 29, 1970–1983 (2021).
Vogel, A. B. et al. Self-amplifying RNA vaccines give equivalent protection against influenza to mRNA vaccines but at much lower doses. Mol. Ther. 26, 446–455 (2018).
Tang, W. et al. Silencing CDR1as inhibits colorectal cancer progression through regulating microRNA-7. Onco Targets Ther. 10, 2045–2056 (2017).
Yang, X., Xiong, Q., Wu, Y., Li, S. & Ge, F. Quantitative proteomics reveals the regulatory networks of circular RNA CDR1as in hepatocellular carcinoma cells. J. Proteome Res. 16, 3891–3902 (2017).
Zhao, Z., Ji, M., Wang, Q., He, N. & Li, Y. Circular RNA Cdr1as upregulates SCAI to suppress cisplatin resistance in ovarian cancer via miR-1270 suppression. Mol. Ther. Nucleic Acids 18, 24–33 (2019).
Yuan, W. et al. Circular RNA Cdr1as sensitizes bladder cancer to cisplatin by upregulating APAF1 expression through miR-1270 inhibition. Mol. Oncol. 13, 1559–1576 (2019).
Chen, X. et al. Circular RNA circHIPK3 modulates autophagy via MIR124-3p-STAT3-PRKAA/AMPKα signaling in STK11 mutant lung cancer. Autophagy 16, 659–671 (2020).
Zeng, K. et al. CircHIPK3 promotes colorectal cancer growth and metastasis by sponging miR-7. Cell Death Dis. 9, 417 (2018).
Wang, Z. et al. Novel circular RNA circNF1 acts as a molecular sponge, promoting gastric cancer by absorbing miR-16. Endocr. Relat. Cancer 26, 265–277 (2019).
Du, W. W. et al. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur. Heart J. 38, 1402–1412 (2017).
Fang, L. et al. Enhanced breast cancer progression by mutant p53 is inhibited by the circular RNA circ-Ccnb1. Cell Death Differ. 25, 2195–2208 (2018).
Du, W. W. et al. The circular RNA circSKA3 binds integrin β1 to induce invadopodium formation enhancing breast cancer invasion. Mol. Ther. 28, 1287–1298 (2020).
Wu, W., Zhao, F. & Zhang, J. circAtlas 3.0: a gateway to 3 million curated vertebrate circular RNAs based on a standardized nomenclature scheme. Nucleic Acids Res. 52, D52–D60 (2024).
Rophina, M., Sharma, D., Poojary, M. & Scaria, V. Circad: a comprehensive manually curated resource of circular RNA associated with diseases. Database 2020, baaa019 (2020).
Fan, C. et al. CircR2Disease v2.0: an updated web server for experimentally validated circRNA–disease associations and its application. Genomics Proteom. Bioinform. 20, 435–445 (2022).
Sun, Z.-Y. et al. circRNADisease v2.0: an updated resource for high-quality experimentally supported circRNA–disease associations. Nucleic Acids Res. 52, D1193–D1200 (2024).
Liu, M., Wang, Q., Shen, J., Yang, B. B. & Ding, X. Circbank: a comprehensive database for circRNA with standard nomenclature. RNA Biol. 16, 899–905 (2019).
Dong, R., Ma, X.-K., Li, G.-W. & Yang, L. CIRCpedia v2: an updated database for comprehensive circular RNA annotation and expression comparison. Genomics Proteom. Bioinform. 16, 226–233 (2018).
Ruan, H. et al. Comprehensive characterization of circular RNAs in ~1000 human cancer cell lines. Genome Med. 11, 55 (2019).
Lai, H. et al. exoRBase 2.0: an atlas of mRNA, lncRNA and circRNA in extracellular vesicles from human biofluids. Nucleic Acids Res. 50, D118–D128 (2022).
Xia, S. et al. Comprehensive characterization of tissue-specific circular RNAs in the human and mouse genomes. Brief. Bioinform. 18, 984–992 (2017).
Wang, S. et al. TCCIA: a comprehensive resource for exploring CircRNA in cancer immunotherapy. J. Immunother. Cancer 12, e008040 (2024).
Chen, Y. et al. CircNet 2.0: an updated database for exploring circular RNA regulatory networks in cancers. Nucleic Acids Res. 50, D93–D101 (2022).
Sanger, H. L., Klotz, G., Riesner, D., Gross, H. J. & Kleinschmidt, A. K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc. Natl Acad. Sci. USA 73, 3852–3856 (1976).
Hsu, M. T. & Coca-Prados, M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature 280, 339–340 (1979).
Burd, C. E. et al. Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet. 6, e1001233 (2010).
Holdt, L. M. et al. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat. Commun. 7, 12429 (2016).
Feng, J. et al. CSCD2: an integrated interactional database of cancer-specific circular RNAs. Nucleic Acids Res. 50, D1179–D1183 (2022).
Xu, C. & Zhang, J. Mammalian circular RNAs result largely from splicing errors. Cell Rep. 36, 109439 (2021).
Santos-Rodriguez, G., Voineagu, I. & Weatheritt, R. J. Evolutionary dynamics of circular RNAs in primates. eLife 10, e69148 (2021).
Ji, P. et al. Expanded expression landscape and prioritization of circular RNAs in mammals. Cell Rep. 26, 3444–3460.e5 (2019).
Gao, Y., Zhang, J. & Zhao, F. Circular RNA identification based on multiple seed matching. Brief. Bioinform. 19, 803–810 (2018).
Song, X. et al. Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res. 44, e87 (2016).
