Abstract
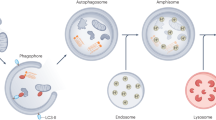
Autophagy is a powerful tool that host cells use to defend against viral infection. Double-membrane vesicles, termed autophagosomes, deliver trapped viral cargo to the lysosome for degradation. Specifically, autophagy initiates an innate immune response by cooperating with pattern recognition receptor signalling to induce interferon production. It also selectively degrades immune components associated with viral particles. Following degradation, autophagy coordinates adaptive immunity by delivering virus-derived antigens for presentation to T lymphocytes. However, in an ongoing evolutionary arms race, viruses have acquired the potent ability to hijack and subvert autophagy for their benefit. In this Review, we focus on the key regulatory steps during viral infection in which autophagy is involved and discuss the specific molecular mechanisms that diverse viruses use to repurpose autophagy for their life cycle and pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Levine, B., Mizushima, N. & Virgin, H. W. Autophagy in immunity and inflammation. Nature 469, 323–335 (2011).
Levine, B. & Kroemer, G. Autophagy in the pathogenesis of disease. Cell 132, 27–42 (2008).
Mizushima, N., Yoshimori, T. & Ohsumi, Y. The role of Atg proteins in autophagosome formation. Annu. Rev. Cell. Dev. Biol. 27, 107–132 (2011).
Mijaljica, D., Prescott, M. & Devenish, R. J. Microautophagy in mammalian cells: revisiting a 40-year-old conundrum. Autophagy 7, 673–682 (2011).
Sahu, R. et al. Microautophagy of cytosolic proteins by late endosomes. Dev. Cell. 20, 131–139 (2011).
Orenstein, S. J. & Cuervo, A. M. Chaperone-mediated autophagy: molecular mechanisms and physiological relevance. Semin. Cell. Dev. Biol. 21, 719–726 (2010).
Shibutani, S. T. & Yoshimori, T. A current perspective of autophagosome biogenesis. Cell Res. 24, 58–68 (2014).
Lamb, C. A., Yoshimori, T. & Tooze, S. A. The autophagosome: origins unknown, biogenesis complex. Nat. Rev. Mol. Cell. Biol. 14, 759–774 (2013).
Mizushima, N. & Levine, B. Autophagy in mammalian development and differentiation. Nat. Cell. Biol. 12, 823–830 (2010).
Senft, D. & Ronai, Z. A. UPR, autophagy, and mitochondria crosstalk underlies the ER stress response. Trends. Biochem. Sci. 40, 141–148 (2015).
Deretic, V., Saitoh, T. & Akira, S. Autophagy in infection, inflammation and immunity. Nat. Rev. Immunol. 13, 722–737 (2013).
Romao, S., Gannage, M. & Munz, C. Checking the garbage bin for problems in the house, or how autophagy assists in antigen presentation to the immune system. Semin. Cancer. Biol. 23, 391–396 (2013).
Paludan, C. et al. Endogenous MHC class II processing of a viral nuclear antigen after autophagy. Science 307, 593–596 (2005). In this study, autophagy-mediated lysosomal processing contributes to MHC class II-restricted surveillance of EBV to generate EBNA1-specific CD4+ T cells.
Levine, B. Eating oneself and uninvited guests: autophagy-related pathways in cellular defense. Cell 120, 159–162 (2005).
Heaton, N. S. & Randall, G. Dengue virus and autophagy. Viruses 3, 1332–1341 (2011).
Lee, M. S. & Min, Y. J. Signaling pathways downstream of pattern-recognition receptors and their cross talk. Annu. Rev. Biochem. 76, 447–480 (2007).
Yang, Q. et al. TRIM32-TAX1BP1-dependent selective autophagic degradation of TRIF negatively regulates TLR3/4-mediated innate immune responses. PLoS Pathog. 13, e1006600 (2017).
Lee, H. K., Lund, J. M., Ramanathan, B., Mizushima, N. & Iwasaki, A. Autophagy-dependent viral recognition by plasmacytoid dendritic cells. Science 315, 1398–1401 (2007). This study reveals that autophagy occurs in pDCs and is required for recognition of viral infection and production of type I IFNs, highlighting the importance of autophagy in innate immunity against viral pathogenesis.
Yordy, B., Tal, M. C., Hayashi, K., Arojo, O. & Iwasaki, A. Autophagy and selective deployment of Atg proteins in antiviral defense. Int. Immunol. 25, 1–10 (2013).
Chiang, J. J., Davis, M. E. & Gack, M. U. Regulation of RIG-I-like receptor signaling by host and viral proteins. Cytokine Growth Factor Rev. 25, 491–505 (2014).
Jounai, N. et al. The Atg5 Atg12 conjugate associates with innate antiviral immune responses. Proc. Natl. Acad. Sci. USA 104, 14050–14055 (2007).
Lei, Y. et al. The mitochondrial proteins NLRX1 and TUFM form a complex that regulates type I interferon and autophagy. Immunity. 36, 933–946 (2012).
