Abstract
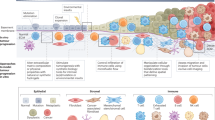
Early detection and intervention of cancer or precancerous lesions hold great promise to improve patient survival. However, the processes of cancer initiation and the normal–precancer–cancer progression within a non-cancerous tissue context remain poorly understood. This is, in part, due to the scarcity of early-stage clinical samples or suitable models to study early cancer. In this Review, we introduce clinical samples and model systems, such as autochthonous mice and organoid-derived or stem cell-derived models that allow longitudinal analysis of early cancer development. We also present the emerging techniques and computational tools that enhance our understanding of cancer initiation and early progression, including direct imaging, lineage tracing, single-cell and spatial multi-omics, and artificial intelligence models. Together, these models and techniques facilitate a more comprehensive understanding of the poorly characterized early malignant transformation cascade, holding great potential to unveil key drivers and early biomarkers for cancer development. Finally, we discuss how these new insights can potentially be translated into mechanism-based strategies for early cancer detection and prevention.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Crosby, D. et al. Early detection of cancer. Science 375, eaay9040 (2022).
Jamieson, C. H. M. & Weissman, I. L. Stem-cell aging and pathways to precancer evolution. N. Engl. J. Med. 389, 1310–1319 (2023).
Jassim, A., Rahrmann, E. P., Simons, B. D. & Gilbertson, R. J. Cancers make their own luck: theories of cancer origins. Nat. Rev. Cancer 23, 710–724 (2023).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Kakiuchi, N. & Ogawa, S. Clonal expansion in non-cancer tissues. Nat. Rev. Cancer 21, 239–256 (2021).
Li, R. et al. A body map of somatic mutagenesis in morphologically normal human tissues. Nature 597, 398–403 (2021).
Kuipers, E. J. et al. Colorectal cancer. Nat. Rev. Dis. Primers 1, 15065 (2015).
Herbst, R. S., Heymach, J. V. & Lippman, S. M. Lung cancer. N. Engl. J. Med. 359, 1367–1380 (2008).
Yuan, S., Almagro, J. & Fuchs, E. Beyond genetics: driving cancer with the tumour microenvironment behind the wheel. Nat. Rev. Cancer 24, 274–286 (2024).
Pashayan, N. & Pharoah, P. D. P. The challenge of early detection in cancer. Science 368, 589–590 (2020).
Le Magnen, C., Dutta, A. & Abate-Shen, C. Optimizing mouse models for precision cancer prevention. Nat. Rev. Cancer 16, 187–196 (2016).
Srivastava, S. et al. The making of a precancer atlas: promises, challenges, and opportunities. Trends Cancer 4, 523–536 (2018).
Gengenbacher, N., Singhal, M. & Augustin, H. G. Preclinical mouse solid tumour models: status quo, challenges and perspectives. Nat. Rev. Cancer 17, 751–765 (2017).
Chen, X. X. et al. Genomic comparison of esophageal squamous cell carcinoma and its precursor lesions by multi-region whole-exome sequencing. Nat. Commun. 8, 524 (2017).
Chitsazzadeh, V. et al. Cross-species identification of genomic drivers of squamous cell carcinoma development across preneoplastic intermediates. Nat. Commun. 7, 12601 (2016).
Wils, L. J. et al. Elucidating the genetic landscape of oral leukoplakia to predict malignant transformation. Clin. Cancer Res. 29, 602–613 (2023).
Ganz, J. et al. Rates and patterns of clonal oncogenic mutations in the normal human brain. Cancer Discov. 12, 172–185 (2022).
Moore, L. et al. The mutational landscape of normal human endometrial epithelium. Nature 580, 640–646 (2020).
Lawson, A. R. J. et al. Extensive heterogeneity in somatic mutation and selection in the human bladder. Science 370, 75–82 (2020).
Yokoyama, A. et al. Age-related remodelling of oesophageal epithelia by mutated cancer drivers. Nature 565, 312–317 (2019).
Lee-Six, H. et al. The landscape of somatic mutation in normal colorectal epithelial cells. Nature 574, 532–537 (2019).
Brunner, S. F. et al. Somatic mutations and clonal dynamics in healthy and cirrhotic human liver. Nature 574, 538–542 (2019).
Suda, K. et al. Clonal expansion and diversification of cancer-associated mutations in endometriosis and normal endometrium. Cell Rep. 24, 1777–1789 (2018).
Martincorena, I. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Martincorena, I. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015).
Li, R. et al. Macroscopic somatic clonal expansion in morphologically normal human urothelium. Science 370, 82–89 (2020).
Robles, A. I., Jen, J. & Harris, C. C. Clinical outcomes of TP53 mutations in cancers. Cold Spring Harb. Perspect. Med. 6, a026294 (2016).
Ren, X. et al. Single-cell transcriptomic analysis highlights origin and pathological process of human endometrioid endometrial carcinoma. Nat. Commun. 13, 6300 (2022).
Liao, G. et al. Single-cell transcriptomics provides insights into the origin and microenvironment of human oesophageal high-grade intraepithelial neoplasia. Clin. Transl. Med. 12, e874 (2022).
Owen, R. P. et al. Single cell RNA-seq reveals profound transcriptional similarity between Barrett’s oesophagus and oesophageal submucosal glands. Nat. Commun. 9, 4261 (2018).
