Abstract
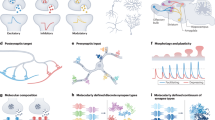
Synapses are highly specialized neuronal structures that are essential for neurotransmission, and they are dynamically regulated throughout the lifetime. Although accumulating evidence indicates that these structures are crucial for information processing and storage in the brain, their precise roles beyond neurotransmission are yet to be fully appreciated. Genetically encoded fluorescent tools have deepened our understanding of synaptic structure and function, but developing an ideal methodology to selectively visualize, label and manipulate synapses remains challenging. Here, we provide an overview of currently available synapse labelling techniques and describe their extension to enable synapse manipulation. We categorize these approaches on the basis of their conceptual bases and target molecules, compare their advantages and limitations and propose potential modifications to improve their effectiveness. These methods have broad utility, particularly for investigating mechanisms of synaptic function and synaptopathy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Aimino, M. A., Humenik, J., Parisi, M. J., Duhart, J. C. & Mosca, T. J. SynLight: a bicistronic strategy for simultaneous active zone and cell labeling in the Drosophila nervous system. G3 13, jkad221 (2023).
Talay, M. et al. Transsynaptic mapping of second-order taste neurons in flies by trans-Tango. Neuron 96, 783–795 (2017).
Oh, J.-Y. et al. Labeling dual presynaptic inputs using cfork anterograde tracing system. Exp. Neurobiol. 29, 219–229 (2020).
Cachero, S. et al. BAcTrace, a tool for retrograde tracing of neuronal circuits in Drosophila. Nat. Methods 17, 1254–1261 (2020). This work uses an engineered form of the Clostridium botulinum neurotoxin A (Botox) that is capable of retrograde trans-synaptic labelling via the activation of a silent transcription factor in the neuron.
Huang, T.-H. et al. Tracing neuronal circuits in transgenic animals by transneuronal control of transcription (TRACT). eLife 6, e32027 (2017).
Choi, J. H. et al. Interregional synaptic maps among engram cells underlie memory formation. Science 360, 430–435 (2018). This paper introduced an advance in the GRASP technique that allowed multicolour labelling of engram and non-engram synapses.
Kim, J. et al. mGRASP enables mapping mammalian synaptic connectivity with light microscopy. Nat. Methods 9, 96–102 (2012). In this work, the GRASP technique was upgraded so that it was applicable to mammalian neurons, in which it labels synapses using split green fluorescent protein expressed as part of presynaptic and postsynaptic partner proteins.
Son, S. et al. Real-time visualization of structural dynamics of synapses in live cells in vivo. Nat. Methods 21, 353–360 (2024). This paper introduced a method for real-time visualization of the structural plasticity of synapses in vivo using dimerization-dependent fluorescent proteins.
Feinberg, E. H. et al. GFP reconstitution across synaptic partners (GRASP) defines cell contacts and synapses in living nervous systems. Neuron 57, 353–363 (2008). This paper initiated the concept of using split green fluorescent protein to label presynaptic and postsynaptic interactions.
Kim, T. et al. Activated somatostatin interneurons orchestrate memory microcircuits. Neuron 112, 201–208 (2023).
Trimmer, J. S. Genetically encoded intrabodies as high-precision tools to visualize and manipulate neuronal function. Semin. Cell Dev. Biol. 126, 117–124 (2022).
Ferro, M. et al. Functional mapping of brain synapses by the enriching activity-marker SynaptoZip. Nat. Commun. 8, 1229 (2017).
Chen, Y. et al. Cell-type-specific labeling of synapses in vivo through synaptic tagging with recombination. Neuron 81, 280–293 (2014).
Perez-Alvarez, A. et al. Freeze-frame imaging of synaptic activity using SynTagMA. Nat. Commun. 11, 2464 (2020).
Lee, C. et al. Hippocampal engram networks for fear memory recruit new synapses and modify pre-existing synapses in vivo. Curr. Biol. 33, 507–516 (2023).
Ma, S. & Zuo, Y. Synaptic modifications in learning and memory — a dendritic spine story. Semin. Cell Dev. Biol. 125, 84–90 (2022).
Ultanir, S. K. et al. Regulation of spine morphology and spine density by NMDA receptor signaling in vivo. Proc. Natl Acad. Sci. USA 104, 19553–19558 (2007).
Sørensen, A. T. et al. A robust activity marking system for exploring active neuronal ensembles. eLife 5, e13918 (2016).
Ortega-de San Luis, C. & Ryan, T. J. Understanding the physical basis of memory: molecular mechanisms of the engram. J. Biol. Chem. 298, 101866 (2022).
Rao-Ruiz, P. et al. Engram-specific transcriptome profiling of contextual memory consolidation. Nat. Commun. 10, 2232 (2019).
Josselyn, S. A., Köhler, S. & Frankland, P. W. Finding the engram. Nat. Rev. Neurosci. 16, 521–534 (2015).