Talloczy, Z. et al. Regulation of starvation- and virus-induced autophagy by the eIF2alpha kinase signaling pathway. Proc. Natl. Acad. Sci. USA 99, 190–195 (2002).
Sun, L. J., Wu, J. X., Du, F. H., Chen, X. & Chen, Z. J. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Liang, Q. M. et al. Crosstalk between the cGAS DNA sensor and Beclin-1 autophagy protein shapes innate antimicrobial immune responses. Cell Host Microbe 15, 228–238 (2014). In this study, the interaction between cGAS and Beclin 1 suppresses cGAMP synthesis and enhances the autophagic degradation of cytosolic DNA to prevent persistent immune activation.
Konno, H., Konno, K. & Barber, G. N. Cyclic dinucleotides trigger ULK1 (ATG1) phosphorylation of STING to prevent sustained innate immune signaling. Cell 155, 688–698 (2013).
Saitoh, T. et al. Atg9a controls dsDNA-driven dynamic translocation of STING and the innate immune response. Proc. Natl. Acad. Sci. USA 106, 20842–20846 (2009).
Hu, M. M. et al. Sumoylation promotes the stability of the DNA sensor cGAS and the adaptor STING to regulate the kinetics of response to DNA virus. Immunity. 45, 555–569 (2016).
Kessler, D. S., Veals, S. A., Fu, X. Y. & Levy, D. E. Interferon-alpha regulates nuclear translocation and DNA-binding affinity of ISGF3, a multimeric transcriptional activator. Genes Dev. 4, 1753–1765 (1990).
Platanias, L. C., Uddin, S., Yetter, A., Sun, X. J. & White, M. F. The type I interferon receptor mediates tyrosine phosphorylation of insulin receptor substrate 2. J. Biol. Chem. 271, 278–282 (1996).
Burfoot, M. S. et al. Janus kinase-dependent activation of insulin receptor substrate 1 in response to interleukin-4, oncostatin M, and the interferons. J. Biol. Chem. 272, 24183–24190 (1997).
Mammucari, C., Schiaffino, S. & Sandri, M. Downstream of Akt: FoxO3 and mTOR in the regulation of autophagy in skeletal muscle. Autophagy 4, 524–526 (2008).
Liang, X. H. et al. Protection against fatal Sindbis virus encephalitis by beclin, a novel Bcl-2-interacting protein. J. Virol. 72, 8586–8596 (1998). This study is the first to identify one of the most important mammalian autophagy proteins, Beclin 1, as a BCL-2-binding protein upon viral infection.
Orvedahl, A. et al. Autophagy protects against Sindbis virus infection of the central nervous system. Cell Host Microbe 7, 115–127 (2010).
Orvedahl, A. et al. Image-based genome-wide siRNA screen identifies selective autophagy factors. Nature 480, 113–117 (2011). This study identifies molecular determinants of selective autophagy in mammalian cells using a small interfering RNA (siRNA) screen and identifies SMURF1 as a mediator of viral autophagy and mitophagy.
Sumpter, R. Jr. et al. Fanconi anemia proteins function in mitophagy and immunity. Cell 165, 867–881 (2016).
Sumpter, R. Jr. & Levine, B. Selective autophagy and viruses. Autophagy 7, 260–265 (2011).
Staring, J. et al. PLA2G16 represents a switch between entry and clearance of Picornaviridae. Nature 541, 412–416 (2017). This study uncovers how host and virus compete during viral entry: activation of a pore-activated (galectin 8-mediated) autophagic degradation pathway and recruitment of the host factor PLA2G16 by the virus to enable viral genome release.
Shi, J. et al. Cleavage of sequestosome 1/p62 by an enteroviral protease results in disrupted selective autophagy and impaired NFkB signaling. Autophagy 9, 1591–1603 (2013).
Ait-Goughoulte, M. et al. Hepatitis C virus genotype 1a growth and induction of autophagy. J. Virol. 82, 2241–2249 (2008).
Kim, N. et al. Interferon-inducible protein SCOTIN interferes with HCV replication through the autolysosomal degradation of NS5A. Nat. Commun. 7, e10631 (2016).
Marin, M., Rose, K. M., Kozak, S. L. & Kabat, D. HIV-1 Vif protein binds the editing enzyme APOBEC3G and induces its degradation. Nat. Med. 9, 1398–1403 (2003).
Valera, M. S. et al. The HDAC6/APOBEC3G complex regulates HIV-1 infectiveness by inducing Vif autophagic degradation. Retrovirology 12, 53 (2015).
Sagnier, S. et al. Autophagy restricts HIV-1 infection by selectively degrading Tat in CD4+ T lymphocytes. J. Virol. 89, 615–625 (2015).