Joyce, R. et al. Identification of aberrant luminal progenitors and mTORC1 as a potential breast cancer prevention target in BRCA2 mutation carriers. Nat. Cell Biol. 26, 138–152 (2024).
Nowicki-Osuch, K. et al. Molecular phenotyping reveals the identity of Barrett’s esophagus and its malignant transition. Science 373, 760–767 (2021).
Chen, B. et al. Differential pre-malignant programs and microenvironment chart distinct paths to malignancy in human colorectal polyps. Cell 184, 6262–6280.e26 (2021).
Liu, R. et al. Co-evolution of tumor and immune cells during progression of multiple myeloma. Nat. Commun. 12, 2559 (2021).
Becker, W. R. et al. Single-cell analyses define a continuum of cell state and composition changes in the malignant transformation of polyps to colorectal cancer. Nat. Genet. 54, 985–995 (2022).
Liu, Z. et al. Single-cell chromatin accessibility analysis reveals the epigenetic basis and signature transcription factors for the molecular subtypes of colorectal cancers. Cancer Discov. 14, 1082–1105 (2024).
Zhang, P. et al. Dissecting the single-cell transcriptome network underlying gastric premalignant lesions and early gastric cancer. Cell Rep. 27, 1934–1947.e5 (2019).
Huang, K. K. et al. Spatiotemporal genomic profiling of intestinal metaplasia reveals clonal dynamics of gastric cancer progression. Cancer Cell 41, 2019–2037.e8 (2023).
Wang, Z. et al. Deciphering cell lineage specification of human lung adenocarcinoma with single-cell RNA sequencing. Nat. Commun. 12, 6500 (2021).
Zhang, T. et al. Identification of cervical cancer stem cells using single-cell transcriptomes of normal cervix, cervical premalignant lesions, and cervical cancer. EBioMedicine 92, 104612 (2023).
Zou, D. D. et al. Single-cell sequencing highlights heterogeneity and malignant progression in actinic keratosis and cutaneous squamous cell carcinoma. eLife 12, e85270 (2023).
Choi, J. H. et al. Single-cell transcriptome profiling of the stepwise progression of head and neck cancer. Nat. Commun. 14, 1055 (2023).
Liu, X. et al. Spatial transcriptomics analysis of esophageal squamous precancerous lesions and their progression to esophageal cancer. Nat. Commun. 14, 4779 (2023).
Carpenter, E. S. et al. Analysis of donor pancreata defines the transcriptomic signature and microenvironment of early neoplastic lesions. Cancer Discov. 13, 1324–1345 (2023).
Li, J. et al. Genomic and transcriptomic profiling of carcinogenesis in patients with familial adenomatous polyposis. Gut 69, 1283–1293 (2020).
Chang, J. et al. Genomic alterations driving precancerous to cancerous lesions in esophageal cancer development. Cancer Cell 41, 2038–2050.e5 (2023). This paper delineates the inactivation of TP53 as a preliminary step in early carcinogenesis of esophageal SCC.
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Erickson, A. et al. Spatially resolved clonal copy number alterations in benign and malignant tissue. Nature 608, 360–367 (2022).
Cui Zhou, D. et al. Spatially restricted drivers and transitional cell populations cooperate with the microenvironment in untreated and chemo-resistant pancreatic cancer. Nat. Genet. 54, 1390–1405 (2022).
Heiser, C. N. et al. Molecular cartography uncovers evolutionary and microenvironmental dynamics in sporadic colorectal tumors. Cell 186, 5620–5637.e16 (2023).
Luebeck, J. et al. Extrachromosomal DNA in the cancerous transformation of Barrett’s oesophagus. Nature 616, 798–805 (2023). This paper reveals that patients with Barrett’s esophagus who advanced to esophageal adenocarcinoma displayed significantly increased levels of ecDNA, housing a wide array of oncogenes and immunomodulatory genes.
Zavidij, O. et al. Single-cell RNA sequencing reveals compromised immune microenvironment in precursor stages of multiple myeloma. Nat. Cancer 1, 493–506 (2020).
Liu, C. et al. Single-cell dissection of cellular and molecular features underlying human cervical squamous cell carcinoma initiation and progression. Sci. Adv. 9, eadd8977 (2023).
Liu, W. et al. An immune cell map of human lung adenocarcinoma development reveals an anti-tumoral role of the Tfh-dependent tertiary lymphoid structure. Cell Rep. Med. 5, 101448 (2024).
Yanagawa, J. et al. Single-cell characterization of pulmonary nodules implicates suppression of immunosurveillance across early stages of lung adenocarcinoma. Cancer Res. 83, 3305–3319 (2023).
Hu, S. et al. TDO2+ myofibroblasts mediate immune suppression in malignant transformation of squamous cell carcinoma. J. Clin. Invest. 132, e157649 (2022).
Chen, Y. et al. Epithelial cells activate fibroblasts to promote esophageal cancer development. Cancer Cell 41, 903–918.e8 (2023).
Nowicki-Osuch, K. et al. Single-cell RNA sequencing unifies developmental programs of esophageal and gastric intestinal metaplasia. Cancer Discov. 13, 1346–1363 (2023).