Choi, D. I. & Kaang, B. K. Interrogating structural plasticity among synaptic engrams. Curr. Opin. Neurobiol. 75, 102552 (2022). This review delves into the concept of engram synapses and the approaches used to label them.
Ryan, T. J., Roy, D. S., Pignatelli, M., Arons, A. & Tonegawa, S. Engram cells retain memory under retrograde amnesia. Science 348, 1007–1013 (2015).
Farhy-Tselnicker, I. & Allen, N. J. Astrocytes, neurons, synapses: a tripartite view on cortical circuit development. Neural Dev. 13, 1–12 (2018).
Silbereis, J. C., Pochareddy, S., Zhu, Y., Li, M. & Sestan, N. The cellular and molecular landscapes of the developing human central nervous system. Neuron 89, 248–268 (2016).
Waites, C. L., Craig, A. M. & Garner, C. C. Mechanisms of vertebrate synaptogenesis. Annu. Rev. Neurosci. 28, 251–274 (2005).
Lin, Y. C. & Koleske, A. J. Mechanisms of synapse and dendrite maintenance and their disruption in psychiatric and neurodegenerative disorders. Annu. Rev. Neurosci. 33, 349–378 (2010).
Peça, J. & Feng, G. Cellular and synaptic network defects in autism. Curr. Opin. Neurobiol. 22, 866–872 (2012).
Rubinov, M. & Bullmore, E. Fledgling pathoconnectomics of psychiatric disorders. Trends Cogn. Sci. 17, 641–647 (2013).
Mevel, K. & Fransson, P. The functional brain connectome of the child and autism spectrum disorders. Acta Paediatr. 105, 1024–1035 (2016).
Narr, K. L. & Leaver, A. M. Connectome and schizophrenia. Curr. Opin. Psychiatry 28, 229–235 (2015).
Sacai, H. et al. Autism spectrum disorder-like behavior caused by reduced excitatory synaptic transmission in pyramidal neurons of mouse prefrontal cortex. Nat. Commun. 11, 5140 (2020).
Guang, S. et al. Synaptopathology involved in autism spectrum disorder. Front. Cell Neurosci. 12, 470 (2018).
Dean, C. & Dresbach, T. Neuroligins and neurexins: linking cell adhesion, synapse formation and cognitive function. Trends Neurosci. 29, 21–29 (2006).
Teng, S.-W. et al. Altered fear engram encoding underlying conditioned versus unconditioned stimulus-initiated memory updating. Sci. Adv. 9, eadf0284 (2023).
Dempsey, W. P. et al. Regional synapse gain and loss accompany memory formation in larval zebrafish. Proc. Natl Acad. Sci. USA 119, e2107661119 (2022).
O’Leary, J. D., Bruckner, R., Autore, L. & Ryan, T. J. Natural forgetting reversibly modulates engram expression. eLife 12, e92860.1 (2023).
Jessica, L., Raphael, Z., Laura, H. C., Michael, S. F. & Bryce, V. Engram size varies with learning and reflects memory content and precision. J. Neurosci. 41, 4120 (2021).
Choi, J.-E., Kim, J. & Kim, J. Capturing activated neurons and synapses. Neurosci. Res. 152, 25–34 (2020). This review highlights the key techniques used to label activated neurons and synapses.
Minorikawa, S. & Nakayama, M. Recombinase-mediated cassette exchange (RMCE) and BAC engineering via VCre/VloxP and SCre/SloxP systems. Biotechniques 50, 235–246 (2011).
Fenno, L. E. et al. Targeting cells with single vectors using multiple-feature Boolean logic. Nat. Methods 11, 763–772 (2014).
Barnea, G. et al. The genetic design of signaling cascades to record receptor activation. Proc. Natl Acad. Sci. USA 105, 64–69 (2008).
Jagadish, S., Barnea, G., Clandinin, T. R. & Axel, R. Identifying functional connections of the inner photoreceptors in Drosophila using Tango-Trace. Neuron 83, 630–644 (2014).
Nässel, D. R. Histamine in the brain of insects: a review. Microsc. Res. Tech. 44, 121–136 (1999).
Hong, S.-T. et al. Histamine and its receptors modulate temperature-preference behaviors in Drosophila. J. Neurosci. 26, 7245–7256 (2006).
Fortin, J.-P. et al. Membrane-tethered ligands are effective probes for exploring class B1 G protein-coupled receptor function. Proc. Natl Acad. Sci. USA 106, 8049–8054 (2009).
Krstenansky, J. L., Trivedi, D., Johnson, D. & Hruby, V. J. Conformational considerations in the design of a glucagon analog with increased receptor binding and adenylate cyclase potencies. J. Am. Chem. Soc. 108, 1696–1698 (1986).
Potter, C. J., Tasic, B., Russler, E. V., Liang, L. & Luo, L. The Q system: a repressible binary system for transgene expression, lineage tracing, and mosaic analysis. Cell 141, 536–548 (2010).