Ribeiro, C. M. et al. Receptor usage dictates HIV-1 restriction by human TRIM5alpha in dendritic cell subsets. Nature 540, 448–452 (2016). In this study, a known HIV-1 restriction factor, TRIM5α, is connected to the autophagic degradation pathway through the C-type lectin langerin.
Baldridge, M. T., Turula, H. & Wobus, C. E. Norovirus regulation by host and microbe. Trends Mol. Med. 22, 1047–1059 (2016).
Hwang, S. et al. Nondegradative role of Atg5-Atg12/ Atg16L1 autophagy protein complex in antiviral activity of interferon gamma. Cell Host Microbe 11, 397–409 (2012).
Cadwell, K. et al. Virus-plus-susceptibility gene interaction determines Crohn’s disease gene Atg16L1 phenotypes in intestine. Cell 141, 1135–1145 (2010).
Biering, S. B. et al. Viral replication complexes are targeted by LC3-guided interferon-inducible GTPases. Cell Host Microbe 22, 74–85.e7 (2017). This study represents an example of how autophagy (the LC3-conjugation system) and the innate immune response (IFN-inducible GTPases) cooperate to control viral infection.
Dengjel, J. et al. Autophagy promotes MHC class II presentation of peptides from intracellular source proteins. Proc. Natl. Acad. Sci. USA 102, 7922–7927 (2005).
Kyei, G. B. et al. Autophagy pathway intersects with HIV-1 biosynthesis and regulates viral yields in macrophages. J. Cell. Biol. 186, 255–268 (2009).
Blanchet, F. P. et al. Human immunodeficiency virus-1 inhibition of immunoamphisomes in dendritic cells impairs early innate and adaptive immune responses. Immunity 32, 654–669 (2010).
Loi, M. et al. Macroautophagy proteins control MHC class I levels on dendritic cells and shape anti-viral CD8(+) T cell responses. Cell Rep. 15, 1076–1087 (2016).
Parekh, V. V. et al. Autophagy-related protein Vps34 controls the homeostasis and function of antigen cross-presenting CD8alpha(+) dendritic cells. Proc. Natl Acad. Sci. USA 114, E6371–E6380 (2017).
Verweij, M. C. et al. Viral inhibition of the transporter associated with antigen processing (TAP): a striking example of functional convergent evolution. PLoS Pathog. 11, e1004743 (2015).
English, L. et al. Autophagy enhances the presentation of endogenous viral antigens on MHC class I molecules during HSV-1 infection. Nat. Immunol. 10, 480–487 (2009). This study indicates that autophagy benefits the host by providing an additional pathway for the degradation of endogenous HSV-1 proteins for antigen presentation.
Tey, S. K. & Khanna, R. Autophagy mediates transporter associated with antigen processing-independent presentation of viral epitopes through MHC class I pathway. Blood 120, 994–1004 (2012).
Tey, S. K., Goodrum, F. & Khanna, R. CD8+ T-cell recognition of human cytomegalovirus latency-associated determinant pUL138. J. Gen. Virol. 91, 2040–2048 (2010).
Shibutani, S. T., Saitoh, T., Nowag, H., Munz, C. & Yoshimori, T. Autophagy and autophagy-related proteins in the immune system. Nat. Immunol. 16, 1014–1024 (2015).
Dasari, V. et al. Autophagy and proteasome interconnect to coordinate cross-presentation through MHC class I pathway in B cells. Immunol. Cell. Biol. 94, 964–974 (2016).
Cruz, F. M., Colbert, J. D., Merino, E., Kriegsman, B. A. & Rock, K. L. The biology and underlying mechanisms of cross-presentation of exogenous antigens on MHC-I molecules. Annu. Rev. Immunol. 35, 149–176 (2017).
Mintern, J. D. et al. Differential use of autophagy by primary dendritic cells specialized in cross-presentation. Autophagy 11, 906–917 (2015).
Ravindran, R. et al. Vaccine activation of the nutrient sensor GCN2 in dendritic cells enhances antigen presentation. Science 343, 313–317 (2014). In this study, an unknown mechanism of vaccine-mediated protection reveals that stress-response dependent initiation of autophagy enhances antigen presentation to T cells.
Smed-Sorensen, A. et al. Influenza A virus infection of human primary dendritic cells impairs their ability to cross-present antigen to CD8 T cells. PLoS Pathog. 8, e1002572 (2012).
Mulvey, M., Poppers, J., Sternberg, D. & Mohr, I. Regulation of eIF2alpha phosphorylation by different functions that act during discrete phases in the herpes simplex virus type 1 life cycle. J. Virol. 77, 10917–10928 (2003).
Kanai, R. et al. Effect of gamma34.5 deletions on oncolytic herpes simplex virus activity in brain tumors. J. Virol. 86, 4420–4431 (2012).
Chaumorcel, M. et al. The human cytomegalovirus protein TRS1 inhibits autophagy via its interaction with Beclin 1. J. Virol. 86, 2571–2584 (2012).