Lee, S.-H. et al. Apposition of fibroblasts with metaplastic gastric cells promotes dysplastic transition. Gastroenterology 165, 374–390 (2023).
Wang, R. et al. Evolution of immune and stromal cell states and ecotypes during gastric adenocarcinoma progression. Cancer Cell 41, 1407–1426.e9 (2023). This paper describes a landscape of the tumour microenvironment at various stages of gastric adenocarcinoma, identifying the crucial tumour microenvironment ecotypes associated with the phenotypic progression and results of gastric adenocarcinoma.
Schutz, S. et al. Functionally distinct cancer-associated fibroblast subpopulations establish a tumor promoting environment in squamous cell carcinoma. Nat. Commun. 14, 5413 (2023).
Sun, L. et al. Single-cell and spatial dissection of precancerous lesions underlying the initiation process of oral squamous cell carcinoma. Cell Discov. 9, 28 (2023).
Roelands, J. et al. Transcriptomic and immunophenotypic profiling reveals molecular and immunological hallmarks of colorectal cancer tumourigenesis. Gut 72, 1326–1339 (2022).
Wang, G. et al. Lung cancer scRNA-seq and lipidomics reveal aberrant lipid metabolism for early-stage diagnosis. Sci. Transl. Med. 14, eabk2756 (2022).
Nie, M. et al. Evolutionary metabolic landscape from preneoplasia to invasive lung adenocarcinoma. Nat. Commun. 12, 6479 (2021).
Chiu, C. Y. & Miller, S. A. Clinical metagenomics. Nat. Rev. Genet. 20, 341–355 (2019).
Serrano-Villar, S. et al. Microbiome-derived cobalamin and succinyl-CoA as biomarkers for improved screening of anal cancer. Nat. Med. 29, 1738–1749 (2023).
Leshchiner, I. et al. Inferring early genetic progression in cancers with unobtainable premalignant disease. Nat. Cancer 4, 550–563 (2023). This paper reports a computational method named PhylogicNDT, which can predict the early genetic events in cancers that lack precancerous lesions.
Cao, K. et al. Large-scale pancreatic cancer detection via non-contrast CT and deep learning. Nat. Med. 29, 3033–3043 (2023). This paper reports a deep learning model called PANDA, which effectively detects and classifies early-stage malignancies with high accuracy using non-contrast CT scans.
Thirunavukarasu, A. J. et al. Large language models in medicine. Nat. Med. 29, 1930–1940 (2023).
Lipkova, J. et al. Artificial intelligence for multimodal data integration in oncology. Cancer Cell 40, 1095–1110 (2022).
Condeelis, J. & Weissleder, R. In vivo imaging in cancer. Cold Spring Harb. Perspect. Biol. 2, a003848 (2010).
Yuan, X. L. et al. Effect of an artificial intelligence-assisted system on endoscopic diagnosis of superficial oesophageal squamous cell carcinoma and precancerous lesions: a multicentre, tandem, double-blind, randomised controlled trial. Lancet Gastroenterol. Hepatol. 9, 34–44 (2024).
Bao, H. et al. Artificial intelligence-assisted cytology for detection of cervical intraepithelial neoplasia or invasive cancer: a multicenter, clinical-based, observational study. Gynecol. Oncol. 159, 171–178 (2020).
Dong, Y. et al. A polarization-imaging-based machine learning framework for quantitative pathological diagnosis of cervical precancerous lesions. IEEE Trans. Med. Imaging 40, 3728–3738 (2021).
Fockens, K. N. et al. A deep learning system for detection of early Barrett’s neoplasia: a model development and validation study. Lancet Digit. Health 5, e905–e916 (2023).
Yin, J. et al. Differential diagnosis of DCIS and fibroadenoma based on ultrasound images: a difference-based self-supervised approach. Interdiscip. Sci. 15, 262–272 (2023).
Bhowmik, A. et al. Portable, handheld, and affordable blood perfusion imager for screening of subsurface cancer in resource-limited settings. Proc. Natl Acad. Sci. USA 119, e2026201119 (2022).
Placido, D. et al. A deep learning algorithm to predict risk of pancreatic cancer from disease trajectories. Nat. Med. 29, 1113–1122 (2023).
Ardila, D. et al. End-to-end lung cancer screening with three-dimensional deep learning on low-dose chest computed tomography. Nat. Med. 25, 954–961 (2019).
Salim, M. et al. AI-based selection of individuals for supplemental MRI in population-based breast cancer screening: the randomized ScreenTrustMRI trial. Nat. Med. 30, 2623–2630 (2024).
Chen, Y. et al. Metabolomic machine learning predictor for diagnosis and prognosis of gastric cancer. Nat. Commun. 15, 1657 (2024).
Deng, Z. et al. Early detection of hepatocellular carcinoma via no end-repair enzymatic methylation sequencing of cell-free DNA and pre-trained neural network. Genome Med. 15, 93 (2023).
Korber, V. et al. Evolutionary trajectories of IDH(WT) glioblastomas reveal a common path of early tumorigenesis instigated years ahead of initial diagnosis. Cancer Cell 35, 692–704.e12 (2019).
Wu, L. et al. Natural coevolution of tumor and immunoenvironment in glioblastoma. Cancer Discov. 12, 2820–2837 (2022).