Riabinina, O. et al. Improved and expanded Q-system reagents for genetic manipulations. Nat. Methods 12, 219–222 (2015).
Nam, H., Hwang, B. J., Choi, D. Y., Shin, S. & Choi, M. Tobacco etch virus (TEV) protease with multiple mutations to improve solubility and reduce self-cleavage exhibits enhanced enzymatic activity. FEBS Open Biol. 10, 619–626 (2020).
Pauli, A. et al. Cell-type-specific TEV protease cleavage reveals cohesin functions in Drosophila neurons. Dev. Cell 14, 239–251 (2008).
Washbourne, P., Pellizzari, R., Baldini, G., Wilson, M. C. & Montecucco, C. Botulinum neurotoxin types A and E require the SNARE motif in SNAP-25 for proteolysis. FEBS Lett. 418, 1–5 (1997).
Tomasoni, R. et al. SNAP-25 regulates spine formation through postsynaptic binding to p140Cap. Nat. Commun. 4, 2136 (2013).
Kádková, A. et al. SNAP25 disease mutations change the energy landscape for synaptic exocytosis due to aberrant SNARE interactions. eLife 12, RP88619 (2024).
Gordon, W. R. et al. Mechanical allostery: evidence for a force requirement in the proteolytic activation of Notch. Dev. Cell 33, 729–736 (2015).
Morsut, L. et al. Engineering customized cell sensing and response behaviors using synthetic notch receptors. Cell 164, 780–791 (2016).
Huang, T.-H., Velho, T. & Lois, C. Monitoring cell–cell contacts in vivo in transgenic animals. Development 143, 4073–4084 (2016).
Fujimoto, M., Poe, J. C., Inaoki, M. & Tedder, T. F. CD19 regulates B lymphocyte responses to transmembrane signals. Semin. Immunol. 10, 267–277 (1998).
Kochenderfer, J. N. et al. Construction and pre-clinical evaluation of an anti-CD19 chimeric antigen receptor. J. Immunother. 32, 689 (2009).
Kovall, R. A., Gebelein, B., Sprinzak, D. & Kopan, R. The canonical Notch signaling pathway: structural and biochemical insights into shape, sugar, and force. Dev. Cell 41, 228–241 (2017).
Sprinzak, D. et al. Cis-interactions between Notch and Delta generate mutually exclusive signalling states. Nature 465, 86–90 (2010).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Bellen, H. J., Tong, C. & Tsuda, H. 100 years of Drosophila research and its impact on vertebrate neuroscience: a history lesson for the future. Nat. Rev. Neurosci. 11, 514–522 (2010).
St Johnston, D. The art and design of genetic screens: Drosophila melanogaster. Nat. Rev. Genet. 3, 176–188 (2002).
Fetcho, J. R. & Liu, K. S. Zebrafish as a model system for studying neuronal circuits and behavior. Ann. N. Y. Acad. Sci. 860, 333–345 (1998).
Patton, E. E. & Zon, L. I. The art and design of genetic screens: zebrafish. Nat. Rev. Genet. 2, 956–966 (2001).
Anderson, K. V. & Ingham, P. W. The transformation of the model organism: a decade of developmental genetics. Nat. Genet. 33, 285–293 (2003).
Kile, B. T. & Hilton, D. J. The art and design of genetic screens: mouse. Nat. Rev. Genet. 6, 557–567 (2005).
Südhof, T. C. The presynaptic active zone. Neuron 75, 11–25 (2012).
Fouquet, W. et al. Maturation of active zone assembly by Drosophila Bruchpilot. J. Cell Biol. 186, 129–145 (2009).
Wang, Y., Liu, X., Biederer, T. & Südhof, T. C. A family of RIM-binding proteins regulated by alternative splicing: implications for the genesis of synaptic active zones. Proc. Natl Acad. Sci. USA 99, 14464–14469 (2002).
Ohtsuka, T. et al. Cast: a novel protein of the cytomatrix at the active zone of synapses that forms a ternary complex with RIM1 and munc13-1. J. Cell Biol. 158, 577–590 (2002).
Lee, T., Lee, A. & Luo, L. Development of the Drosophila mushroom bodies: sequential generation of three distinct types of neurons from a neuroblast. Development 126, 4065–4076 (1999).
Aimino, M. A., DePew, A. T., Restrepo, L. & Mosca, T. J. Synaptic development in diverse olfactory neuron classes uses distinct temporal and activity-related programs. J. Neurosci. 43, 28–55 (2023).
Mosca, T. J. & Luo, L. Synaptic organization of the Drosophila antennal lobe and its regulation by the Teneurins. eLife 3, e03726 (2014).
Christiansen, F. et al. Presynapses in Kenyon cell dendrites in the mushroom body calyx of Drosophila. J. Neurosci. 31, 9696–9707 (2011).