Mouna, L. et al. Analysis of the role of autophagy inhibition by two complementary human cytomegalovirus BECN1/Beclin 1-binding proteins. Autophagy 12, 327–342 (2016).
Cuconati, A. & White, E. Viral homologs of BCL-2: role of apoptosis in the regulation of virus infection. Genes Dev. 16, 2465–2478 (2002).
Pattingre, S. et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 122, 927–939 (2005). This study reveals that cellular and viral (KSHV) BCL-2 interact with Beclin 1 and inhibit the Beclin 1-associated class III PI3K activity, resulting in the inhibition of autophagy.
Ku, B. et al. Structural and biochemical bases for the inhibition of autophagy and apoptosis by viral BCL-2 of murine gamma-herpesvirus 68. PLoS. Pathog. 4, e25 (2008).
Wei, Y., Sinha, S. & Levine, B. Dual role of JNK1-mediated phosphorylation of Bcl-2 in autophagy and apoptosis regulation. Autophagy 4, 949–951 (2008).
Yamamoto, K., Ichijo, H. & Korsmeyer, S. J. BCL-2 is phosphorylated and inactivated by an ASK1/Jun N-terminal protein kinase pathway normally activated at G(2)/M. Mol. Cell. Biol. 19, 8469–8478 (1999).
Altmann, M. & Hammerschmidt, W. Epstein-Barr virus provides a new paradigm: a requirement for the immediate inhibition of apoptosis. PLoS Biol. 3, e404 (2005).
Liang, Q. et al. Kaposi’s sarcoma-associated herpesvirus K7 modulates Rubicon-mediated inhibition of autophagosome maturation. J. Virol. 87, 12499–12503 (2013).
Lee, J. S. et al. FLIP-mediated autophagy regulation in cell death control. Nat. Cell. Biol. 11, 1355–1362 (2009).
Campbell, G. R., Rawat, P., Bruckman, R. S. & Spector, S. A. Human immunodeficiency virus type 1 Nef inhibits autophagy through transcription factor EB sequestration. PLoS. Pathog. 11, e1005018 (2015).
Dales, S., Eggers, H. J., Tamm, I. & Palade, G. E. Electron microscopic study of the formation of poliovirus. Virology. 26, 379–389 (1965). This study is the first to represent virus-mediated membrane-enclosed bodies in the cytoplasm upon poliovirus infection; later studies reveal these as DMVs.
Jackson, W. T. et al. Subversion of cellular autophagosomal machinery by RNA viruses. PLoS Biol. 3, e156 (2005).
Robinson, S. M. et al. Coxsackievirus B exits the host cell in shed microvesicles displaying autophagosomal markers. PLoS. Pathog. 10, e1004045 (2014).
Berryman, S. et al. Foot-and-mouth disease virus induces autophagosomes during cell entry via a class III phosphatidylinositol 3-kinase-independent pathway. J. Virol. 86, 12940–12953 (2012).
Wong, J. et al. Autophagosome supports coxsackievirus B3 replication in host cells. J. Virol. 82, 9143–9153 (2008).
O’Donnell, V. et al. Foot-and-mouth disease virus utilizes an autophagic pathway during viral replication. Virology. 410, 142–150 (2011).
Prentice, E., Jerome, W. G., Yoshimori, T., Mizushima, N. & Denison, M. R. Coronavirus replication complex formation utilizes components of cellular autophagy. J. Biol. Chem. 279, 10136–10141 (2004).
de Haan, C. A. & Reggiori, F. Are nidoviruses hijacking the autophagy machinery? Autophagy 4, 276–279 (2008).
Zhao, Z. J. et al. Coronavirus replication does not require the autophagy gene ATG5. Autophagy 3, 581–585 (2007).
Reggiori, F. et al. Coronaviruses Hijack the LC3-I-positive EDEMosomes, ER-derived vesicles exporting short-lived ERAD regulators, for replication. Cell. Host. Microbe. 7, 500–508 (2010).
Paul, D. & Bartenschlager, R. Flaviviridae replication organelles: oh, what a tangled web we weave. Annu. Rev. Virol. 2, 289–310 (2015).
Miller, S., Kastner, S., Krijnse-Locker, J., Buhler, S. & Bartenschlager, R. The non-structural protein 4A of dengue virus is an integral membrane protein inducing membrane alterations in a 2K-regulated manner. J. Biol. Chem. 282, 8873–8882 (2007).
Blazquez, A. B., Martin-Acebes, M. A. & Saiz, J. C. Amino acid substitutions in the non-structural proteins 4A or 4B modulate the induction of autophagy in west nile virus infected cells independently of the activation of the unfolded protein response. Front. Microbiol. 5, e797 (2014).
Cortese, M. et al. Ultrastructural characterization of zika virus replication factories. Cell Rep. 18, 2113–2123 (2017).
Offerdahl, D. K., Dorward, D. W., Hansen, B. T. & Bloom, M. E. Cytoarchitecture of zika virus infection in human neuroblastoma and aedes albopictus cell lines. Virology. 501, 54–62 (2017).