Stangis, M. M. et al. The hallmarks of precancer. Cancer Discov. 14, 683–689 (2024).
Greener, J. G., Kandathil, S. M., Moffat, L. & Jones, D. T. A guide to machine learning for biologists. Nat. Rev. Mol. Cell Biol. 23, 40–55 (2022).
Kersten, K., de Visser, K. E., van Miltenburg, M. H. & Jonkers, J. Genetically engineered mouse models in oncology research and cancer medicine. EMBO Mol. Med. 9, 137–153 (2017).
Hebert, J. D., Neal, J. W. & Winslow, M. M. Dissecting metastasis using preclinical models and methods. Nat. Rev. Cancer 23, 391–407 (2023).
Sanchez-Rivera, F. J. et al. Rapid modelling of cooperating genetic events in cancer through somatic genome editing. Nature 516, 428–431 (2014).
Zhao, H. & Zhou, B. Dual genetic approaches for deciphering cell fate plasticity in vivo: more than double. Curr. Opin. Cell Biol. 61, 101–109 (2019).
Schonhuber, N. et al. A next-generation dual-recombinase system for time- and host-specific targeting of pancreatic cancer. Nat. Med. 20, 1340–1347 (2014).
Robles-Oteiza, C. et al. Recombinase-based conditional and reversible gene regulation via XTR alleles. Nat. Commun. 6, 8783 (2015).
Hingorani, S. R. et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell 4, 437–450 (2003).
Hingorani, S. R. et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 7, 469–483 (2005).
Aguirre, A. J. et al. Activated Kras and Ink4a/Arf deficiency cooperate to produce metastatic pancreatic ductal adenocarcinoma. Genes Dev. 17, 3112–3126 (2003).
Kemp, C. J. Animal models of chemical carcinogenesis: driving breakthroughs in cancer research for 100 years. Cold Spring Harb. Protoc. 2015, 865–874 (2015).
Hill, W. et al. Lung adenocarcinoma promotion by air pollutants. Nature 616, 159–167 (2023). This paper reports that PM2.5 contributes to lung cancer by impacting cells that already have oncogenic events in healthy lung tissue.
Balmain, A. The critical roles of somatic mutations and environmental tumor-promoting agents in cancer risk. Nat. Genet. 52, 1139–1143 (2020).
Colom, B. et al. Spatial competition shapes the dynamic mutational landscape of normal esophageal epithelium. Nat. Genet. 52, 604–614 (2020).
Murai, K. p53 mutation in normal esophagus promotes multiple stages of carcinogenesis but is constrained by clonal competition. Nat. Commun. 13, 6206 (2022).
Scheele, C. et al. Multiphoton intravital microscopy of rodents. Nat. Rev. Methods Primers 2, 89 (2022).
Entenberg, D., Oktay, M. H. & Condeelis, J. S. Intravital imaging to study cancer progression and metastasis. Nat. Rev. Cancer 23, 25–42 (2023).
Xin, T. et al. Oncogenic Kras induces spatiotemporally specific tissue deformation through converting pulsatile into sustained ERK activation. Nat. Cell Biol. 26, 859–867 (2024). This paper reports that the KrasG12D mutation induces epithelial tissue deformation in a spatiotemporally specific manner, primarily through the continuous activation of ERK signals.
Almagro, J., Messal, H. A., Zaw Thin, M., van Rheenen, J. & Behrens, A. Tissue clearing to examine tumour complexity in three dimensions. Nat. Rev. Cancer 21, 718–730 (2021).
Messal, H. A. et al. Tissue curvature and apicobasal mechanical tension imbalance instruct cancer morphogenesis. Nature 566, 126–130 (2019).
Chen, P. et al. Olfactory sensory experience regulates gliomagenesis via neuronal IGF1. Nature 606, 550–556 (2022).
Iwano, S. et al. Single-cell bioluminescence imaging of deep tissue in freely moving animals. Science 359, 935–939 (2018).
Su, Y. et al. An optimized bioluminescent substrate for non-invasive imaging in the brain. Nat. Chem. Biol. 19, 731–739 (2023).
Hosein, A. N. et al. Cellular heterogeneity during mouse pancreatic ductal adenocarcinoma progression at single-cell resolution. JCI Insight 5, e129212 (2019).
Yao, J. et al. Single-cell transcriptomic analysis in a mouse model deciphers cell transition states in the multistep development of esophageal cancer. Nat. Commun. 11, 3715 (2020).
Li, D. et al. ETV4 mediates dosage-dependent prostate tumor initiation and cooperates with p53 loss to generate prostate cancer. Sci. Adv. 9, eadc9446 (2023).
Yuan, S. et al. Ras drives malignancy through stem cell crosstalk with the microenvironment. Nature 612, 555–563 (2022).
Abu El Maaty, M. A. et al. Single-cell analyses unravel cell type-specific responses to a vitamin D analog in prostatic precancerous lesions. Sci. Adv. 7, eabg5982 (2021).
Abu El Maaty, M. A. et al. Hypoxia-mediated stabilization of HIF1A in prostatic intraepithelial neoplasia promotes cell plasticity and malignant progression. Sci. Adv. 8, eabo2295 (2022).
Prieto, L. I. et al. Senescent alveolar macrophages promote early-stage lung tumorigenesis. Cancer Cell 41, 1261–1275.e6 (2023).