Kremer, M. C. et al. Structural long-term changes at mushroom body input synapses. Curr. Biol. 20, 1938–1944 (2010).
Toegel, M. et al. A multiplexable TALE-based binary expression system for in vivo cellular interaction studies. Nat. Commun. 8, 1663 (2017).
Mosca, T. J., Luginbuhl, D. J., Wang, I. E. & Luo, L. Presynaptic LRP4 promotes synapse number and function of excitatory CNS neurons. eLife 6, e27347 (2017).
Pfeiffer, B. D., Truman, J. W. & Rubin, G. M. Using translational enhancers to increase transgene expression in Drosophila. Proc. Natl Acad. Sci. USA 109, 6626–6631 (2012).
Yu, D. et al. Optogenetic activation of intracellular antibodies for direct modulation of endogenous proteins. Nat. Methods 16, 1095–1100 (2019).
Marzilli, A. M., McMahan, J. B. & Ngo, J. T. Precision control of intrabodies in live cells. Nat. Methods 17, 259–260 (2020).
Farrants, H. et al. Chemogenetic control of nanobodies. Nat. Methods 17, 279–282 (2020).
Gross, G. G. et al. Recombinant probes for visualizing endogenous synaptic proteins in living neurons. Neuron 78, 971–985 (2013).
Bensussen, S. et al. A viral toolbox of genetically encoded fluorescent synaptic tags. iScience 23, 101330 (2020).
Mani, M., Kandavelou, K., Dy, F. J., Durai, S. & Chandrasegaran, S. Design, engineering, and characterization of zinc finger nucleases. Biochem. Biophys. Res. Commun. 335, 447–457 (2005).
Gabriel, R. et al. An unbiased genome-wide analysis of zinc-finger nuclease specificity. Nat. Biotechnol. 29, 816–823 (2011).
El-Husseini, A. E.-D., Schnell, E., Chetkovich, D. M., Nicoll, R. A. & Bredt, D. S. PSD-95 involvement in maturation of excitatory synapses. Science 290, 1364–1368 (2000).
O’Shea, E. K., Lumb, K. J. & Kim, P. S. Peptide ‘Velcro’: design of a heterodimeric coiled coil. Curr. Biol. 3, 658–667 (1993).
Wang, C. et al. Different regions of synaptic vesicle membrane regulate VAMP2 conformation for the SNARE assembly. Nat. Commun. 11, 1531 (2020).
Salpietro, V. et al. Mutations in the neuronal vesicular SNARE VAMP2 affect synaptic membrane fusion and impair human neurodevelopment. Am. J. Hum. Genet. 104, 721–730 (2019).
Koo, S. J. et al. Vesicular synaptobrevin/VAMP2 levels guarded by AP180 control efficient neurotransmission. Neuron 88, 330–344 (2015).
Bhattacharya, S. et al. Members of the synaptobrevin/vesicle-associated membrane protein (VAMP) family in Drosophila are functionally interchangeable in vivo for neurotransmitter release and cell viability. Proc. Natl Acad. Sci. USA 99, 13867–13872 (2002).
Warming, S., Costantino, N., Court, D. L., Jenkins, N. A. & Copeland, N. G. Simple and highly efficient BAC recombineering using galK selection. Nucleic Acids Res. 33, e36 (2005).
Venken, K. J. T., He, Y., Hoskins, R. A. & Bellen, H. J. P[acman]: a BAC transgenic platform for targeted insertion of large DNA fragments in D. melanogaster. Science 314, 1747–1751 (2006).
Southern, J. A., Young, D. F., Heaney, F., Baumgärtner, W. K. & Randall, R. E. Identification of an epitope on the P and V proteins of simian virus 5 that distinguishes between two isolates with different biological characteristics. J. Gen. Virol. 72, 1551–1557 (1991).
Golic, K. G. & Lindquist, S. The FLP recombinase of yeast catalyzes site-specific recombination in the Drosophila genome. Cell 59, 499–509 (1989).
Lai, S.-L. & Lee, T. Genetic mosaic with dual binary transcriptional systems in Drosophila. Nat. Neurosci. 9, 703–709 (2006).
Ryan, M. D. & Drew, J. Foot‐and‐mouth disease virus 2A oligopeptide mediated cleavage of an artificial polyprotein. EMBO J. 13, 928–933 (1994).
Gengs, C. et al. The target of Drosophila photoreceptor synaptic transmission is a histamine-gated chloride channel encoded byort (hclA). J. Biol. Chem. 277, 42113–42120 (2002).
Gao, S. et al. The neural substrate of spectral preference in Drosophila. Neuron 60, 328–342 (2008).
Nern, A., Pfeiffer, B. D., Svoboda, K. & Rubin, G. M. Multiple new site-specific recombinases for use in manipulating animal genomes. Proc. Natl Acad. Sci. USA 108, 14198–14203 (2011).