Hamel, R. et al. Biology of zika virus infection in human skin cells. J. Virol. 89, 8880–8896 (2015).
Cao, B., Parnell, L. A., Diamond, M. S. & Mysorekar, I. U. Inhibition of autophagy limits vertical transmission of zika virus in pregnant mice. J. Exp. Med. 214, 2303–2313 (2017).
Liang, Q. et al. Zika virus NS4A and NS4B proteins deregulate Akt-mTOR signaling in human fetal neural stem cells to inhibit neurogenesis and induce autophagy. Cell. Stem. Cell. 19, 663–671 (2016).
Lennemann, N. J. & Coyne, C. B. Dengue and zika viruses subvert reticulophagy by NS2B3-mediated cleavage of FAM134B. Autophagy 13, 322–332 (2017).
Khaminets, A. et al. Regulation of endoplasmic reticulum turnover by selective autophagy. Nature 522, 354–358 (2015). This study shows that reticulon protein FAM134B targets the ER to autophagosomes and regulates ER structure and turnover as a determinant for reticulophagy.
Guevin, C. et al. Autophagy protein ATG5 interacts transiently with the hepatitis C virus RNA polymerase (NS5B) early during infection. Virology. 405, 1–7 (2010).
Wang, L., Tian, Y. & Ou, J. H. HCV induces the expression of Rubicon and UVRAG to temporally regulate the maturation of autophagosomes and viral replication. PLoS. Pathog. 11, e1004764 (2015).
Gregoire, I. P. et al. IRGM is a common target of RNA viruses that subvert the autophagy network. PLoS Pathog. 7, e1002422 (2011).
Hansen, M. D. et al. Hepatitis C virus triggers Golgi fragmentation and autophagy through the immunity-related GTPase M. Proc. Natl. Acad. Sci. USA 114, 3462–3471 (2017).
Chauhan, S., Mandell, M. A. & Deretic, V. IRGM governs the core autophagy machinery to conduct antimicrobial defense. Mol. Cell. 58, 507–521 (2015). This study, together with references 100 and 101, identifies IRGM as a host factor targeted by RNA viruses for their optimal infection. Data analysis reveals that RNA viruses commonly target IRGM in the autophagic pathway. HCV utilizes IRGM for the induction of autophagy and membrane remodelling.
Zhou, Z. et al. Autophagy is involved in influenza A virus replication. Autophagy 5, 321–328 (2009).
Chen, B. J., Leser, G. P., Jackson, D. & Lamb, R. A. The influenza virus M2 protein cytoplasmic tail interacts with the M1 protein and influences virus assembly at the site of virus budding. J. Virol. 82, 10059–10070 (2008).
Beale, R. et al. A LC3-interacting motif in the influenza A virus M2 protein is required to subvert autophagy and maintain virion stability. Cell. Host. Microbe. 15, 239–247 (2014).
Zhirnov, O. P. & Klenk, H. D. Influenza A virus proteins NS1 and hemagglutinin along with M2 are involved in stimulation of autophagy in infected cells. J. Virol. 87, 13107–13114 (2013).
Ma, J. H., Sun, Q., Mi, R. F. & Zhang, H. B. Avian influenza A virus H5N1 causes autophagy-mediated cell death through suppression of mTOR signaling. J. Genet. Genom. 38, 533–537 (2011).
Ding, B. et al. Phosphoprotein of human parainfluenza virus type 3 blocks autophagosome-lysosome fusion to increase virus production. Cell Host Microbe 15, 564–577 (2014).
Itakura, E., Kishi-Itakura, C. & Mizushima, N. The hairpin-type tail-anchored SNARE syntaxin 17 targets to autophagosomes for fusion with endosomes/lysosomes. Cell 151, 1256–1269 (2012).
Ding, B. et al. The matrix protein of human parainfluenza virus type 3 induces mitophagy that suppresses interferon responses. Cell. Host. Microbe. 21, 538–547.e4 (2017).
Singh, R. et al. Autophagy regulates lipid metabolism. Nature 458, 1131–1135 (2009). In this study, lipid droplets and autophagic components associate during nutrient deprivation, and inhibition of autophagy increases triglyceride in lipid droplets, suggesting a critical function for autophagy in lipid metabolism.
Thiele, C. & Spandl, J. Cell biology of lipid droplets. Curr. Opin. Cell. Biol. 20, 378–385 (2008).
Liu, K. & Czaja, M. J. Regulation of lipid stores and metabolism by lipophagy. Cell Death Differ. 20, 3–11 (2013).
Samsa, M. M. et al. Dengue virus capsid protein usurps lipid droplets for viral particle formation. PLoS Pathog. 5, e1000632 (2009). This study reports that DENV infection increases the number of lipid droplets and accumulates mature capsid protein on the surface of lipid droplets for viral replication.