Haston, S. et al. Clearance of senescent macrophages ameliorates tumorigenesis in KRAS-driven lung cancer. Cancer Cell 41, 1242–1260.e6 (2023).
Kolodkin-Gal, D. et al. Senolytic elimination of Cox2-expressing senescent cells inhibits the growth of premalignant pancreatic lesions. Gut 71, 345–355 (2022).
Yeo, A. T. et al. Single-cell RNA sequencing reveals evolution of immune landscape during glioblastoma progression. Nat. Immunol. 23, 971–984 (2022).
Rajendran, S. et al. Single-cell RNA sequencing reveals immunosuppressive myeloid cell diversity during malignant progression in a murine model of glioma. Cell Rep. 42, 112197 (2023).
Wagner, D. E. & Klein, A. M. Lineage tracing meets single-cell omics: opportunities and challenges. Nat. Rev. Genet. 21, 410–427 (2020).
Visvader, J. E. Cells of origin in cancer. Nature 469, 314–322 (2011).
Liu, C. et al. Mosaic analysis with double markers reveals tumor cell of origin in glioma. Cell 146, 209–221 (2011).
Sket, T., Falcomata, C. & Saur, D. Dual recombinase-based mouse models help decipher cancer biology and targets for therapy. Cancer Res. 83, 2279–2282 (2023).
Boone, P. G. et al. A cancer rainbow mouse for visualizing the functional genomics of oncogenic clonal expansion. Nat. Commun. 10, 5490 (2019).
Sankaran, V. G., Weissman, J. S. & Zon, L. I. Cellular barcoding to decipher clonal dynamics in disease. Science 378, eabm5874 (2022).
Kalhor, R. et al. Developmental barcoding of whole mouse via homing CRISPR. Science 361, eaat9804 (2018).
Bowling, S. et al. An engineered CRISPR-Cas9 mouse line for simultaneous readout of lineage histories and gene expression profiles in single cells. Cell 181, 1410–1422.e27 (2020).
Ratz, M. et al. Clonal relations in the mouse brain revealed by single-cell and spatial transcriptomics. Nat. Neurosci. 25, 285–294 (2022).
Baslan, T. et al. Ordered and deterministic cancer genome evolution after p53 loss. Nature 608, 795–802 (2022). This paper reports a PDAC mouse model, which allows tracing precancerous cells by monitoring the loss of heterozygosity of the second wild-type Trp53 allele, and uncovers a sequential pattern of genome evolution throughout tumorigenesis.
Yao, P. et al. Protein-level mutant p53 reporters identify druggable rare precancerous clones in noncancerous tissues. Nat. Cancer 4, 1176–1192 (2023). This paper reports a protein-level mutant p53 reporter that effectively replicates the functionality of mutant p53 proteins in vivo, facilitating the detection and monitoring of rare precancerous clones in deep non-cancerous tissues.
Johnsson, A. E. et al. The Rac-FRET mouse reveals tight spatiotemporal control of Rac activity in primary cells and tissues. Cell Rep. 6, 1153–1164 (2014).
Burd, C. E. et al. Monitoring tumorigenesis and senescence in vivo with a p16(INK4a)-luciferase model. Cell 152, 340–351 (2013).
Yum, M. K. et al. Tracing oncogene-driven remodelling of the intestinal stem cell niche. Nature 594, 442–447 (2021).
Yang, D. et al. Lineage tracing reveals the phylodynamics, plasticity, and paths of tumor evolution. Cell 185, 1905–1923.e25 (2022). This paper reports a lung mouse model that enables continuous cell lineage tracing and identifies rare clonal expansion during tumour development.
Ceresa, D. et al. Early clonal extinction in glioblastoma progression revealed by genetic barcoding. Cancer Cell 41, 1466–1479.e9 (2023).
Colom, B. et al. Mutant clones in normal epithelium outcompete and eliminate emerging tumours. Nature 598, 510–514 (2021).
Jiang, Z. et al. Tff2 defines transit-amplifying pancreatic acinar progenitors that lack regenerative potential and are protective against Kras-driven carcinogenesis. Cell Stem Cell 30, 1091–1109.e7 (2023).
Chen, Y. et al. Club cells employ regeneration mechanisms during lung tumorigenesis. Nat. Commun. 13, 4557 (2022).
Taylor, M. A. et al. Stem-cell states converge in multistage cutaneous squamous cell carcinoma development. Science 384, eadi7453 (2024).
Marjanovic, N. D. et al. Emergence of a high-plasticity cell state during lung cancer evolution. Cancer Cell 38, 229–246.e13 (2020).
Rajbhandari, N. et al. Single-cell mapping identifies MSI+ cells as a common origin for diverse subtypes of pancreatic cancer. Cancer Cell 41, 1989–2005.e9 (2023).
Schlesinger, Y. et al. Single-cell transcriptomes of pancreatic preinvasive lesions and cancer reveal acinar metaplastic cells’ heterogeneity. Nat. Commun. 11, 4516 (2020).
Burdziak, C. et al. Epigenetic plasticity cooperates with cell-cell interactions to direct pancreatic tumorigenesis. Science 380, eadd5327 (2023).
Li, Y. et al. Mutant Kras co-opts a proto-oncogenic enhancer network in inflammation-induced metaplastic progenitor cells to initiate pancreatic cancer. Nat. Cancer 2, 49–65 (2021).