Mikuni, T., Nishiyama, J., Sun, Y., Kamasawa, N. & Yasuda, R. High-throughput, high-resolution mapping of protein localization in mammalian brain by in vivo genome editing. Cell 165, 1803–1817 (2016).
Nishiyama, J., Mikuni, T. & Yasuda, R. Virus-mediated genome editing via homology-directed repair in mitotic and postmitotic cells in mammalian brain. Neuron 96, 755–768.e755 (2017).
Gao, Y. et al. Plug-and-play protein modification using homology-independent universal genome engineering. Neuron 103, 583–597.e8 (2019).
Willems, J. et al. ORANGE: a CRISPR/Cas9-based genome editing toolbox for epitope tagging of endogenous proteins in neurons. PLOS Biol. 18, e3000665 (2020).
Fang, H., Bygrave, A. M., Roth, R. H., Johnson, R. C. & Huganir, R. L. An optimized CRISPR/Cas9 approach for precise genome editing in neurons. eLife 10, e65202 (2021).
Zhong, H. et al. High-fidelity, efficient, and reversible labeling of endogenous proteins using CRISPR-based designer exon insertion. eLife 10, e64911 (2021).
Ghosh, I., Hamilton, A. D. & Regan, L. Antiparallel leucine zipper-directed protein reassembly: application to the green fluorescent protein. J. Am. Chem. Soc. 122, 5658–5659 (2000).
Zhang, S., Ma, C. & Chalfie, M. Combinatorial marking of cells and organelles with reconstituted fluorescent proteins. Cell 119, 137–144 (2004).
Pédelacq, J.-D., Cabantous, S., Tran, T., Terwilliger, T. C. & Waldo, G. S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Cabantous, S., Terwilliger, T. C. & Waldo, G. S. Protein tagging and detection with engineered self-assembling fragments of green fluorescent protein. Nat. Biotechnol. 23, 102–107 (2005). In this work, soluble and self-associating fragments of green fluorescent protein were engineered, which are capable of labelling soluble and insoluble proteins without changing their properties.
Gordon, M. D. & Scott, K. Motor control in a Drosophila taste circuit. Neuron 61, 373–384 (2009).
Fairless, R. et al. Polarized targeting of neurexins to synapses is regulated by their C-terminal sequences. J. Neurosci. 28, 12969–12981 (2008).
Zhang, C. et al. Neurexins physically and functionally interact with GABAA receptors. Neuron 66, 403–416 (2010).
Tang, W. et al. Faithful expression of multiple proteins via 2A-peptide self-processing: a versatile and reliable method for manipulating brain circuits. J. Neurosci. 29, 8621–8629 (2009).
Martell, J. D. et al. A split horseradish peroxidase for the detection of intercellular protein–protein interactions and sensitive visualization of synapses. Nat. Biotechnol. 34, 774–780 (2016). This paper reports the engineering of a split horse peroxidase capable of reconstitution into an active form upon intercellular protein–protein interaction. The use of this technique provides sensitive labelling of synapses.
Macpherson, L. J. et al. Dynamic labelling of neural connections in multiple colours by trans-synaptic fluorescence complementation. Nat. Commun. 6, 10024 (2015).
Donaldson, L. W., Gish, G., Pawson, T., Kay, L. E. & Forman-Kay, J. D. Structure of a regulatory complex involving the Abl SH3 domain, the Crk SH2 domain, and a Crk-derived phosphopeptide. Proc. Natl Acad. Sci. USA 99, 14053–14058 (2002).
Pisabarro, M. T., Serrano, L. & Wilmanns, M. Crystal structure of the abl-SH3 domain complexed with a designed high-affinity peptide ligand: implications for SH3–ligand interactions. J. Mol. Biol. 281, 513–521 (1998).
Hou, T., Chen, K., McLaughlin, W. A., Lu, B. & Wang, W. Computational analysis and prediction of the binding motif and protein interacting partners of the Abl SH3 domain. PLOS Comput. Biol. 2, e1 (2006).
Pisabarro, M. T. & Serrano, L. Rational design of specific high-affinity peptide ligands for the Abl-SH3 domain. Biochemistry 35, 10634–10640 (1996).
Nguyen, Q.-A., Horn, M. E. & Nicoll, R. A. Distinct roles for extracellular and intracellular domains in neuroligin function at inhibitory synapses. eLife 5, e19236 (2016).
Südhof, T. C. Neuroligins and neurexins link synaptic function to cognitive disease. Nature 455, 903–911 (2008). This review delves into the role of the synaptic partner proteins neuroligins and neurexins in cognitive disorders.
Chih, B., Engelman, H. & Scheiffele, P. Control of excitatory and inhibitory synapse formation by neuroligins. Science 307, 1324–1328 (2005).
Fredj, A. et al. The single T65S mutation generates brighter cyan fluorescent proteins with increased photostability and pH insensitivity. PLoS ONE 7, e49149 (2012).