Travis, A. J., Moody, J., Helwak, A., Tollervey, D. & Kudla, G. Hyb: a bioinformatics pipeline for the analysis of CLASH (crosslinking, ligation and sequencing of hybrids) data. Methods 65, 263–273 (2014).
Vromman, M. et al. Large-scale benchmarking of circRNA detection tools reveals large differences in sensitivity but not in precision. Nat. Methods 20, 1159–1169 (2023).
Conn, V. M. et al. Use of synthetic circular RNA spike-ins (SynCRS) for normalization of circular RNA sequencing data. Nat. Protoc. (in the press).
Xin, R. et al. isoCirc catalogs full-length circular RNA isoforms in human transcriptomes. Nat. Commun. 12, 266 (2021).
Bustin, S. A. et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622 (2009).
Zirkel, A. & Papantonis, A. Detecting circular RNAs by RNA fluorescence in situ hybridization. Methods Mol. Biol. 1724, 69–75 (2018).
Conn, V. & Conn, S. J. SplintQuant: a method for accurately quantifying circular RNA transcript abundance without reverse transcription bias. RNA 25, 1202–1210 (2019).
Hansen, E. B. et al. The transcriptional landscape and biomarker potential of circular RNAs in prostate cancer. Genome Med. 14, 8 (2022).
Dahl, M. et al. Expression patterns and prognostic potential of circular RNAs in mantle cell lymphoma: a study of younger patients from the MCL2 and MCL3 clinical trials. Leukemia 36, 177–188 (2022).
Acknowledgements
The authors gratefully acknowledge funding from the Australian National Health and Medical Research Council (GNT1198014 and research fellowship to S.J.C.) and Tour de Cure (RSP-089-2020 to S.J.C. and V.M.C.). A.M.C. was supported by the following National Cancer Institute grants U2C CA271854, P50 CA186786 and R35 CA231996.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Circad: https://clingen.igib.res.in/circad/
CircAtlas: https://ngdc.cncb.ac.cn/circatlas/
Circbank: http://www.circbank.cn/
circBase: http://www.circbase.org/
circNet2.0: https://awi.cuhk.edu.cn/~CircNet/php/index.php
CIRCpedia v2: http://yang-laboratory.com/circpedia/
CircR2Disease v2.0: http://bioinfo.snnu.edu.cn/CircR2Disease_v2.0/
CircRic: https://hanlaboratory.com/cRic/
CircRNADisease v2.0: http://cgga.org.cn:9091/circRNADisease/
CSCD2.0: http://geneyun.net/CSCD2
exoRBase 2.0: http://www.exorbase.org/
FL-circAS: https://cosbi.ee.ncku.edu.tw/FL-circAS/
MiOncoCirc: https://mioncocirc.github.io/
Tissue-specific circRNA database: http://gb.whu.edu.cn/TSCD/
Glossary
- Backsplice junction
-
(bsj). Characteristic structural feature found in circRNAs, in which a downstream 3′ splice site is joined to an upstream 5′ splice site as a chimaera.
- Endoribonucleases
-
Enzymes that cleave the phosphodiester bonds within an RNA molecule, typically at specific recognition sequences and/or structured regions.
- Epithelial-to-mesenchymal transition
-
(EMT). A change in cell phenotype associated with embryonic differentiation and wound healing that enables tumour cells to metastasize.
- miRNA sponging
-
A regulatory mechanism in which specific transcripts, often circular RNAs or long non-coding RNAs, bind to microRNAs, preventing them from targeting their usual mRNA targets.
- Polyribosomes
-
Clusters of ribosomes simultaneously translating a single mRNA or circular RNA in eukaryotic cells; also known as polysomes.
- Rapid amplification of cDNA ends
-
(RACE). A molecular biology technique used to target either the 5′ or the 3′ end of the transcript, enabling the identification of unknown sequences, including transcription start sites or polyadenylation sites, facilitating gene annotation and understanding of transcriptional regulation.
- Spliceosome
-
Multimegadalton ribonucleoprotein complex coordinating RNA splicing.
- Splicing enhancer
-
Specific regions within a gene sequence that promote the inclusion of certain exons during mRNA splicing and that contain regulatory elements recognized by splicing factors, influencing the splicing process.
- Splicing silencer
-
Regions within a gene sequence that inhibit the inclusion of specific exons during mRNA splicing and that contain regulatory elements that bind to splicing factors, interfering with the splicing process.
- Vasculogenic mimicry
-
When cancer cells form fluid-conducting channels resembling blood vessels, allowing them to acquire nutrients and oxygen independently of traditional angiogenesis.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Conn, V.M., Chinnaiyan, A.M. & Conn, S.J. Circular RNA in cancer. Nat Rev Cancer 24, 597–613 (2024). https://doi.org/10.1038/s41568-024-00721-7
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41568-024-00721-7
This article is cited by
-
A Multi-Center Cohort-Based circRNA Diagnostic Model for Detection of Gastric Cancer
Biological Procedures Online (2026)
-
CircKIAA1617 promotes stemness via USP14/PGRMC1-mediated autophagy and lipid metabolism reprogramming in ER-positive breast cancer
Molecular Cancer (2026)
-
Circular RNA hsa_circ_0001829 promotes pancreatic ductal adenocarcinoma through miR-7113–3p/DTX4 axis
World Journal of Surgical Oncology (2026)
-
CircZBTB46 alleviates metabolic dysfunction–associated steatotic liver disease by targeting miRNA-326/FGF1 axis
Cell Death Discovery (2026)
-
Dysfunctional circular RNA network in major depressive disorder: dissecting the cell identity and potential clinical applications
Molecular Psychiatry (2026)