Heaton, N. S. & Randall, G. Dengue virus-induced autophagy regulates lipid metabolism. Cell Host Microbe 8, 422–432 (2010).
Jordan, T. X. & Randall, G. Dengue virus activates the AMP kinase-mTOR axis to stimulate a proviral lipophagy. J. Virol. 91, e02020–16 (2017).
McLauchlan, J. Lipid droplets and hepatitis C virus infection. Biochim. Biophys. Acta. 1791, 552–559 (2009).
Kim, J. Y., Wang, L., Lee, J. & Ou, J. J. Hepatitis C virus induces the localization of lipid rafts to autophagosomes for its RNA replication. J. Virol. 91, e00541–00517 (2017).
Saxena, V., Lai, C. K., Chao, T. C., Jeng, K. S. & Lai, M. M. C. Annexin A2 is involved in the formation of hepatitis C virus replication complex on the lipid raft. J. Virol. 86, 4139–4150 (2012).
Munz, C. The autophagic machinery in viral exocytosis. Front. Microbiol. 8, 269 (2017). This review article presents the relationships between autophagy and exocytosis and how viruses exit infected cells in packages wrapped into autophagic membranes.
Johnson, D. C. & Baines, J. D. Herpesviruses remodel host membranes for virus egress. Nat. Rev. Microbiol. 9, 382–394 (2011).
Nowag, H. et al. Macroautophagy proteins assist Epstein Barr virus production and get incorporated into the virus particles. EBioMedicine 1, 116–125 (2014).
Buckingham, E. M., Jarosinski, K. W., Jackson, W., Carpenter, J. E. & Grose, C. Exocytosis of varicella-zoster virus virions involves a convergence of endosomal and autophagy pathways. J. Virol. 90, 8673–8685 (2016).
Chen, Y. H. et al. Phosphatidylserine vesicles enable efficient en bloc transmission of enteroviruses. Cell 160, 619–630 (2015).
Bird, S. W., Maynard, N. D., Covert, M. W. & Kirkegaard, K. Nonlytic viral spread enhanced by autophagy components. Proc. Natl. Acad. Sci. USA 111, 13081–13086 (2014).
Metz, P. et al. Dengue virus inhibition of autophagic flux and dependency of viral replication on proteasomal degradation of the autophagy receptor p62. J. Virol. 89, 8026–8041 (2015).
Shrivastava, S. et al. Knockdown of autophagy inhibits infectious hepatitis C virus release by the exosomal pathway. J. Virol. 90, 1387–1396 (2015).
Orvedahl, A. et al. HSV-1 ICP34.5 confers neurovirulence by targeting the Beclin 1 autophagy protein. Cell. Host. Microbe. 1, 23–35 (2007). In this study, a mutant HSV-1 lacking the Beclin 1-binding domain of ICP34.5 fails to inhibit autophagy and demonstrates impaired viral neurovirulence.
Cabrera, C. V. et al. Herpes simplex virus DNA sequences in the CNS of latently infected mice. Nature 288, 288–290 (1980).
Chou, J., Kern, E. R., Whitley, R. J. & Roizman, B. Mapping of herpes simplex virus-1 neurovirulence to gamma 134.5, a gene nonessential for growth in culture. Science 250, 1262–1266 (1990).
Yordy, B., Iijima, N., Huttner, A., Leib, D. & Iwasaki, A. A neuron-specific role for autophagy in antiviral defense against herpes simplex virus. Cell Host Microbe 12, 334–345 (2012).
Zhong, Z., Sanchez-Lopez, E. & Karin, M. Autophagy, inflammation, and immunity: a troika governing cancer and its treatment. Cell 166, 288–298 (2016).
Hung, C. H. et al. Regulation of autophagic activation by Rta of Epstein-Barr virus via the extracellular signal-regulated kinase pathway. J. Virol. 88, 12133–12145 (2014).
Granato, M. et al. Epstein-barr virus blocks the autophagic flux and appropriates the autophagic machinery to enhance viral replication. J. Virol. 88, 12715–12726 (2014).
De Leo, A. et al. Inhibition of autophagy in EBV-positive Burkitt’s lymphoma cells enhances EBV lytic genes expression and replication. Cell Death Dis. 6, e1876 (2015).
Liu, B. et al. Hepatitis B virus X protein inhibits autophagic degradation by impairing lysosomal maturation. Autophagy 10, 416–430 (2014).
Dash, S. et al. Hepatitis C virus infection induces autophagy as a prosurvival mechanism to alleviate hepatic ER-stress response. Viruses 8, 150 (2016).
Dash, S. et al. Autophagy in hepatocellular carcinomas: from pathophysiology to therapeutic response. Hepat. Med. 8, 9–20 (2016).
Delorme-Axford, E. et al. Human placental trophoblasts confer viral resistance to recipient cells. Proc. Natl. Acad. Sci. USA 110, 12048–12053 (2013).