Griesbeck, O., Baird, G. S., Campbell, R. E., Zacharias, D. A. & Tsien, R. Y. Reducing the environmental sensitivity of yellow fluorescent protein: mechanism and applications. J. Biol. Chem. 276, 29188–29194 (2001).
Rekas, A., Alattia, J.-R., Nagai, T., Miyawaki, A. & Ikura, M. Crystal structure of venus, a yellow fluorescent protein with improved maturation and reduced environmental sensitivity. J. Biol. Chem. 277, 50573–50578 (2002).
Kügler, S., Lingor, P., Schöll, U., Zolotukhin, S. & Bähr, M. Differential transgene expression in brain cells in vivo and in vitro from AAV-2 vectors with small transcriptional control units. Virology 311, 89–95 (2003).
Kügler, S., Kilic, E. & Bähr, M. Human synapsin 1 gene promoter confers highly neuron-specific long-term transgene expression from an adenoviral vector in the adult rat brain depending on the transduced area. Gene Ther. 10, 337–347 (2003).
Jin, L. et al. High-efficiency transduction and specific expression of ChR2opt for optogenetic manipulation of primary cortical neurons mediated by recombinant adeno-associated viruses. J. Biotechnol. 233, 171–180 (2016).
Dymecki, S. M. Flp recombinase promotes site-specific DNA recombination in embryonic stem cells and transgenic mice. Proc. Natl Acad. Sci. USA 93, 6191–6196 (1996).
Ryu, H.-H. et al. Excitatory neuron-specific SHP2-ERK signaling network regulates synaptic plasticity and memory. Sci. Signal. 12, eaau5755 (2019).
Tsetsenis, T., Boucard, A. A., Araç, D., Brunger, A. T. & Südhof, T. C. Direct visualization of trans-synaptic neurexin–neuroligin interactions during synapse formation. J. Neurosci. 34, 15083–15096 (2014).
Balleza, E., Kim, J. M. & Cluzel, P. Systematic characterization of maturation time of fluorescent proteins in living cells. Nat. Methods 15, 47–51 (2018).
Alford, S. C., Ding, Y., Simmen, T. & Campbell, R. E. Dimerization-dependent green and yellow fluorescent proteins. ACS Synth. Biol. 1, 569–575 (2012).
Ding, Y. et al. Ratiometric biosensors based on dimerization-dependent fluorescent protein exchange. Nat. Methods 12, 195–198 (2015).
Challis, R. C. et al. Adeno-associated virus toolkit to target diverse brain cells. Annu. Rev. Neurosci. 45, 447–469 (2022).
Dong, J. Y., Fan, P. D. & Frizzell, R. A. Quantitative analysis of the packaging capacity of recombinant adeno-associated virus. Hum. Gene Ther. 7, 2101–2112 (1996).
Fosque, B. F. et al. Labeling of active neural circuits in vivo with designed calcium integrators. Science 347, 755–760 (2015).
Bohra, A. A., Kallman, B. R., Reichert, H. & VijayRaghavan, K. Identification of a single pair of interneurons for bitter taste processing in the Drosophila brain. Curr. Biol. 28, 847–858 (2018).
Moeyaert, B. et al. Improved methods for marking active neuron populations. Nat. Commun. 9, 4440 (2018).
Hayashi-Takagi, A. et al. Labelling and optical erasure of synaptic memory traces in the motor cortex. Nature 525, 333–338 (2015). This paper introduces a novel synaptic optoprobe called as activated synapse targeting photoactivatable Rac1 (As-PaRac1), which labels recently activated spines. In addition, this technique is also able to manipulate the labelled spines optically.
Peter, E., Dick, B. & Baeurle, S. A. Mechanism of signal transduction of the LOV2-Jα photosensor from Avena sativa. Nat. Commun. 1, 122 (2010).
Wu, Y. I. et al. A genetically encoded photoactivatable Rac controls the motility of living cells. Nature 461, 104–108 (2009).
Borovac, J., Bosch, M. & Okamoto, K. Regulation of actin dynamics during structural plasticity of dendritic spines: signaling messengers and actin-binding proteins. Mol. Cell. Neurosci. 91, 122–130 (2018).
Luo, L. et al. Differential effects of the Rac GTPase on Purkinje cell axons and dendritic trunks and spines. Nature 379, 837–840 (1996).
Hayashi-Takagi, A. et al. Disrupted-in-schizophrenia 1 (DISC1) regulates spines of the glutamate synapse via Rac1. Nat. Neurosci. 13, 327–332 (2010).
Arnold, D. B. & Clapham, D. E. Molecular determinants for subcellular localization of PSD-95 with an interacting K+ channel. Neuron 23, 149–157 (1999).
Kobayashi, H., Yamamoto, S., Maruo, T. & Murakami, F. Identification of a cis‐acting element required for dendritic targeting of activity‐regulated cytoskeleton‐associated protein mRNA. Eur. J. Neurosci. 22, 2977–2984 (2005).