Alonso-Curbelo, D. et al. A gene-environment-induced epigenetic program initiates tumorigenesis. Nature 590, 642–648 (2021).
Del Poggetto, E. et al. Epithelial memory of inflammation limits tissue damage while promoting pancreatic tumorigenesis. Science 373, eabj0486 (2021).
Brubaker, D. K. & Lauffenburger, D. A. Translating preclinical models to humans. Science 367, 742–743 (2020).
Dudgeon, C. et al. The evolution of thymic lymphomas in p53 knockout mice. Genes Dev. 28, 2613–2620 (2014).
McFadden, D. G. et al. Genetic and clonal dissection of murine small cell lung carcinoma progression by genome sequencing. Cell 156, 1298–1311 (2014).
Zhao, Z. et al. Organoids. Nat. Rev. Methods Primers 2, 94 (2022).
Tuveson, D. & Clevers, H. Cancer modeling meets human organoid technology. Science 364, 952–955 (2019).
Koster, S. et al. Modelling chlamydia and HPV co-infection in patient-derived ectocervix organoids reveals distinct cellular reprogramming. Nat. Commun. 13, 1030 (2022).
Hu, B. et al. A promising new model: establishment of patient-derived organoid models covering HPV-related cervical pre-cancerous lesions and their cancers. Adv. Sci. 11, e2302340 (2024).
Karlsson, K. et al. Deterministic evolution and stringent selection during preneoplasia. Nature 618, 383–393 (2023). This paper reveals that loss of TP53 results in progressive aneuploidy that follows a specific temporal order.
Yuan, L. et al. Reconstruction of dynamic mammary mini gland in vitro for normal physiology and oncogenesis. Nat. Methods 20, 2021–2033 (2023). This paper presents an organoid system designed for in vitro investigation of tumour initiation and evaluation of prospective cancer therapies.
Bian, S. et al. Genetically engineered cerebral organoids model brain tumor formation. Nat. Methods 15, 631–639 (2018).
Seino, T. et al. Human pancreatic tumor organoids reveal loss of stem cell niche factor dependence during disease progression. Cell Stem Cell 22, 454–467.e6 (2018).
Boretto, M. et al. Patient-derived organoids from endometrial disease capture clinical heterogeneity and are amenable to drug screening. Nat. Cell Biol. 21, 1041–1051 (2019).
Goto, N. et al. SOX17 enables immune evasion of early colorectal adenomas and cancers. Nature 627, 636–645 (2024).
Breunig, M. et al. Modeling plasticity and dysplasia of pancreatic ductal organoids derived from human pluripotent stem cells. Cell Stem Cell 28, 1105–1124.e19 (2021).
Matano, M. et al. Modeling colorectal cancer using CRISPR-Cas9-mediated engineering of human intestinal organoids. Nat. Med. 21, 256–262 (2015).
O’Rourke, K. P. et al. Transplantation of engineered organoids enables rapid generation of metastatic mouse models of colorectal cancer. Nat. Biotechnol. 35, 577–582 (2017).
Na, F. et al. KMT2C deficiency promotes small cell lung cancer metastasis through DNMT3A-mediated epigenetic reprogramming. Nat. Cancer 3, 753–767 (2022).
Sun, L. et al. Modelling liver cancer initiation with organoids derived from directly reprogrammed human hepatocytes. Nat. Cell Biol. 21, 1015–1026 (2019).
Huang, L. et al. Commitment and oncogene-induced plasticity of human stem cell-derived pancreatic acinar and ductal organoids. Cell Stem Cell 28, 1090–1104.e6 (2021).
Min, J. et al. Heterogeneity and dynamics of active Kras-induced dysplastic lineages from mouse corpus stomach. Nat. Commun. 10, 5549 (2019).
Xu, Y. et al. Reconstitution of human PDAC using primary cells reveals oncogenic transcriptomic features at tumor onset. Nat. Commun. 15, 818 (2024).
Yucer, N. et al. Human iPSC-derived fallopian tube organoids with BRCA1 mutation recapitulate early-stage carcinogenesis. Cell Rep. 37, 110146 (2021).
Clevers, H. Modeling development and disease with organoids. Cell 165, 1586–1597 (2016).
Guo, W. et al. Single-cell transcriptomics identifies a distinct luminal progenitor cell type in distal prostate invagination tips. Nat. Genet. 52, 908–918 (2020).
Lorenzo-Martin, L. F. et al. Spatiotemporally resolved colorectal oncogenesis in mini-colons ex vivo. Nature 629, 450–457 (2024). This paper reports mini-colons, a 3D organoid culture system that enables spatial and temporal control of tumorigenesis through blue light exposure, allowing real-time tracking of emerging colon tumours at single-cell resolution.
Wu, B. et al. Single-cell transcriptome analyses reveal critical roles of RNA splicing during leukemia progression. PLoS Biol. 21, e3002088 (2023).
Wang, X. et al. Sequential fate-switches in stem-like cells drive the tumorigenic trajectory from human neural stem cells to malignant glioma. Cell Res. 31, 684–702 (2021). This paper reports the oncogenic event-induced gliomagenic trajectories in human neural stem cells and discovers a sustained neural stem cell-like group that drives tumour progression throughout all stages of tumorigenesis.