Screaton, G., Mongkolsapaya, J., Yacoub, S. & Roberts, C. New insights into the immunopathology and control of dengue virus infection. Nat. Rev. Immunol. 15, 745–759 (2015).
Huang, X. W. et al. Antibody-dependent enhancement of dengue virus infection inhibits RLR-mediated Type-I IFN-independent signalling through upregulation of cellular autophagy. Sci. Rep. 6, 22303 (2016).
Fang, Y. T. et al. Autophagy facilitates antibody-enhanced dengue virus infection in human pre-basophil/mast cells. PLoS. ONE. 9, e110655 (2014).
Kemball, C. C. et al. Coxsackievirus infection induces autophagy-like vesicles and megaphagosomes in pancreatic acinar cells in vivo. J. Virol. 84, 12110–12124 (2010).
Alirezaei, M., Flynn, C. T., Wood, M. R. & Whitton, J. L. Pancreatic acinar cell-specific autophagy disruption reduces coxsackievirus replication and pathogenesis in vivo. Cell. Host. Microbe. 11, 298–305 (2012). This study shows that coxsackievirus replication and pathology is decreased in the absence of autophagy due to ATG5 deficiency and that there is a reduction of virus-mediated membrane vesicles.
Ohsumi, Y. Historical landmarks of autophagy research. Cell Res. 24, 9–23 (2014).
Hara, T. et al. FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells. J. Cell. Biol. 181, 497–510 (2008).
Hosokawa, N. et al. Atg101, a novel mammalian autophagy protein interacting with Atg13. Autophagy 5, 973–979 (2009).
Mercer, C. A., Kaliappan, A. & Dennis, P. B. A novel, human Atg13 binding protein, Atg101, interacts with ULK1 and is essential for macroautophagy. Autophagy 5, 649–662 (2009).
Itakura, E., Kishi, C., Inoue, K. & Mizushima, N. Beclin 1 forms two distinct phosphatidylinositol 3-kinase complexes with mammalian Atg14 and UVRAG. Mol. Biol. Cell. 19, 5360–5372 (2008).
Matsunaga, K. et al. Two Beclin 1-binding proteins, Atg14L and Rubicon, reciprocally regulate autophagy at different stages. Nat. Cell. Biol. 11, 385–396 (2009).
Fujita, N. et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol. Biol. Cell. 19, 2092–2100 (2008).
Russell, R. C. et al. ULK1 induces autophagy by phosphorylating Beclin-1 and activating VPS34 lipid kinase. Nat. Cell. Biol. 15, 741–750 (2013).
Egan, D. F. et al. Small molecule inhibition of the autophagy kinase ULK1 and identification of ULK1 substrates. Mol. Cell. 59, 285–297 (2015).
Ge, L., Melville, D., Zhang, M. & Schekman, R. The ER-Golgi intermediate compartment is a key membrane source for the LC3 lipidation step of autophagosome biogenesis. eLife 2, e00947 (2013).
Hailey, D. W. et al. Mitochondria supply membranes for autophagosome biogenesis during starvation. Cell 141, 656–667 (2010).
Hamasaki, M. et al. Autophagosomes form at ER-mitochondria contact sites. Nature 495, 389–393 (2013).
Hayashi-Nishino, M. et al. A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nat. Cell. Biol. 11, 1433–1437 (2009).
Ravikumar, B., Moreau, K., Jahreiss, L., Puri, C. & Rubinsztein, D. C. Plasma membrane contributes to the formation of pre-autophagosomal structures. Nat. Cell. Biol. 12, 747–757 (2010).
Proikas-Cezanne, T., Takacs, Z., Donnes, P. & Kohlbacher, O. WIPI proteins: essential PtdIns3P effectors at the nascent autophagosome. J. Cell. Sci. 128, 207–217 (2015).
Ichimura, Y. et al. A ubiquitin-like system mediates protein lipidation. Nature 408, 488–492 (2000).
Kirisako, T. et al. The reversible modification regulates the membrane-binding state of Apg8/Aut7 essential for autophagy and the cytoplasm to vacuole targeting pathway. J. Cell. Biol. 151, 263–276 (2000).
Tanida, I., Ueno, T. & Kominami, E. LC3 conjugation system in mammalian autophagy. Int. J. Biochem. Cell. Biol. 36, 2503–2518 (2004).
Mizushima, N. et al. A protein conjugation system essential for autophagy. Nature 395, 395–398 (1998). This paper is the first to describe the ubiquitylation-like conjugation system in yeast at the molecular level, discovering the E1-like enzyme ATG7.
Nakatogawa, H., Ichimura, Y. & Ohsumi, Y. Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell 130, 165–178 (2007).
McEwan, D. G. et al. PLEKHM1 regulates autophagosome-lysosome fusion through HOPS complex and LC3/GABARAP proteins. Mol. Cell. 57, 39–54 (2015).