Steward, O., Wallace, C. S., Lyford, G. L. & Worley, P. F. Synaptic activation causes the mRNA for the IEG Arc to localize selectively near activated postsynaptic sites on dendrites. Neuron 21, 741–751 (1998).
Steward, O. & Worley, P. F. Selective targeting of newly synthesized Arc mRNA to active synapses requires NMDA receptor activation. Neuron 30, 227–240 (2001).
Bulina, M. E. et al. Chromophore-assisted light inactivation (CALI) using the phototoxic fluorescent protein KillerRed. Nat. Protoc. 1, 947–953 (2006). This study elaborates on the use of the fluorescent protein Killer Red to achieve targeted protein inactivation.
Takemoto, K. et al. SuperNova, a monomeric photosensitizing fluorescent protein for chromophore-assisted light inactivation. Sci. Rep. 3, 2629 (2013).
Suzuki, H. et al. In vivo imaging of C. elegans mechanosensory neurons demonstrates a specific role for the MEC-4 channel in the process of gentle touch sensation. Neuron 39, 1005–1017 (2003).
Bosch, M. et al. Structural and molecular remodeling of dendritic spine substructures during long-term potentiation. Neuron 82, 444–459 (2014).
Goto, A. et al. Stepwise synaptic plasticity events drive the early phase of memory consolidation. Science 374, 857–863 (2021).
Okamoto, K.-I., Nagai, T., Miyawaki, A. & Hayashi, Y. Rapid and persistent modulation of actin dynamics regulates postsynaptic reorganization underlying bidirectional plasticity. Nat. Neurosci. 7, 1104–1112 (2004).
Goto, A. Synaptic plasticity during systems memory consolidation. Neurosci. Res. 183, 1–6 (2022).
Ziv, Y. et al. Long-term dynamics of CA1 hippocampal place codes. Nat. Neurosci. 16, 264–266 (2013).
Liu, W., Montana, V., Chapman, E. R., Mohideen, U. & Parpura, V. Botulinum toxin type B micromechanosensor. Proc. Natl Acad. Sci. USA 100, 13621–13625 (2003).
Elliott, M. et al. Engineered botulinum neurotoxin B with improved binding to human receptors has enhanced efficacy in preclinical models. Sci. Adv. 5, eaau7196 (2019).
Tao, L. et al. Engineered botulinum neurotoxin B with improved efficacy for targeting human receptors. Nat. Commun. 8, 53 (2017).
Liu, Q. et al. A photoactivatable botulinum neurotoxin for inducible control of neurotransmission. Neuron 101, 863–875.e6 (2019).
Schiavo, G. G. et al. Tetanus and botulinum-B neurotoxins block neurotransmitter release by proteolytic cleavage of synaptobrevin. Nature 359, 832–835 (1992). This work describes the proteolytic activity of zinc endopeptidases (tetanus and botulinum toxins serotype B), which mediate the cleavage of synatobrevin-2 to inhibit neurotransmitter release.
Guntas, G. et al. Engineering an improved light-induced dimer (iLID) for controlling the localization and activity of signaling proteins. Proc. Natl Acad. Sci. USA 112, 112–117 (2015).
Zimmerman, S. P. et al. Tuning the binding affinities and reversion kinetics of a light inducible dimer allows control of transmembrane protein localization. Biochemistry 55, 5264–5271 (2016).
Segelke, B., Knapp, M., Kadkhodayan, S., Balhorn, R. & Rupp, B. Crystal structure of Clostridium botulinum neurotoxin protease in a product-bound state: evidence for noncanonical zinc protease activity. Proc. Natl Acad. Sci. USA 101, 6888–6893 (2004).
Won, J. et al. Opto-vTrap, an optogenetic trap for reversible inhibition of vesicular release, synaptic transmission, and behavior. Neuron 110, 423–435.e4 (2022).
Vettkötter, D. et al. Rapid and reversible optogenetic silencing of synaptic transmission by clustering of synaptic vesicles. Nat. Commun. 13, 7827 (2022).
Vogt, N. Improving retrograde labeling of neurons. Nat. Methods 15, 571–571 (2018).
Oyibo, H. K., Znamenskiy, P., Oviedo, H. V., Enquist, L. W. & Zador, A. M. Long-term Cre-mediated retrograde tagging of neurons using a novel recombinant pseudorabies virus. Front. Neuroanat. 8, 86 (2014).
Catapano, J. et al. Retrograde labeling of regenerating motor and sensory neurons using silicone caps. J. Neurosci. Methods 259, 122–128 (2016).
Beier, K. T. et al. Anterograde or retrograde transsynaptic labeling of CNS neurons with vesicular stomatitis virus vectors. Proc. Natl Acad. Sci. USA 108, 15414–15419 (2011).