Haag, D. et al. H3.3-K27M drives neural stem cell-specific gliomagenesis in a human iPSC-derived model. Cancer Cell 39, 407–422.e13 (2021).
Funato, K., Major, T., Lewis, P. W., Allis, C. D. & Tabar, V. Use of human embryonic stem cells to model pediatric gliomas with H3.3K27M histone mutation. Science 346, 1529–1533 (2014).
Flanagan, D. J. et al. NOTUM from Apc-mutant cells biases clonal competition to initiate cancer. Nature 594, 430–435 (2021).
van Neerven, S. M. et al. Apc-mutant cells act as supercompetitors in intestinal tumour initiation. Nature 594, 436–441 (2021).
Dost, A. F. M. et al. Organoids model transcriptional hallmarks of oncogenic KRAS activation in lung epithelial progenitor cells. Cell Stem Cell 27, 663–678.e8 (2020).
Min, J. et al. Dysplastic stem cell plasticity functions as a driving force for neoplastic transformation of precancerous gastric mucosa. Gastroenterology 163, 875–890 (2022).
Tao, Y. et al. Aging-like spontaneous epigenetic silencing facilitates Wnt activation, stemness, and BrafV600E-induced tumorigenesis. Cancer Cell 35, 315–328.e6 (2019).
Hofer, M. & Lutolf, M. P. Engineering organoids. Nat. Rev. Mater. 6, 402–420 (2021).
Hicks, M. R. & Pyle, A. D. The emergence of the stem cell niche. Trends Cell Biol. 33, 112–123 (2023).
Kirschenbaum, D. et al. Time-resolved single-cell transcriptomics defines immune trajectories in glioblastoma. Cell 187, 149–165.e23 (2024).
Dang, M. et al. Single cell clonotypic and transcriptional evolution of multiple myeloma precursor disease. Cancer Cell 41, 1032–1047.e4 (2023).
Chen, L. et al. Aberrant epithelial cell interaction promotes esophageal squamous-cell carcinoma development and progression. Signal Transduct. Target. Ther. 8, 453 (2023).
Roulis, M. et al. Paracrine orchestration of intestinal tumorigenesis by a mesenchymal niche. Nature 580, 524–529 (2020).
Sethi, N. S. et al. Early TP53 alterations engage environmental exposures to promote gastric premalignancy in an integrative mouse model. Nat. Genet. 52, 219–230 (2020).
Zhao, H. et al. Generation and multiomic profiling of a TP53/CDKN2A double-knockout gastroesophageal junction organoid model. Sci. Transl. Med. 14, eabq6146 (2022).
Acknowledgements
Y.W. gratefully acknowledges funding from the National Natural Science Foundation of China (82425037, 92359303, 82273117), the National Key R&D Program of China, Stem Cell and Translational Research (2022YFA1105200), Sichuan Science and Technology Program (2023ZYD0128, 2024NSFSC0059) and West China Hospital (ZYYC23023). R.Z. thanks the National Natural Science Foundation of China (82303975), the China Postdoctoral Science Foundation (2022TQ0226 and 2023M742492) and West China Hospital (2023HXBH100). After completing the final manuscript, the authors utilized ChatGPT (OpenAI, https://chat.openai.com/) to proofread the final draft.
Author information
Authors and Affiliations
Contributions
R.Z. and X.T. researched data for the article. R.Z. and Y.W. wrote the article. All authors reviewed or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Cancer thanks Toshiro Sato, Rebecca Fitzgerald and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
ATCC: https://www.atcc.org
inferCNV: https://github.com/broadinstitute/infercnv
PreCancer Atlas: https://prevention.cancer.gov/major-programs/pre-cancer-atlas-pca
Glossary
- Acinar-to-ductal metaplasia
-
The transformation of pancreatic acinar cells into duct-like cells in response to pancreatic injury or chronic stress that is considered a precursor to pancreatic intraepithelial neoplasia and pancreatic ductal adenocarcinoma.
- Actinic keratosis
-
A rough, dry and scaly patch or plaque on the skin, considered a precursor to squamous cell carcinoma of the skin, that is associated with risk factors such as sun exposure, human papillomavirus, fair skin, immunosuppressive therapy and age.
- Adenomas
-
Benign tumours originating from glandular epithelial tissue in various organs, including the colon, pituitary gland, thyroid, adrenal glands and liver.
- Assay for transposase-accessible chromatin with sequencing
-
(ATAC-seq). A widely utilized method to assess the accessibility of chromatin within cells.
- Assembloids
-
3D in vitro tissue models that integrates multiple organoid types or cell lineages to replicate the complex interactions and architecture of tissues.
- Atypical adenomatous hyperplasia
-
A term predominantly used in lung pathology to describe precursor lesions of lung adenocarcinoma; however, it can occasionally be applied to other organs to denote abnormal, precancerous growth patterns in glandular tissues.
- Atypical endometrial hyperplasia
-
Also known as endometrial intraepithelial neoplasia. A precancerous condition characterized by the abnormal proliferation of the cells lining the endometrium with atypical cellular features.
- Autochthonous mouse models
-
Mouse models in which cancer arises naturally within the mouse, typically induced through genetic modifications or exposure to carcinogens.