Kim, I., Rodriguez-Enriquez, S. & Lemasters, J. J. Selective degradation of mitochondria by mitophagy. Arch. Biochem. Biophys. 462, 245–253 (2007).
Kraft, C., Deplazes, A., Sohrmann, M. & Peter, M. Mature ribosomes are selectively degraded upon starvation by an autophagy pathway requiring the Ubp3p/Bre5p ubiquitin protease. Nat. Cell. Biol. 10, 602–610 (2008).
Hutchins, M. U., Veenhuis, M. & Klionsky, D. J. Peroxisome degradation in Saccharomyces cerevisiae is dependent on machinery of macroautophagy and the Cvt pathway. J. Cell. Sci. 112, 4079–4087 (1999).
Thurston, T. L., Ryzhakov, G., Bloor, S., von Muhlinen, N. & Randow, F. The TBK1 adaptor and autophagy receptor NDP52 restricts the proliferation of ubiquitin-coated bacteria. Nat. Immunol. 10, 1215–1221 (2009).
Wild, P. et al. Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science 333, 228–233 (2011).
Zheng, Y. T. et al. The adaptor protein p62/SQSTM1 targets invading bacteria to the autophagy pathway. J. Immunol. 183, 5909–5916 (2009).
Kirkin, V. et al. A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol. Cell. 33, 505–516 (2009).
Thurston, T. L., Wandel, M. P., von Muhlinen, N., Foeglein, A. & Randow, F. Galectin 8 targets damaged vesicles for autophagy to defend cells against bacterial invasion. Nature 482, 414–418 (2012).
Acknowledgements
During the preparation of this Review article, the authors were supported by grants from the US National Institutes of Health (CA200422, CA180779, DE023926, AI073099, AI116585 and AI129496) and the Fletcher Jones Foundation. The authors sincerely apologize for not being able to cite all papers related to this topic owing to space limitations.
Author information
Authors and Affiliations
Contributions
Y.C. and J.U.J. substantially contributed to the discussion of content. All authors wrote the article and reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Multivesicular bodies
-
(MVBs). A specialized subset of endosomes that contain membrane-bound intraluminal vesicles.
- Unfolded protein response
-
(UPR). A cellular stress response activated in response to an accumulation of unfolded or aggregated proteins in the endoplasmic reticulum.
- Type I interferon
-
(IFN). A subset of interferons, including IFNα, IFNβ and IFNω, that have potent antiviral roles by binding to a common receptor (interferon receptor (IFNAR)).
- Plasmacytoid dendritic cells
-
(pDCs). A subset of dendritic cells that are distinct from conventional dendritic cells and that produce large amounts of interferon.
- IFN-stimulated genes
-
(ISGs). A subset of genes induced by the activation of the interferon receptor-mediated JAK–STAT pathway.
- E3-ubiquitin ligase
-
An enzyme catalysing transfer of ubiquitin from the activated site of an E2 enzyme to a primary amine on a lysine side chain or amino terminus of a protein.
- Restriction factor
-
An antiviral protein that is produced in the host and blocks viral replication.
- Langerhans cells
-
Immune cells found in the epidermis that function as antigen-presenting cells that bind antigens entering through the skin.
- Crohn’s disease
-
A chronic inflammatory disease of the gastrointestinal tract.
- Puncta
-
A structural marker that is involved in autophagosome formation by the dynamic process of LC3 association.
- LC3 lipidation
-
A conjugation of phosphatidylethanolamine (PE) to LC3 to form LC3–PE, which is recruited to autophagosomal membranes.
- ER-associated degradation
-
(ERAD). A process through which misfolded proteins are transported from the endoplasmic reticulum to the cytoplasm for proteasomal degradation.
- SNAP receptor
-
(SNARE). A key component of the cellular membrane fusion machinery that mediates vesicular fusion events in autophagosome–lysosome fusion.
- Lipid raft
-
A small heterogeneous cellular compartment enriched with sphingolipids and cholesterols.
Rights and permissions
About this article
Cite this article
Choi, Y., Bowman, J.W. & Jung, J.U. Autophagy during viral infection — a double-edged sword. Nat Rev Microbiol 16, 341–354 (2018). https://doi.org/10.1038/s41579-018-0003-6
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41579-018-0003-6
This article is cited by
-
Tetrandrine-driven autophagy suppresses SARS-CoV-2 replication by modulating cholesterol and IGF signaling pathways
Cell Death Discovery (2026)
-
Ubiquitination and autophagy in host–pathogen interactions: from immune surveillance to therapeutic targeting
Nature Reviews Immunology (2026)
-
Histone deacetylase 6 deacetylates and ubiquitinates ATG3 to regulate autophagy
Cell Death & Differentiation (2026)
-
Dynamic regulation of TBK1 lactylation shapes antiviral immune responses
Cellular & Molecular Immunology (2026)
-
Mechanistic study of miR-148a-3p-mediated AMPK/mTOR/S6-dependent autophagy in hepatitis B virus replication
Virus Genes (2026)