Berezin, M. Y. & Achilefu, S. Fluorescence lifetime measurements and biological imaging. Chem. Rev. 110, 2641–2684 (2010).
Lanciego, J. L. & Wouterlood, F. G. A half century of experimental neuroanatomical tracing. J. Chem. Neuroanat. 42, 157–183 (2011).
Nassi, J. J., Cepko, C. L., Born, R. T. & Beier, K. T. Neuroanatomy goes viral! Front. Neuroanat. 9, 80 (2015).
Ugolini, G., Kuypers, H. & Strick, P. L. Transneuronal transfer of herpes virus from peripheral nerves to cortex and brainstem. Science 243, 89–91 (1989).
Callaway, E. M. Transneuronal circuit tracing with neurotropic viruses. Curr. Opin. Neurobiol. 18, 617–623 (2008). This review provides a comprehensive discussion on the use of viruses to target neuronal circuits composed of specific neuron types.
Lo, L. & Anderson, D. J. A Cre-dependent, anterograde transsynaptic viral tracer for mapping output pathways of genetically marked neurons. Neuron 72, 938–950 (2011).
Zingg, B. et al. AAV-mediated anterograde transsynaptic tagging: mapping corticocollicular input-defined neural pathways for defense behaviors. Neuron 93, 33–47 (2017).
Palmer, A. E., Qin, Y., Park, J. G. & McCombs, J. E. Design and application of genetically encoded biosensors. Trends Biotechnol. 29, 144–152 (2011).
Son, J.-H. et al. Transgenic FingRs for live mapping of synaptic dynamics in genetically-defined neurons. Sci. Rep. 6, 18734 (2016).
Acknowledgements
This work was supported by a grant from the Institute of Basic Science (IBS-R001-E1-2023-a00). Figures were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Neuroscience thanks Hyung-Bae Kwon and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Active zone
-
The part of the presynaptic membrane that is the major site for the fusion of synaptic vesicles and neurotransmitter release.
- Anterograde tracing
-
A method used to trace axonal projections from the cell body to the axon terminals.
- Binary expression system
-
A transgene expression system that is based on a single exogenous DNA-binding protein (driver) binding to a unique recognition motif (responder). For example, LexA protein recognizes the LexA operator, whereas QF DNA-binding protein acts on QUAS sequence.
- CRISPR–Cas9
-
A genome editing tool that acts as ‘molecular scissors’ by using CRISPR sequences as a guide to recognize and cut DNA at a specific location, enabling the manipulation of genetic DNA.
- Engram cells
-
Neurons that are activated during memory formation and are believed to be the neuronal substrate of memory storage.
- Engram synapses
-
Activated synapses between pairs of engram cells that serve as a marker of memory formation.
- Fusion proteins
-
Proteins derived by combining two or more distinct protein domains that are encoded by separate genes but are transcribed and translated as a single polypeptide chain.
- Gal4 driver line
-
A transgenic line of fruitflies expressing the Gal4 transcription factor in specific cell populations or at specific times. Gal4 can activate the expression of target genes under the control of upstream activating sequence elements.
- Genome editing
-
A genetic engineering method in which the DNA of an organism or a cell is manipulated via the insertion, deletion or replacement of sequences in the genome.
- Guide RNAs
-
Specific RNA sequences that recognize the regions of interest within DNA and act as a guide for the Cas proteins that target precise DNA editing.
- Homology-directed repair
-
(HDR). A mechanism to repair a double-strand DNA break in cells by using a homologous sequence of sister chromatids as a repair template.
- Immediate early genes
-
(IEGs). Genes that are activated rapidly and transiently upon the neuronal activation in response to various cellular stimuli.
- Intrabodies
-
Antibodies targeting the endogenous proteins that interact within a cell.
- Non-homologous end-joining
-
(NHEJ). The predominant pathway for the repair of double-strand breaks in DNA, regardless of the sequence homology.
- Optogenetic actuators
-
Light-sensitive proteins that can control specific molecular or cellular activity.
- Reactive oxygen species
-
A type of highly reactive molecules and free radicals derived from molecular oxygen with one or more unpaired electrons in the outer orbital. ROS serves as signalling molecules in normal biological processes.
- Recombinases
-
Enzymes that catalyse site-specific DNA recombination events, which allow the exchange of DNA fragments.
- Retrograde tracing
-
The process by which neuronal processes are traced from the axon terminals towards the cell body.
- Synaptopathy
-
Dysfunction of the nervous system (including the brain, spinal cord and peripheral nervous system) that is related to altered synapse formation and function.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Timalsina, B., Lee, S. & Kaang, BK. Advances in the labelling and selective manipulation of synapses. Nat. Rev. Neurosci. 25, 668–687 (2024). https://doi.org/10.1038/s41583-024-00851-9
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41583-024-00851-9
This article is cited by
-
A brief history of insect neuropeptide and peptide hormone research
Cell and Tissue Research (2025)