- Barrett’s oesophagus
-
Barrett’s oesophagus precedes the onset of oesophageal adenocarcinoma and involves a metaplastic change in the mucosal cells within the lower part of the oesophagus, a response to damage from gastro-oesophageal reflux.
- Chronic atrophic gastritis
-
A long-term condition characterized by the chronic inflammation of the stomach lining, leading to the gradual loss of gastric glandular cells, resulting in stomach lining atrophy that can be accompanied by changes in the structure of the stomach lining, potentially leading to intestinal metaplasia and an increased risk of gastric cancer.
- Club cells
-
A type of non-ciliated epithelial cells located in the bronchioles of the lungs, which is essential for maintaining the health and function of the respiratory epithelium.
- Dysplasia
-
An abnormal growth or development of cells within a tissue, exhibiting irregularities in size, shape, organization and cellular structure, with severity ranging from mild to severe, with high-grade dysplasia carrying a risk of progressing to cancer.
- Extrachromosomal DNA
-
(ecDNA). Any DNA that is often larger than 1 Mb and is located outside the chromosomes.
- Familial adenomatous polyposis
-
A type of syndromic polyp associated with genetic syndromes that predisposes individuals to the development of hundreds to thousands of adenomatous polyps in the colon and rectum, which almost inevitably progresses to colorectal cancer.
- Field cancerization
-
A process through which a broad region of cells within a tissue or organ undergoes genetic and epigenetic alterations, thereby predisposing the entire field to an elevated risk of developing cancer.
- Hyperplasia
-
Hyperplasia is a reversible process that involves an increase in the number of cells in a tissue, which can occur owing to physiological or pathological triggers.
- Intravital microscopy
-
(IVM). A technique to visualize cells within a living organism using fluorescent markers or dyes that encompasses various methods, including confocal microscopy, two-photon microscopy and multiphoton microscopy.
- Large language models
-
Advanced machine learning algorithms designed to grasp the intricacies, patterns and subtleties of human language by analysing data sets containing billions of words.
- Leukoplakia
-
White patches or plaques in the oral or genital regions with the potential to progress to squamous cell carcinoma, which is associated with risk factors including tobacco use, alcohol consumption, chronic irritation, viral infections such as HPV, and age and gender.
- Low-grade intraepithelial neoplasia
-
(LGIN). A precancerous condition characterized by the presence of mildly abnormal epithelial cells confined to the epithelial layer of the oesophagus that has a lower risk of progression to oesophageal cancer than high-grade intraepithelial neoplasia (HGIN).
- Metaplasia
-
A reversible process through which one differentiated cell is replaced by another cell type, typically in response to persistent irritation or inflammation.
- Monoclonal gammopathy of undetermined significance
-
A precancerous condition with a modest risk of progression to more severe plasma cell disorders that is characterized by the presence of an abnormal monoclonal protein in the blood, produced by a clone of plasma cells.
- Neoplastic polyps
-
Polyps with the potential to develop into colorectal cancer that can be classified into adenomatous polyps (including tubular, tubulovillous and villous types) and serrated polyps (including sessile serrated lesions and traditional serrated adenomas).
- Pancreatic intraepithelial neoplasia
-
(PanIN). These precursor lesions to pancreatic ductal adenocarcinoma that encompass a spectrum of dysplastic changes are microscopic abnormalities found in the epithelial cells lining the pancreatic ducts and are not detectable through standard imaging techniques.
- Precancerous cells
-
Cancer precursor cells that harbour common cancer driver mutations but exist in tissues without clinical evidence of precursor lesions.
- Precancerous lesions
-
Morphological changes in tissues resulting from the acquisition of driver events in individual cells that undergo positive selection and clonal expansion within normal tissues, leading to tissue remodelling.
- Pulmonary nodules
-
Small, abnormal areas that appear in the lung tissue that can result from infections or scarring, which could potentially lead to lung cancer.
- Single-cell multi-omics
-
An advanced genomic technique that enables the simultaneous analysis of multiple types of biological signals such as the genome, transcriptome, proteome and metabolome within the same cell.
- Smouldering multiple myeloma
-
An asymptomatic, precancerous condition that is considered an intermediate stage between monoclonal gammopathy of undetermined significance and symptomatic multiple myeloma and is characterized by elevated levels of monoclonal protein and a higher percentage of abnormal plasma cells in the bone marrow.
- Spatial transcriptomics
-
A method that enables the spatial visualization and quantification of gene expression within a tissue.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, R., Tang, X. & Wang, Y. Emerging strategies to investigate the biology of early cancer. Nat Rev Cancer 24, 850–866 (2024). https://doi.org/10.1038/s41568-024-00754-y
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41568-024-00754-y
This article is cited by
-
Signaling pathways and targeted interventions for precancers
Signal Transduction and Targeted Therapy (2026)
-
Plasma untargeted metabolomics reveals the metabolic landscape of NKTCL and aids in liquid biopsy for disease detection
European Journal of Medical Research (2025)
-
Mitophagy’s impacts on cancer and neurodegenerative diseases: implications for future therapies
Journal of Hematology & Oncology (2025)
-
PCMR: a comprehensive precancerous molecular resource
Scientific Data (2025)
-
Engineering and biofabrication of early cancer models
Nature Reviews Bioengineering (2025)